Abstract
In the past decade, crinviruses have gained interest due to their rapid widespread and destructive nature for cucurbit cultivation. Several members of the genus Crinivirus are considered emerging viruses. Currently, four criniviruses: Beet pseudo-yellows virus, Cucurbit chlorotic yellows virus, Cucurbit yellow stunting disorder virus and Lettuce infectious yellows virus have been reported to infect field- or greenhouse- grown cucurbits. Apart from their cucurbit hosts, criniviruses infect other cash crops and weeds. Criniviruses are exclusively transmitted by whiteflies. The virion titer and the vector genus or species complex are predominant factors affecting virus transmission. These criniviruses maintain genetic stability with limited intra-species variability. They share similar core genome structure and replication strategies with some variations in the non-core proteins and downstream replication processes. Management of the diseases induced by criniviruses relies on integrated disease management strategies and on resistant varieties, when available. This review will cover their epidemiology, molecular biology, detection and management.
Keywords: Cucurbit yellow stunting disorder virus, Lettuce infectious yellows virus, Cucurbit chlorotic yellows virus, Beet pseudo yellows virus
Introduction
Cucurbits are susceptible to at least 59 plant viruses [44]. The majority of these viruses are vector-borne and transmitted by either aphids or whiteflies. Cucurbits are infected by many whitefly-transmitted viruses in the genera Crinivirus, Begomovirus and Ipomovirus. Viruses of the genus Crinivirus are emerging and considered an obstacle to crop production [61]. Crinviruses are single stranded RNA (ssRNA) viruses that belong to the family Closteroviridae which includes the genera Ampelovirus, Closterovirus, Crinivirus and the newly-proposed Velarivirus [8, 52]. The family Closteroviridae includes a diverse group of plant viruses based on their distinctive particle morphology, length, semi-persistent transmission by hemipteran vectors, phloem-limitation, cytopathology, genome organization and expression [37]. Criniviruses are transmitted by whiteflies; ampeloviruses and closteroviruses are transmitted by mealybugs and aphids, respectively; whereas velariviruses mode of transmission remains undetermined [8, 52]. So far, only four out of 14 criniviruses (listed in caption of Fig. 3) were reported to infect cucurbits; Cucurbit yellow stunting disorder virus (CYSDV), Cucurbit chlorotic yellows virus (CCYV), Beet pseudo yellows virus (BPYV) and Lettuce infectious yellows virus (LIYV). In the past decade the emergence of viruses in the crinivirus group has attracted researchers and growers due to their damaging effect on cultivated crops. Also, there is a lack of a comprehensive up to date review on the status of cucurbit-infecting criniviruses. In this paper, the epidemiology, biology and disease management of cucurbit-infecting criniviruses will be reviewed.
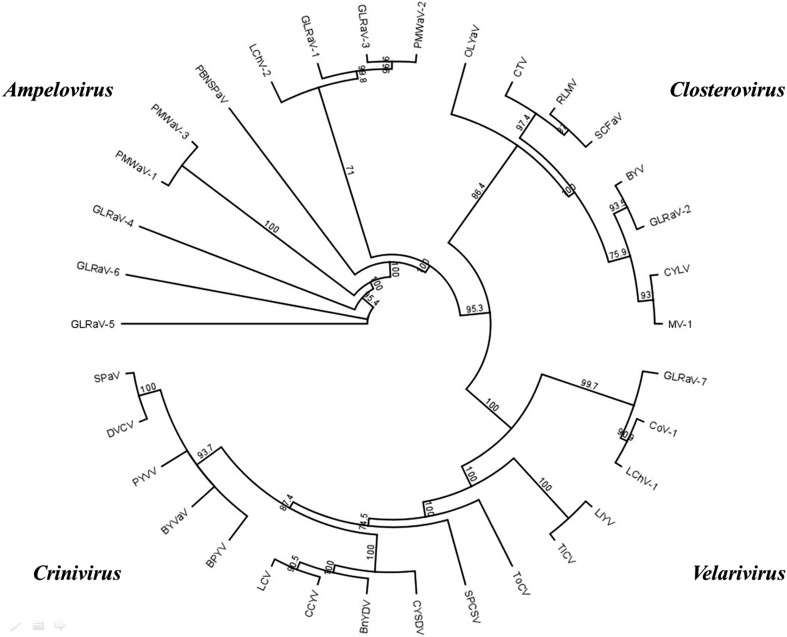
Fig. 3.
Unrooted phylogenetic tree showing the relationships between the species and genera of the family Closteroviridae based on the Hsp70h amino acid sequence. A multiple sequence alignment of the amino acid sequence of the Hsp70h was conducted using CLUSTALW. The phylogenetic tree was constructed using the neighbour joining method and bootstrapped 1000 times. The number on nodes represents bootstrap confidence percentages. Abbreviations of viruses and accession numbers are given in parentheses Bean yellow disorder virus (BnYDV; EU191905); Beet pseudo-yellows virus (BPYV; AY330919); Beet yellows virus (BYV; AF190581); Blackberry yellow vein-associated virus (BYVaV; AY776335); Carrot yellow leaf virus (CYLV; FJ869862); Citrus tristeza virus (CTV; U56902); Cordyline virus-1 (CoV-1; HM588723); Cucurbit chlorotic yellows virus (CCYV; AB523789); Cucurbit yellow stunting disorder virus (CYSDV; AJ439690); Diodia vein chlorosis virus (DVCV; GQ376201); Grapevine leafroll-associated virus-1 (GLRaV-1; JQ023131); Grapevine leafroll-associated virus-2 (GLRaV-2; AY881628); Grapevine leafroll-associated virus-3 (GLRaV-3; AF037268); Grapevine leafroll-associated virus-4 (GLRaV-4; FJ467503); Grapevine leafroll-associated virus-5 (GLRaV-5; JX559639); Grapevine leafroll-associated virus-6 (GLRaV-6; FJ467504); Grapevine leafroll-associated virus-7 (GLRaV-7; HE588185); Lettuce chlorosis virus (LCV; FJ380119); Lettuce infectious yellows virus (LIYV; NC_003618); Little cherry virus-1 (LChV-1; Y10237); Little cherry virus-2 (LChV-2; AF531505); Mint virus-1 (MV-1; AY792620); Olive leaf yellowing-associated virus (OLYaV; AJ440010); Pineapple mealybug wilt-associated virus-1 (PMWaV-1; AF414119); Pineapple mealybug wilt-associated virus-2 (PMWaV-2; AF283103); Pineapple mealybug wilt-associated virus-3 (PMWaV-3; DQ399259); Plum bark necrosis stem pitting-associated virus (PBNSPaV; EF546442); Potato yellow vein virus (PYVV; AJ557129); Raspberry leaf mottle virus (RLMV; DQ357218); Strawberry chlorotic fleck-associated virus (SCFaV; DQ860839); Strawberry pallidosis-associated virus (SPaV; AY488138); Sweet potato chlorotic stunt virus (SPCSV; AJ428555); Tomato chlorosis virus (ToCV; AY903448); Tomato infectious chlorosis virus (TICV; FJ815441). GLRaV-7, LChV-1, and CoV-1 are assigned to a tentative new species, ‘Velarivirus ’. OLYaV is an unassigned species in the family
Epidemiology of cucurbit-infecting criniviruses
Viral distribution and economic impact
Crinviruses are categorized as emerging viruses and their rapid spread is attributed to the rapid increase and spread of whitefly populations. This upsurge of whitefly populations may be correlated with their resistance to pesticides and global warming [61]. Indirectly, man-made actions exacerbate virus spread through intensified agricultural practices and introduction of exotic pests and viruses through movement of vegetative materials [61].
From the epidemiological and economical point of view, CYSDV is the most threatening to cultivated cucurbits, whereas LIYV is the least significant. CYSDV was first reported in the United Arab Emirates in the early 1990s [32] and has spread globally to regions in North America, North Africa, Europe and Asia over the past two decades. In the new world, CYSDV was initially reported in southern Texas and northern Mexico in 1999, but recently it has surfaced as a widespread emergent virus in several southern and western US states [13, 36, 42, 70].
In 2004, yellowing symptoms in melon and cucumber were observed in Japan and few years later, CCYV was identified as the causal agent [29]. This virus has recently moved from Japan in eastern Asia to Sudan, Lebanon and Iran [5, 30]. So far CCYV seems to be spreading competitively with CYSDV in Lebanon; where about 60 % of the cucumber samples collected in 2012/2013 showed mixed infections of CCYV and CYSDV (P.E. Abrahamian and Y. Abou-Jawdah, unpublished data).
BPYV is the earliest cucurbit infecting Crinivirus isolated from the Salinas valley in California [94]. BPYV later spread through southern Europe, where its vector, the greenhouse whitefly (Trialeurodes) predominated. However, following its initial establishment, reports from southern Spain showed the progressive displacement of BPYV by CYSDV [11]; under the prevailing climatic conditions, Bemisia tabaci, the more polyphagous vector, was more competitive than Trialeurodes spp. [94].
LIYV has been reported only in North America and its incidence has decreased drastically to negligible levels [23, 94]. This was attributed to the introduction of an exotic B. tabaci MEAM-1 (B) species complex which completely displaced the indigenous LIYV vector, B. tabaci New World species complex (biotype A) [73, 94]. CYSDV is the most widely spread followed by BPYV and CCYV (Table 1).
Table 1.
Major characteristics and geographical distribution of cucurbit infecting criniviruses
| Virus | Acronym | Non-cucurbit affected cropsa | Vectorb | Persistence Days |
Distribution | References |
|---|---|---|---|---|---|---|
| Cucurbit yellow stunting disorder virus | CYSDV | Alfalfa, lettuce, snap bean | Bemisia tabaci MEAM-1 (B)/MED (Q) | 9 | Africa (Egypt, Morocco, Sudan Tunisia), Asia (China, United Arab Emirates, Saudi Arabia, Iran, Turkey, Syria, Jordan, Israel, Lebanon), Europe (Cyprus, Franced, Greece, Spain, Portugal), North America (Mexico, USA—California, Arizona, Texas, Florida) | [3, 10, 12, 15, 19, 20, 22, 32, 36, 42, 43, 46, 48, 68, 70, 91, 95] |
| Cucurbit chlorotic yellows virus | CCYV | Lettuce, spinach, sugarbeet | Bemisia tabaci MEAM-1 (B)/MED (Q) | Unknown | Asia (China, Iran, Japan, Lebanon, Taiwan), Africa (Sudan) | [5, 10, 28, 30, 34, 64, 98] |
| Beet-pseudo yellows virus | BPYV | Carrot, endive, lettuce, flax, spinach, sugarbeet, strawberry | Trialeurodes vaporariorum | 7 | Asia (Japan), Europe (Bulgaria, Crete, Cyprus, France, Greece, Italy, Netherlands, Spain), North America (Canada–British Columbia, Costa Rica, USA—California, Florida, Oregon), Oceania (Australia, New Zealand) | [11, 12, 17, 31, 68, 71, 73, 84, 85, 94] |
| Lettuce infectious yellows virus | LIYV | Carrot, lettuce, radish, spinach, sugarbeet | Bemisia tabaci New World (A)/MEAM-1 (B)c | 3 | North America (USA—California, Arizona) | [23, 94] |
aThe crops indicated are either susceptible under natural or experimental conditions
bAcronyms following B. tabaci are species complexes whereas letters indicated in parenthesis refer to the biotype
cThe efficiency of transmission by MEAM-1 (B) is very low
dEradicated in 2003 and 2004
All four criniviruses have been reported to infect cucurbit crops with moderate to high incidence [12, 13, 23, 28]. CCYV and CYSDV may occur in mixed infections [5, 10]. In terms of economic losses, all criniviruses cause significant yield reductions and impede plant growth. For example, in cucumber CYSDV may cause a 30–50 % yield reduction whereas in melon losses up to 80 % have been reported [3, 13, 47].
CCYV, an emerging crinivirus, causes lower yield losses in melon than CYSDV and range between 10 and 20 % [28]. Economic losses due to BPYV infections may fluctuate according to the cropping season and vector populations which may affect disease severity and virus concentration in plants. For example, significant decrease in fruit weight and a high incidence of abortion has been observed in pumpkin fields [89]. Currently, LIYV has been eliminated as a threat to cucurbit production in the US after the elimination of its indigenous vector [94].
Host range and symptomology
In the Cucurbitaceae family, several major cucurbits such as melon (Cucumis melo), cucumber (C. sativus), winter squash (Cucurbita maxima); summer squash (C. pepo), pumpkin (C. moschata) and watermelon (Citrullus lanatus) are susceptible to CYSDV, CCYV, LIYV and BPYV [12, 15, 23, 64]. Also, several cucurbit-infecting criniviruses have been detected in weeds and non-cucurbitaceous economic crops (Table 1) [23, 64, 71, 91].
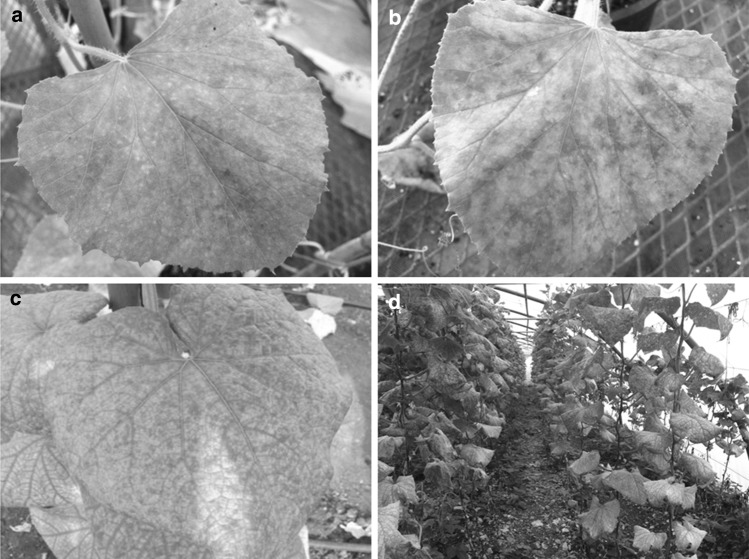
The difficulty in differentiating among viruses based on symptoms is a common characteristic of criniviruses. All four criniviruses induce similar yellowing symptoms on cucurbits. Interveinal chlorotic spots appear on lower leaves and coalesce to give a bright yellow color, while leaves curl downward slightly, remain turgid and become brittle (Fig. 1). On cucumber and melon, yellowing symptoms are evident after three to 4 weeks post-inoculation. Yellowing becomes more severe as leaves grow older. Symptoms spread towards new growth but never reach the young leaves [3]. Fruit shape and color are not altered but fruit number, total weight and sugar content are significantly reduced [3, 47, 65]. Other cucurbits, including melon and watermelon, show similar yellowing symptoms on lower leaves. According to Gil-Salas et al. [27], zucchini squash is a symptomless host of CYSDV. The yellowing symptoms induced by mixed viral infections of CCYV and CYSDV in melon or cucumber are indistinguishable from singly infected plants with either virus [5].
Fig. 1.
Symptoms caused by crinivirus infection on cucurbits under experimental (a, b, c) or natural conditions (d). The early (a) and advanced (b) symptom development of Cucurbit chlorotic yellows virus (CCYV) are shown on melon, whereas advanced symptoms of Cucurbit yellow stunting disorder virus (CYSDV) on cucumber (c). Cucumber-grown in a commercial greenhouse showing a high incidence of yellowing symptoms caused by criniviruses (d)
Yellowing symptoms may often be confused with physiological and/or nutritional disorders, effects of extreme soil pH, micronutrient deficiencies, natural senescence, and pesticide phytotoxicity. As a result, the visual recognition of criniviruses as causal agents of yellowing symptoms is difficult for growers and researchers [94]. The key distinguishing characteristics of CYSDV infection are leaf thickening and brittleness.
Criniviruses induce similar cytopathological effects in plants characterized by altering the chloroplasts and by the presence of Beet yellows virus (BYV)-type inclusion bodies in companion cells [56]. Sieve tubes and vascular parenchyma cells harbor scattered virus-like particles whereas companion cells show large masses of cross-banded inclusions [56]. A unique feature of LIYV cytopathology characteristics is the formation of conical plasmalemma deposits [56]. Detection of the CYSDV RNA in melon infected plants by in situ hybridization, showed virus specific signals in the vesicular phloem of vascular bundles, particularly in the bundle-associated cells [50]. High virus signals in the bundle-associated cells of CYSDV-infected cucumber and melon were also observed using in situ serological assays [33]. Unexpectedly, in situ immunostains showed that CCYV localizes in the leaf lamina and not only near the leaf veins [41].
Transmission
Criniviruses are transmitted in the semi-persistent manner and are retained in their vector for a relatively short time (Table 1). CYSDV and CCYV are transmitted by the sweet potato whitefly, B. tabaci, MEAM-1 (B) and MED (Q) species complexes [11, 15, 23, 29], while the New World (A) is an inefficient vector of CYSDV [93]. B. tabaci New World (A) is a vastly better vector than MEAM-1 (B) for transmission of LIYV [93, 94]. Unlike the former three viruses, BPYV is transmitted by the greenhouse whitefly, Trialeurodes vaporariorum. CCYV transmission parameters (acquisition and inoculation feeding access periods and persistence) have not been studied yet, however, they might be more likely similar to Lettuce chlorosis virus (LCV), based on gene homology of the transmission determinants, coat protein (CP), 76.9 %, and minor coat protein (CPm), 59.7 %. CYSDV transmission efficiencies ranged between 85 and 100 % in two different studies using a high number of whiteflies [11, 15]. Such variation may be due to the age, gender and feeding behavior of vector in addition to virus genetic variations.
In vitro transmission studies for members of Closteroviridae family are usually difficult to achieve and reproduce. Nevertheless, whiteflies were capable of feeding and acquiring LIYV in vitro from purified virions [62]. This implicated a capsid strategy, which is the direct attachment of virions to vector cuticles, but not a helper strategy involved in the recruitment of a viral protein as an intermediate factor for virion retention [62].Consistent transmission was also obtained by whiteflies feeding through membranes on purified virions isolated from infected protoplasts or plants [62]. Virion concentrations as low as 0.1 ng μl−1 resulted in low transmission efficiency in contrast to high efficiency with 100 or 1000 ng μl−1 using the same number of whiteflies [62]. Also, whitefly population directly affected transmission efficiency thus showing that transmission is dependent on whitefly numbers and virion concentrations [62]. The semi-persistent mechanism of transmission has been reviewed by Ng and Falk [62].
Molecular biology
Genome organization
LIYV, the type species of the genus Crinivirus, has been extensively studied at the molecular level. Therefore, the LIYV genome organization and genes implicated in virus replication and pathogenicity will be discussed. The similarities and differences among the genes in the four criniviruses will be described and discussed (Fig. 2). Crinivirus genomes consist of a linear, positive sense, ssRNA 15.6–17.9 kb in size, segmented into two (or three) molecules that are needed for infectivity and are separately encapsidated [52]. Virion particles of cucurbit criniviruses are flexuous, filamentous, with cross-banding and open helix pattern; their length and diameter range between 650–850 and 12–14 nm, respectively [45]. The genome is structurally divided into two parts: RNA1 and RNA2. RNA 1 encodes proteins associated with RNA replication, whereas RNA 2 typically contains a five gene hallmark array of the family Closteroviridae, which include a ~5 or 6 kDa hydrophobic protein (p5/6), heat shock protein 70 homolog (Hsp70h), CP (59 kDa), CPm and a 59 kDa protein (p60) [21, 52].
Fig. 2.
Genome organization of the Crinivirus type member Lettuce infectious yellows virus (LIYV; RNA 1: NC_003617; RNA 2: NC_003618), Cucurbit yellow stunting disorder virus (CYSDV; RNA 1: AJ537493; RNA 2: AJ439690), Cucurbit chlorotic yellows virus (CCYV; RNA 1: AB523788; RNA 2: AB523789) and Beet pseudo-yellows virus (BPYV; RNA 1: AY330918; RNA 2: AY330919). ORF 1 is divided into two parts 1a and 1b, where the latter is expressed via a +1 ribosomal frameshift. The names of putative proteins are given by the letter ‘P’ followed by the molecular weight × 103. P-Pro papain-like protease, MTR methyltransferase, HEL helicase, RdRp RNA-dependent RNA polymerase, Hsp70h heat shock protein 70 homolog, CP coat protein, CPm coat protein minor
LIYV RNA 1 is 8118 nucleotides (nt) long and encodes for three open reading frames (ORFs): ORF1a, ORF1b and ORF2. The first ORF yields a polyprotein (217 kDa) which encodes a papain-like protease (P-Pro), methyltransferase (MTR) and RNA helicase (HEL) [39]. Conserved amino acid residues in P-Pro directly cleave the polyprotein into a putative proteinase [7]. RNA-dependent RNA polymerase (RdRp, 55 kDa) expression from ORF1b is a result of a+1 frameshift translation mechanism. The resulting LIYV polyprotein (276 kDa) is slightly smaller than those of CYSDV (289 kDa) and CCYV (285 kDa). Downstream ORF1b, LIYV encodes a putative 34 kDa protein (ORF 2) which is a trans enhancer for replication and accumulation of RNA 2, but not RNA 1 [97]. Moreover, p34 binds to ssRNA in a sequence non-specific manner [87]. Yet, no similarity to p34 is found among the 3′-RNA 1 proteins encoded by the three other criniviruses. Unlike LIYV, CYSDV encodes other RNA 1, 3′-proximal proteins such as p5 (ORF 2), p25 (ORF3) and p22 (ORF 4) (Fig. 2). The small hydrophobic protein (CYSDV-p5) includes an N-terminal signal peptide likely targeting the endoplasmic reticulum [7]. The CYSDV p25 protein has been identified as a suppressor of post-transcriptional gene silencing (PTGS) in plants [38]. Nevertheless, p25 is a weak suppressor of PTGS as compared to RNA 1-encoded p22 of Tomato chlorosis virus (ToCV) [38]. It is likely that CYSDV-p25 does not interact with siRNAs, as Sweet potato chlorotic stunt virus (SPCSV)-encoded RNase III, due to the high amount of recovered short-interfering RNAs (siRNAs) [38]. Moreover, p25 does not prevent the spread of the silencing signal. While the study only screened RNA 1-encoded viral genes it is likely that RNA 2 has additional silencing suppressors. The CYSDV-p22 might have a similar function to SPCSV RNase III in PTGS [38]. CCYV and BPYV both encode putative proteins of low molecular weight with transmembrane (TM) domains [64, 83]. Additionally, CCYV encodes a 22 kDa protein (ORF 3) similar in size to ToCV-p22 [64].
LIYV RNA2 is 7193 nt long and encodes seven ORFs: p5 (ORF1), heat shock protein 70 homolog (Hsp70h; ORF 2), p59 (ORF3), p9 (ORF 4), CP (ORF 5), CPm (ORF 6) and p26 (ORF 7) [39]. It should be noted that the small putative protein(s) of the quintuple gene block varies in position and size among crinivirus species. LIYV possess a p5 hydrophobic protein with a TM domain. P5 might be a movement protein that localizes to the ER [21]. Hsp70h is a multi-functional protein which possesses several conserved motifs such as those involved in ATPase activity [39]. The crinivirus-encoded Hsp70h may be associated with virion tail assembly and disassembly similar to the closterovirus Beet yellows virus (BYV) [9]. Further, Hsp70h may be involved in plasmodesmatal attachment and virion translocation through ATPase activity [9]. However, the precise functions remain unknown for crinivirus-encoded Hsp70h. P59 is also implicated in tail assembly and may be associated in virion movement [60]. The CP encapsidates and protects the RNA from cellular degradation. The CPm is a whitefly-transmission determinant needed for plant-to-plant transmission for LIYV [16, 79, 81]. CPm constitutes an integral part of the tail protein complex, whereby CPm binds to the anterior foregut of whitefly epithelial cells [79]. Examination of CYSDV, BPYV and CCYV RNA 2 genome organization showed similar features to LIYV except for an additional small 6 kDa putative protein (p6) positioned between Hsp70h and p59. This is common for criniviruses where several mysterious ORFs and gene shuffling occurs in the ‘quintuple gene block’ [21]. Surprisingly, the small putative proteins (p4.9 and p6) do not have any TM regions, whereas only one TM region (amino acid position 187–207) was found in p26. The p26 protein was present in the membranes and cell wall protein fraction [57]. The p26 showed to aggregate, localize and induce production of plasmalemma deposits [57, 78]. Under the same expression system, CYSDV and BPYV 26-kDa proteins exhibit different localization patterns, although they share similar biochemical properties to LIYV p26 [78].
In the terminal sequence region, LIYV RNA 1 includes a 97 and 219 nt as 5′ and 3′ un-translated region (UTR), respectively [39]. LIYV RNA 2 has a 326 and 191 nt as 5′ and 3′ UTRs, respectively. The first five nucleotides are identical in the 5′-UTRs of RNA 1 and RNA 2 in addition to a stretch of 23 nucleotides extending into ORF 1a of RNA 1. On the contrary, the LIYV 3′-UTRs of both RNAs do not share significant homology as compared to CCYV, CYSDV and BPYV 3′-UTRs. Further, the 3′-UTR region of the latter criniviruses have variable secondary structure which may have different functional requirements [7, 64, 83].
Following inoculation, virions disassemble and replicate in the cytoplasm. Virions possibly associate with ER-derived membranous vesicles and vesiculated mitochondria [52]. Replication is believed to be similar to other closteroviruses, where the genome expression is based on several strategies. Initially RNA 1 is directly translated into the polyprotein encoded by ORF1a/b followed by the +1 ribosomal frameshift for the expression of the RdRp domain encoded by ORF1b. Subsequently, coterminal 3′-ORFs are expressed through subgenomic RNA. RNA 1 and RNA 2 replication and accumulation occurs through cis- and trans-preferential replication, respectively [88, 97]. RNA 1 does not support trans replication of artificial defective RNA (D-RNA) derived from RNA 1 [88]. LIYV-encoded p34 binds to RNA 2 and subsequently enhances RNA 2 accumulation levels [87]. Accumulated RNA is assembled with viral proteins into virions and translocated into adjacent cell to initiate a new replication cycle [reviewed in 21]. A byproduct of LIYV replication is the de novo production of non-interfering defective (D)-RNAs derived exclusively from genomic RNA 2 [74]. The absence of D-RNA derived from genomic RNA 1 is due to the cis-preferential replication which eliminates the accumulation of these potential D-RNAs [88]. These RNA 2-derived D-RNA species are vector transmissible, encapsidated, capable of replication in trans in the presence of RNA 1 and expression of intact 3′-ORFs [62, 74, 96].
LCV RNA 2 replication enhancement in trans is supplemented by LCV-encoded p23 which is similar in function but not homologous to LIYV p34 [75]. However, unlike LIYV, RNA replication kinetics proceed in symmetrical accumulation of LCV genomic RNA probably due to the highly homologous 3′ UTR ends between both RNA segments [75]. It is likely that each of the closely related CCYV, CYSDV and Bean yellow disorder virus (BnYDV), which possess highly homologous 3′ UTRs, replicate in the same manner of LCV. LIYV replication mechanism is not universal as evident by the de novo generation and replication in trans of D-RNA derived from LCV RNA 1 [58].
Genetic variability
CYSDV genetic diversity has been more extensively studied among the four criniviruses. Early studies based on single-strand conformational polymorphism (SSCP) and sequencing of the Hsp70h and CP gene of CYSDV isolates from different geographical areas showed slightly distinct CP and Hsp70h patterns [51, 72, 73, 80]. Hsp70h-based analysis showed that CYSDV population in the same plant was closely related but had one predominant variant independent of the host [73]. Analysis of CYSDV CP showed that the studied isolates had low genetic diversity in the CP and complemented previous studies [72]. Thus, CYSDV could be divided into two divergent subpopulations: a ‘western’ (isolates from Jordan, Lebanon, North America, Spain, France, Tunisia, Greece and Turkey) and an ‘eastern’ subpopulation (isolates form Saudi Arabia and Iran) [12, 72, 73, 80, 95]. Five coding and one non-coding genomic regions of a CYSDV population were examined in different cucurbit hosts in Spain [51]. CYSDV showed high genetic stability with negligible nucleotide diversity over a period of 8 years regardless of the genomic region or host [51]. Exceptionally, the low genetic variability resulted in positive selection of an amino acid in the CP [51]. A suggested model infers that the initial few CYSDV genotypes (in Spain) were rapidly dispersed through whiteflies and continuous cucurbit cultivation which hindered the generation of new variants [51].
Currently, two fully sequenced BPYV strains exist, separately isolated from strawberry and cucumber, with striking difference in genome structure [31, 83]. In the past, several strains such as melon yellows virus and cucumber chlorotic spot virus have been reported as distinct species, nevertheless they are now considered as BPYV variants [94]. Similarly, Cucumber yellows virus, is currently recognized as a strain of BPYV with amino acid identities ranging from 94 to 100 % [83]. Two major differences are an insertion in ORF1a and an additional 3′-terminal ORF in RNA 1 [83]. The BPYV Hsp70h showed a homogenous population among a few number of isolates [73]. BPYV isolates recovered from cucurbits in Costa Rica showed high Hsp70h or CPm homology to the strawberry strain [71]. The distribution of these dissimilar strains remains unclear due to the low number of BPYV isolates analyzed.
CCYV genetic variability has not been studied yet. A low number of available sequences of full CCYV genomes or partial genes show limited variability. Unexpectedly, CCYV isolates from Iran show the only significant difference from all other distant geographic isolates at about 5 % nt difference [10]. Nevertheless, the partial Hsp70h sequence of the Iranian isolate is not enough for further assumptions. It is likely that CCYV has a similar genetic pattern to CYSDV (i.e. subgroups Western and Eastern) but with potentially different subgroup naming. LIYV genetic variability was not studied probably due to its restricted distribution and currently reduced incidence. Several factors contribute to the relatively low genetic variability of these criniviruses such as negative selection pressure and constraints on RNA secondary structure to maintain virus function and genetic bottlenecks caused by founder effects [51, 59].
Phylogenetic relationships
In general, RNA 2 genome structure is conserved among criniviruses with some slight reshuffling and presence of small ORFs, whereas RNA 1 has different ORFs downstream the replicase polyprotein. Sequencing data revealed that criniviruses contain limited genetic variability [7, 37]. Phylogenetic analyses were carried out on the amino acid sequences of the conserved Hsp70h gene of clostero-, ampelo-, velari- and criniviruses (Fig. 3). Phylogenetic relationships based on RdRp and CP amino acid sequences were similar with slightly different clustering within the genus Crinivirus only (not shown). Phylogenetic trees are in complete agreement with biological observations where whitefly-transmitted viruses cluster in a distinct group from Closterovirus, Ampelovirus and Velarivirus.
The Hsp70h and RdRp are highly conserved genes and possess several signature motifs [37]. The homology among Hsp70h nucleotide and amino acid pairwise identity of crinivuruses was at 61.3 and 61.7 %, respectively (accession numbers of sequences used are provided in Fig. 3). Similarly, RdRp showed a 61.5 and 61.3 % of nucleotide and amino acid pairwise identity, respectively. Multiple sequence alignment of Crinivirus CP and CPm sequences revealed only limited amino acid identities as compared to Hsp70h. A striking feature is that CPm size varied among criniviruses and ranged between 400 and 600 amino acids [21].
Phylogenetic analysis showed that CCYV and CYSDV are more closely related to each other than to LIYV or BPYV. According to nucleotide substitution rate, CYSDV appears to be more closely related to CCYV and to two other non-cucurbit infecting criniviruses: BnYDV and LCV. Cucurbit- and non-cucurbit-infecting genomes of CYSDV, BPYV, CCYV, LCV, BnYDV, SPCSV, ToCV and Blackberry yellow vein-associated virus share a related feature of having very similar 3′ UTRs in RNAs 1 and 2 [7, 31], although for CYSDV, similarity in primary structures did not result in predictions of equivalent secondary structures. CYSDV, CCYV, LCV and BnYDV share 5 homologous nucleotides at each of the 5′ and 3′ UTR of RNA 1 and RNA 2. BnYDV, a biologically dissimilar virus, appears to be a relative of CYSDV, and shares with it, high homologies in the RdRp, CP, Hsp70h and HEL amino acid sequences [53]. On the contrary, CCYV possesses similar symptom expression and similar host range to CYSDV but shares less homology in the Hsp70h and CP amino acid sequences [64]. Phylogenetic analyses based on conserved genes of the RdRp, CP and Hsp70h, suggested dividing criniviruses into three groups based mainly on vector transmission [86, 92]. Groups 1 and 2 are transmitted by T. vaporariorum and B. tabaci vector, respectively [92]. On the other hand, group 3 is transmitted by different vectors but share many common hosts. CYSDV and CCYV belong to group 2, whereas LIYV belongs to group 3.
Diagnosis
Due to the phloem limitation of criniviruses and their overall low titer in infected plants [94], it is often recommended to rely on more sensitive nucleic acid detection methods, than on serology-based methods. In the past, extraction and purification of virions for antibody production was difficult. However, the utilization of recombinant technology has eased antibody production. Several studies developed antibodies for CYSDV and CCYV CP expressed in E. coli cells [33, 41]. Recently, Steel et al. [77] introduced a novel technique to produce CYSDV CP using “Agrobacterium tumefaciens-mediated transient expression system” in plants. These antibodies have been used successfully in DAS-ELISA, plate trapped antibody-ELISA (PTA-ELISA), tissue-blot immuno assay (TBIA), dot-blot immuno assay (DBIA), immuno-capture reverse transcription polymerase chain reaction (IC-RT-qPCR) and immuno-electron microscopy (IEM) [33, 77]. In general, TBIA is a preferred technique for rapid field surveys and may be just slightly less sensitive than RT-PCR [1, 82].
Double-stranded RNA (dsRNA) analysis serves as a preliminary diagnostic procedure; however, gel interpretations can often be erroneous. Two highly conserved genes, the RdRp and Hsp70h, of the Closteroviridae serve as good candidates to design primers for general detection of criniviruses by RT-PCR. Degenerate primers were used for early detection of criniviruses and followed by other techniques, such as RFLP and sequencing, to identify the specific virus. At the fast rate of emergence of new criniviruses, new detection assays should be developed and evaluated. A recent set of degenerate primers has been designed and tested in different hosts for detection of several criniviruses based on the RdRp gene [90]. BPYV specific primers targeting two regions of the Hsp70h and CP were compared in a wide survey [70]. Primers based on Hsp70h gene were capable of detecting more (30–50 % higher chance) BPYV samples than those of the CP gene [70]. CCYV and CYSDV may often occur in mixed infections, specific primers were designed to distinguish between the two infections in multiplex RT-PCR [4].
In the past few years, the use of real-time PCR for diagnosis, quantification and genotyping has increased considerably. RT-qPCR assays have been developed for CYSDV and CCYV but not for LIYV or BPYV. These assays are either based on the TaqMan® or SYBR Green chemistry [1, 26, 67]. RT-qPCR has been used for sensitive virus detection and quantitation of viral titer. Gil-Salas et al. [26] developed an assay for CYSDV and Cucumber vein yellowing virus (CVYV) detection. Subsequently, the same group quantified CYSDV titer in co-infection with CVYV [27].
Disease management
Integrated disease management
CYSDV is considered a plant quarantine pest in several regions, for example in France, in 2002 and 2003, rapid eradication measures were undertaken [63, 66]. The European and Mediterranean Plant Protection Organization (EPPO) listed CYSDV under its A2 action list no. 324 whereas LIYV is listed under its A1 list [63].
Cultural practices, such as crop free periods, crop residue disposal, planting dates, weed removal, mulches and irrigation can reduce whitefly populations and subsequently delay or reduce crinivirus incidence. Crop free periods proved to be successful in significantly lowering the incidence of LIYV by maintaining a host-free period in North America [94]. In addition, the use of insect-proof nets and double doors reduced CYSDV incidence in greenhouses, whereas field-grown cucurbits gave higher yields when shielded with floating row covers (e.g. Agryl®) [2, 3, 35]. Colored plastic mulches in a variety of colors, including aluminum, silver, transparent, white and yellow, have been proven to be effective in reducing the incidence of whitefly transmitted viruses [76].
An important aspect is to reduce vector populations either through biological or chemical control. Some potent bio-control agents include the entemopathogenic fungus Beauveria bassiana and the parasitoid insects Encarsiapergandiella and Eretmocerusmundus [25]. In general, chemical control of the vectors of closteroviruses has not been effective in preventing the spread of the diseases they cause [11]. Whiteflies have often developed rapid resistance to commercially used chemicals and rendered many insecticides ineffective for their control [49]. A mixture of novel insecticide groups were tested as soil or foliar application and showed potential reduction of whitefly populations and CYSDV incidence [14]. However, the control achieved in the latter study is a matter of time before whiteflies gain insecticide resistance. Several management strategies can be followed during the growing season as recommended by the University of California (Davis) extension guidelines (http://www.ipm.ucdavis.edu/PMG/r116100211.html).
Resistant germplasm
Most of the germplasm tested were of melon and cucumber origin and focused on screening for resistance to CYSDV. Nevertheless, none of the germplasm tested showed immunity but a few were either resistant or tolerant. Generally, screening for resistance relies mainly on two factors: symptom development and virus accumulation. Germplasm screening showed that CYSDV inoculated Cucumis melo TGR-1551 was asymptomatic with virus titers in whitefly-inoculated leaves significantly lower than in leaves of susceptible cultivars [47, 50]. Furthermore, at 6 weeks post-inoculation, CYSDV was not detected in upper leaves of TGR-1551 plants [50]. In grafting experiments, TGR-1551 scion grafted on CYSDV-infected root stock remained asymptomatic up to 28 days after grafting [50]. Nevertheless, positive and negative CYSDV RNA strands were detected in the scion of grafted plants but virus titer decreased gradually from the grafting point [50]. In situ localization showed that CYSDV was occasionally present in petiole and stem tissue but absent from leaf veins [50]. Thus, the suggested resistance involves restriction of virus movement into the vascular system of the plants but not a replication-mediated mechanism [50].
Breeding trials showed that hybrids (F1) of TGR-1551 crossed with a susceptible cultivar were resistant to CYSDV, suggesting that the resistance in TGR-1551 is controlled by a single dominant gene ‘Cys’ as shown in F1 and F2 populations [47]. On the contrary, Sinclair [cited in 55] noted that a Texas strain of CYSDV broke resistance of TGR-1551, and suspected that TGR resistance may be co-dominant or more complex. This was also reported in F2 populations from crosses between TGR1551x‘Deltex’ [69]. A somewhat promising germplasm is PI 313970, C. melo var. acidulus, a salad-type melon, which expressed high resistance level against CYSDV, LIYV and against the sweet potato whitefly [54, 55]. Resistance of PI 313970 was found to be controlled by a single recessive gene for CYSDV and a dominant gene for LIYV [54, 55]. However, their exotic fruit characteristics and sugar content may be a limiting factor for breeding programs where extensive backcrossing is needed to reach a resistant cultivar possessing marketable fruit features.
In a cucumber germplasm screening trial, three accessions were highly tolerant to CYSDV and four other accessions tolerant “under moderate disease pressure” [24]. Tolerant varieties showed milder symptoms with a lower infection rate, but only one accession showed significantly lower virus concentration [1, 24]. Simultaneously, cucumber accessions A1 and A2 exhibited partial resistance, lower virus concentration, and no symptoms under high inoculum pressure [6]. Furthermore, accession A2 exhibited antixenotic characteristics similar to TGR-1551 [6]. Another accession, PI 250147 ‘Khira’ originating from Pakistan showed resistance, but a cross between ‘Khira’ and a commercial variety gave a fully susceptible population, indicating a recessive nature of genetic inheritance [18]. Yet, none of these cucumber accessions show resistance similar to their melon counterpart, TGR-1551. Some cucumber cultivars resistant to CYSDV are commercially available [44], for example the cucumber cultivar ‘Jungla RZ F1’ is resistant to CYSDV and another whitefly-transmitted ipomovirus (Rijk Zwaan, Netherlands). Nevertheless, it would be interesting to study if the co-infection of CCYV and CYSDV would break resistance.
More recently, screening efforts of 51 melon germplasms showed only five promising accessions with mild CCYV symptoms. However, only one germplasm (JP 138332), an Indian melon, showed significant reduction of CCYV RNA levels [65].
Breeding programs may be accelerated by using molecular marker assisted selection. Park et al. [69] have identified 8 random amplified polymorphic DNA (RAPD) markers linked to quantitative trait loci (QTL) in TGR-1551 and ‘Deltex’ for CYSDV resistance. However, only one marker, derived from TGR-1551, was highly associated with resistance and accounted for 25 and 19 % of the phenotypic variation under natural and controlled inoculation conditions, respectively. QTL analysis of an F2 population, derived by crossing the cucumber accession PI 250147 with an elite breeding line, revealed a QTL responsible for 42 % of phenotypic variation in CYSDV resistance [18].
Genetic engineering may present an efficient tool for introducing disease resistance in cases where natural resistance is not present in cultivated or wild crop relatives or where resistance is governed by complex genetics. Pathogen-derived resistance has been applied successfully to develop transgenic cucurbits resistant to some aphid transmitted viruses including Cucumber mosaic virus, Watermelon mosaic virus 2, and Zucchini yellow mosaic virus [44]. To our knowledge no reports were published so far on transgenic cucurbits resistant to criniviruses. Nevertheless, RNA silencing-mediated resistance to the crinivirus Sweet potato chlorotic stunt virus (SPCSV) was successfully obtained in regenerated plants transformed with an intron-spliced hairpin construct targeting the replicase encoding gene. Upon SPCSV inoculation of the transformed plants, no symptoms or only mild symptoms were observed and SPCSV accumulation was significantly reduced [40].
Future prospects
Several aspects on cucurbit crinivirus biology and genetics are still lacking. For instance, a better insight on genetic variability is needed for other criniviruses (e.g. CCYV and BPYV) through multi-locus typing of isolates from different geographic regions. Also, it would be interesting to study the effect of mixed infections between criniviruses and other whitefly transmitted viruses such as geminiviruses or aphid-borne viruses. The replication mechanism briefly discussed here may be loosely applied to other criniviruses, thus the study of replication mechanisms in CYSDV or CCYV would be of interest. The lack of infectious clones for CYSDV, BPYV and CCYV hinder reverse genetics studies to reveal the function of unique viral genes. Also, elucidating the mechanism of crinivirus-host interaction at the cellular levels is still entirely unknown, such as host factors involved in movement and replication if any are involved. Subsequently, the results of these studies should aid on devising control strategies of economically emerging viruses such as CYSDV and CCYV. In addition to strict quarantine measures, the control of criniviruses currently focuses on a traditional integrated pest management approach. The commercialization of CYSDV tolerant or resistant varieties may pave the way for the development of cultivars with resistance to other cucurbit-infecting criniviruses. Screening for additional sources of resistance should be sustained. Although genetic engineering has been successful in generating cucurbits resistant to some aphid transmitted viruses, its efficacy is still not demonstrated for cucurbit criniviruses. Therefore, optimized plant regeneration protocols and efficient transgene constructs are needed. However, it should be noted that transgenic resistance to the crinivirus, SPCSV, has been successfully implemented in sweet potato with a high transformation frequency and biosafety [40]. Criniviruses should be given special attention due to their capability of infecting a wide range of herbaceous plants and rapid transmission by whiteflies. The rate at which new criniviruses are being reported is remarkable.
Acknowledgements
The research on criniviruses has been partially funded by Grants from USAID and USDA. We would like to thank Dr. Elie Barbour for his critical review of the manuscript.
References
- 1.Abou-Jawdah Y, Eid SG, Atamian HS, Havey M. Assessing the movement of Cucurbit yellow stunting disorder virus in susceptible and tolerant cucumber germplasms using serological and nucleic acid-based methods. J Phytopathol. 2008;156:438–445. doi: 10.1111/j.1439-0434.2007.01388.x. [DOI] [Google Scholar]
- 2.Abou-Jawdah Y, Sobh H, El-Zammar S, Fayyad A, Lecoq H. Incidence and management of virus diseases of cucurbits in Lebanon. Crop Prot. 2000;19:217–224. doi: 10.1016/S0261-2194(99)00100-3. [DOI] [Google Scholar]
- 3.Abou-Jawdah Y, Sobh H, Fayyad A, Lecoq H, Delecolle B, Trad-ferre J. Cucurbit yellow stunting disorder virus- a new threat to cucurbits in Lebanon. J Plant Pathol. 2000;82:55–60. [Google Scholar]
- 4.Abrahamian PE, Seblani R, Sobh H, Abou-Jawdah Y. Detection and quantitation of two cucurbit criniviruses in mixed infection by real-time RT-PCR. J Virol Methods. 2013;193:320–326. doi: 10.1016/j.jviromet.2013.06.004. [DOI] [PubMed] [Google Scholar]
- 5.Abrahamian PE, Sobh H, Abou-Jawdah Y. First Report of Cucurbit chlorotic yellows virus on Cucumber in Lebanon. Plant Dis. 2012;96:1704. doi: 10.1094/PDIS-05-12-0486-PDN. [DOI] [PubMed] [Google Scholar]
- 6.Aguilar JM, Abad J, Aranda MA. Resistance to Cucurbit yellow stunting disorder virus in Cucumber. Plant Dis. 2006;90:583–586. doi: 10.1094/PD-90-0583. [DOI] [PubMed] [Google Scholar]
- 7.Aguilar JM, Franco M, Marco CF, Berdiales B, Rodriguez-Cerezo E, Truniger V, Aranda MA. Further variability within the genus Crinivirus, as revealed by determination of the complete RNA genome sequence of Cucurbit yellow stunting disorder virus. J Gen Virol. 2003;84:2555–2564. doi: 10.1099/vir.0.19209-0. [DOI] [PubMed] [Google Scholar]
- 8.Al Rwahnih M, Dolja VV, Daubert S, Koonin EV, Rowhani A. Genomic and biological analysis of Grapevine leafroll-associated virus 7 reveals a possible new genus within the family Closteroviridae. Virus Res. 2012;163:302–309. doi: 10.1016/j.virusres.2011.10.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Alzhanova DV, Napuli AJ, Creamer R, Dolja VV. Cell-to-cell movement and assembly of a plant closterovirus: roles for the capsid proteins and Hsp70 homolog. EMBO J. 2001;20:6997–7007. doi: 10.1093/emboj/20.24.6997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Bananej K, Menzel W, Kianfar N, Vahdat A, Winter S. First report of Cucurbit chlorotic yellows virus infecting cucumber, melon and squash in Iran. Plant Dis. 2013;97:1005. doi: 10.1094/PDIS-01-13-0125-PDN. [DOI] [PubMed] [Google Scholar]
- 11.Berdiales B, Bernal JJ, Sáez E, Woudt B, Beitia F, Rodríguez-Cerezo E. Occurrence of Cucurbit yellow stunting disorder virus (CYSDV) and Beet pseudo-yellows virus in cucurbit crops in Spain and transmission of CYSDV by two biotypes of Bemisia tabaci. Eur J Plant Pathol. 1999;105:211–215. doi: 10.1023/A:1008713629768. [DOI] [Google Scholar]
- 12.Boubourakas IN, Avgelis AD, Kyriakopoulou PE, Katis NI. Occurrence of yellowing viruses (Beet pseudo-yellows virus, Cucurbit yellow stunting disorder virus and Cucurbit aphid-borne yellows virus) affecting cucurbits in Greece. Plant Pathol. 2006;55:276–283. doi: 10.1111/j.1365-3059.2006.01341.x. [DOI] [Google Scholar]
- 13.Brown JK, Guerrero JC, Matheron M, Olsen M, Idris AM. Widespread outbreak of Cucurbit yellow stunting disorder virus in melon, squash, and watermelon crops in the Sonoran Desert of Arizona and Sonora, Mexico. Plant Dis. 2007;91:773. doi: 10.1094/PDIS-91-6-0773A. [DOI] [PubMed] [Google Scholar]
- 14.Castle S, Palumbo J, Prabhaker N. Newer insecticides for plant virus disease management. Virus Res. 2009;141:131–139. doi: 10.1016/j.virusres.2008.12.006. [DOI] [PubMed] [Google Scholar]
- 15.Célix A, Lopez-Sese AI, Almarza N, Gomez-Guillamon ML, Rodriguez-Cerezo E. Characterization of Cucurbit yellow stunting disorder virus, a Bemisia tabaci-transmitted closterovirus. Phytopathology. 1996;86:1370–1376. [Google Scholar]
- 16.Chen AYS, Walker GP, Carter D, Ng JCK. A virus capsid component mediates virion retention and transmission by its insect vector. Proc Natl Acad Sci USA. 2011;108:16777–16782. doi: 10.1073/pnas.1109384108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Clover GRG, Elliott DR, Tang Z, Alexander BJR. Occurrence of Beet pseudo-yellows virus in cucumber in New Zealand. Plant Pathol. 2002;51:389. doi: 10.1046/j.1365-3059.2002.00702.x. [DOI] [Google Scholar]
- 18.de Ruiter W, Hofstede R, Vries Jd, Heuvel Hvd (2008) Combining QTLs for resistance to CYSDV and powdery mildew in a single cucumber line. In: Pitrat M (ed.) Proceedings of the IXth EUCARPIA meeting on genetics and breeding of Cucurbitaceae, Avignon, pp. 181–188.
- 19.Desbiez C, Lecoq H, Aboulama S, Peterschmitt M. First Report of Cucurbit yellow stunting disorder virus in Morocco. Plant Dis. 2000;84:596. doi: 10.1094/PDIS.2000.84.5.596C. [DOI] [PubMed] [Google Scholar]
- 20.Desbiez C, Lecoq H, Girard M, Cotillon AC, Schoen L. First report of Cucurbit yellow stunting disorder virus in commercial cucumber greenhouses in France. Plant Dis. 2003;87:600. doi: 10.1094/PDIS.2003.87.5.600C. [DOI] [PubMed] [Google Scholar]
- 21.Dolja VV, Kreuze JF, Valkonen JPT. Comparative and functional genomics of closteroviruses. Virus Res. 2006;117:38–51. doi: 10.1016/j.virusres.2006.02.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Duffus JE. Whitefly-transmitted yellowing viruses of the Cucurbitaceae. In: Lester GE, Dunlap JR, editors. Cucurbitaceae 94: evaluation and enhancement of Cucurbit Germplasm. South Padre Island: Gateway Printing; 1995. pp. 12–16. [Google Scholar]
- 23.Duffus JE, Larsen RC, Liu HY. Lettuce infectious yellows virus: a new type of whitefly-transmitted virus. Phytopathology. 1986;76:97–100. doi: 10.1094/Phyto-76-97. [DOI] [Google Scholar]
- 24.Eid S, Abou-Jawdah Y, El-Mohtar C, Sobh H, Havey M. Tolerance in Cucumber to Cucurbit yellow stunting disorder virus. Plant Dis. 2006;90:645–649. doi: 10.1094/PD-90-0645. [DOI] [PubMed] [Google Scholar]
- 25.Gerling D, Alomar Ò, Arnò J. Biological control of Bemisia tabaci using predators and parasitoids. Crop Prot. 2001;20:779–799. doi: 10.1016/S0261-2194(01)00111-9. [DOI] [Google Scholar]
- 26.Gil-Salas FM, Morris J, Colyer A, Budge G, Boonham N, Cuadrado IM, Janssen D. Development of real-time RT-PCR assays for the detection of Cucumber vein yellowing virus (CVYV) and Cucurbit yellow stunting disorder virus (CYSDV) in the whitefly vector Bemisia tabaci. J Virol Methods. 2007;146:45–51. doi: 10.1016/j.jviromet.2007.05.032. [DOI] [PubMed] [Google Scholar]
- 27.Gil-Salas FM, Peters J, Boonham N, Cuadrado IM, Janssen D. Yellowing disease in zucchini squash produced by mixed infections of Cucurbit yellow stunting disorder virus and Cucumber vein yellowing virus. Phytopathology. 2011;101:1365–1372. doi: 10.1094/PHYTO-12-10-0343. [DOI] [PubMed] [Google Scholar]
- 28.Gu QS, Liu YH, Wang YH, Huangfu WG, Gu HF, Xu L, Song FM, Brown JK. First Report of Cucurbit chlorotic yellows virus in Cucumber, Melon, and Watermelon in China. Plant Dis. 2011;95:73. doi: 10.1094/PDIS-07-10-0550. [DOI] [PubMed] [Google Scholar]
- 29.Gyoutoku Y, Okazaki S, Furuta A, Etoh T, Mizobe M, Kuno K, Hayashida S, Okuda M. Chlorotic yellows disease of melon caused by Cucurbit chlorotic yellows virus, a new crinivirus. Jpn J Phytopathol. 2009;75:109–111. doi: 10.3186/jjphytopath.75.109. [DOI] [Google Scholar]
- 30.Hamed K, Menzel W, Dafalla G, Gadelseed AMA, Winter S. First report of Cucurbit chlorotic yellows virus infecting muskmelon and cucumber in Sudan. Plant Dis. 2011;95:1321. doi: 10.1094/PDIS-04-11-0349. [DOI] [PubMed] [Google Scholar]
- 31.Hartono S, Natsuaki T, Genda Y, Okuda S. Nucleotide sequence and genome organization of Cucumber yellows virus, a member of the genus Crinivirus. J Gen Virol. 2003;84:1007–1012. doi: 10.1099/vir.0.18605-0. [DOI] [PubMed] [Google Scholar]
- 32.Hassan AA, Duffus JE. A review of a yellowing and stunting disorder of cucurbits in the United Arab Emirates. Em J Agric Sci. 1991;2:1–16. [Google Scholar]
- 33.Hourani H, Abou-Jawdah Y. Immunodiagnosis of Cucurbit yellow stunting disorder virus using polyclonal antibodies developed against recombinant coat protein. J Plant Pathol. 2003;85:197–204. [Google Scholar]
- 34.Huang LH, Tseng HH, Li JT, Chen TC. First Report of Cucurbit chlorotic yellows virus infecting Cucurbits in Taiwan. Plant Dis. 2010;94:1168. doi: 10.1094/PDIS-94-9-1168B. [DOI] [PubMed] [Google Scholar]
- 35.Janssen D, Ruiz L, Cano M, Belmonte A, Martin G, Segundo E, Cuadrado IM. Physical and genetic control of Bemisia tabaci-transmitted Cucurbit yellow stunting disorder virus in cucumber. Bull OILB/SROP. 2003;26:101–106. [Google Scholar]
- 36.Kao J, Jia L, Tian T, Rubio L, Falk BW. First report of Cucurbit yellow stunting disorder virus (genus Crinivirus) in North America. Plant Dis. 2000;84:101. doi: 10.1094/PDIS.2000.84.1.101C. [DOI] [PubMed] [Google Scholar]
- 37.Karasev AV. Genetic diversity and evolution of closteroviruses. Annu Rev Phytopathol. 2000;38:293–324. doi: 10.1146/annurev.phyto.38.1.293. [DOI] [PubMed] [Google Scholar]
- 38.Kataya ARA, Suliman MNS, Kalantidis K, Livieratos IC. Cucurbit yellow stunting disorder virus p25 is a suppressor of post-transcriptional gene silencing. Virus Res. 2009;145:48–53. doi: 10.1016/j.virusres.2009.06.010. [DOI] [PubMed] [Google Scholar]
- 39.Klaassen VA, Boeshore ML, Koonin EV, Tian T, Falk BW. Genome structure and phylogenetic analysis of Lettuce infectious yellows virus, a whitefly-transmitted, bipartite Closterovirus. Virology. 1995;208:99–110. doi: 10.1006/viro.1995.1133. [DOI] [PubMed] [Google Scholar]
- 40.Kreuze JF, Klein IS, Lazaro MU, Chuquiyuri WJC, Morgan GL, MejÍA PGC, Ghislain M, Valkonen JPT. RNA silencing-mediated resistance to a crinivirus (Closteroviridae) in cultivated sweetpotato (Ipomoea batatas L.) and development of sweetpotato virus disease following co-infection with a potyvirus. Mol Plant Pathol. 2008;9:589–598. doi: 10.1111/j.1364-3703.2008.00480.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Kubota K, Usugi T, Tsuda S. Production of antiserum and immunodetection of Cucurbit chlorotic yellows virus, a novel whitefly-transmitted crinivirus. J Gen Plant Pathol. 2011;77:116–120. doi: 10.1007/s10327-010-0287-5. [DOI] [Google Scholar]
- 42.Kuo YW, Rojas MR, Gilbertson RL, Wintermantel WM. First report of Cucurbit yellow stunting disorder virus in California and Arizona, in association with Cucurbit leaf crumple virus and Squash leaf curl virus. Plant Dis. 2007;91:330. doi: 10.1094/PDIS-91-3-0330B. [DOI] [PubMed] [Google Scholar]
- 43.Lecoq H, Dafalla G, Desbiez C, Wipf-Scheibel C, Kheyr-Pour A. A 10-year survey (1993–2002) of cucurbit viruses in Sudan. J Plant Dis Prot. 2003;110:68–69. [Google Scholar]
- 44.Lecoq H, Desbiez C. Viruses of cucurbit crops in the Mediterranean region: an ever-changing picture. In: Gad L, Hervé L, editors. Advances in virus research. London: Academic Press; 2012. pp. 67–126. [DOI] [PubMed] [Google Scholar]
- 45.Liu HY, Wisler GC, Duffus JE. Particle lengths of whitefly-transmitted Criniviruses. Plant Dis. 2000;84:803–805. doi: 10.1094/PDIS.2000.84.7.803. [DOI] [PubMed] [Google Scholar]
- 46.Liu LZ, Chen YY, Zhu WM. First report of Cucurbit yellow stunting disorder virus on melon in China. Plant Dis. 2010;94:485. doi: 10.1094/PDIS-94-4-0485A. [DOI] [PubMed] [Google Scholar]
- 47.López-Sesé AI, Gómez-Guillamón ML. Resistance to Cucurbit yellowing stunting disorder virus (CYSDV) in Cucumis melo L. HortScience. 2000;35:110–113. [Google Scholar]
- 48.Louro D, Vicente M, Vaira AM, Accotto GP, Nolasco G. Cucurbit yellow stunting disorder virus (genus Crinivirus) associated with the yellowing disease of cucurbit crops in Portugal. Plant Dis. 2000;84:1156. doi: 10.1094/PDIS.2000.84.10.1156A. [DOI] [PubMed] [Google Scholar]
- 49.Luo C, Jones CM, Devine G, Zhang F, Denholm I, Gorman K. Insecticide resistance in Bemisia tabaci biotype Q (Hemiptera: Aleyrodidae) from China. Crop Prot. 2010;29:429–434. doi: 10.1016/j.cropro.2009.10.001. [DOI] [Google Scholar]
- 50.Marco CF, Aguilar JM, Abad J, Gómez-Guillamón ML, Aranda MA. Melon resistance to Cucurbit yellow stunting disorder virus is characterized by reduced virus accumulation. Phytopathology. 2003;93:844–852. doi: 10.1094/PHYTO.2003.93.7.844. [DOI] [PubMed] [Google Scholar]
- 51.Marco CF, Aranda MA. Genetic diversity of a natural population of Cucurbit yellow stunting disorder virus. J Gen Virol. 2005;86:815–822. doi: 10.1099/vir.0.80584-0. [DOI] [PubMed] [Google Scholar]
- 52.Martelli GP, Agranovsky AA, Bar-Joseph M, Boscia D, Candresse T, Coutts RHA, Dolja VV, Hu JS, Jelkmann W, Karasev AV, Martin RR, Minafra A, Namba S, Vetten HJ. Closteroviridae. In: King AMQ, Adams MJ, Carstens EB, Lefkowitz EJ, editors. Virus Taxonomy Ninth Report of the International Committee on Taxonomy of Viruses. London: Elsevier; 2012. pp. 987–1001. [Google Scholar]
- 53.Martin G, Velasco L, Segundo E, Cuadrado IM, Janssen D. The complete nucleotide sequence and genome organization of bean yellow disorder virus, a new member of the genus Crinivirus. Arch Virol. 2008;153:999–1001. doi: 10.1007/s00705-008-0077-y. [DOI] [PubMed] [Google Scholar]
- 54.McCreight JD. Inheritance of Resistance to Lettuce infectious yellows virus in Melon. HortScience. 2000;35:1118–1120. [Google Scholar]
- 55.McCreight JD, Wintermantel WM. Genetic Resistance in Melon PI 313970 to Cucurbit yellow stunting disorder virus. HortScience. 2011;46:1582–1587. [Google Scholar]
- 56.Medina V, Rodrigo G, Tian T, Juarez M, Dolja VV, MAchon MA, Falk BW. Comparative cytopathology of Crinivirus infections in different plant hosts. Ann Appl Biol. 2003;143:99–110. doi: 10.1111/j.1744-7348.2003.tb00274.x. [DOI] [Google Scholar]
- 57.Medina V, Sudarshana MR, Tian T, Ralston KS, Yeh HH, Falk BW. The Lettuce infectious yellows virus (LIYV)-encoded P26 is associated with plasmalemma deposits within LIYV-infected cells. Virology. 2005;333:367–373. doi: 10.1016/j.virol.2005.01.012. [DOI] [PubMed] [Google Scholar]
- 58.Mongkolsiriwattana C, Chen AYS, Ng JCK. Replication of Lettuce chlorosis virus (LCV), a crinivirus in the family Closteroviridae, is accompanied by the production of LCV RNA 1-derived novel RNAs. Virology. 2011;420:89–97. doi: 10.1016/j.virol.2011.08.017. [DOI] [PubMed] [Google Scholar]
- 59.Moury B, Desbiez C, Jacquemond M, Lecoq H. Genetic diversity of plant virus populations: towards hypothesis testing in molecular epidemiology. In: Maramorosch K, Shatkin AJ, Murphy FA, Thresh JM, editors. Advances in virus research. London: Academic Press; 2006. pp. 49–87. [DOI] [PubMed] [Google Scholar]
- 60.Napuli AJ, Alzhanova DV, Doneanu CE, Barofsky DF, Koonin EV, Dolja VV. The 64-kilodalton capsid protein homolog of Beet yellows virus is required for assembly of virion tails. J Virol. 2003;77:2377–2384. doi: 10.1128/JVI.77.4.2377-2384.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Navas-Castillo J, Fiallo-Olivé E, Sánchez-Campos S. Emerging virus diseases transmitted by whiteflies. Annu Rev Phytopathol. 2011;49:219–248. doi: 10.1146/annurev-phyto-072910-095235. [DOI] [PubMed] [Google Scholar]
- 62.Ng JCK, Falk BW. Virus-vector interactions mediating nonpersistent and semipersistent transmission of plant viruses. Annu Rev Phytopathol. 2006;44:183–212. doi: 10.1146/annurev.phyto.44.070505.143325. [DOI] [PubMed] [Google Scholar]
- 63.OEPP/EPPO (2005) Data sheets on quarantine pests. Cucurbit yellow stunting disorder crinivirus. OEPP/EPPO Bull 35:442–444.
- 64.Okuda M, Okazaki S, Yamasaki S, Okuda S, Sugiyama M. Host range and complete genome sequence of Cucurbit chlorotic yellows virus, a new member of the genus Crinivirus. Phytopathology. 2010;100:560–566. doi: 10.1094/PHYTO-100-6-0560. [DOI] [PubMed] [Google Scholar]
- 65.Okuda S, Okuda M, Sugiyama M, Sakata Y, Takeshita M, Iwai H. Resistance in melon to Cucurbit chlorotic yellows virus, a whitefly-transmitted crinivirus. Eur J Plant Pathol. 2013;135:313–321. doi: 10.1007/s10658-012-0088-0. [DOI] [Google Scholar]
- 66.Pacheco CA. Six nouveaux virus des cultures légumières. Phytoma. 2005;580:17–19. [Google Scholar]
- 67.Papayiannis LC, Hunter SC, Iacovides T, Brown JK. Detection of Cucurbit yellow stunting disorder virus in cucurbit leaves using sap extracts and real-time TaqMan® reverse transcription (RT) polymerase chain reaction (PCR) J Phytopathol. 2010;158:487–495. doi: 10.1111/j.1439-0434.2009.01647.x. [DOI] [Google Scholar]
- 68.Papayiannis LC, Ioannou N, Boubourakas IN, Dovas CI, Katis NI, Falk BW. Incidence of viruses infecting cucurbits in Cyprus. J Phytopathol. 2005;153:530–535. doi: 10.1111/j.1439-0434.2005.01015.x. [DOI] [Google Scholar]
- 69.Park SO, Crosby KM, Mirkov TE (2007) Detection of loci for Cucurbit yellow stunting disorder virus resistance in Cucumis melo L. In: Drew R (ed) XXVII International Horticultural Congress, Seoul, pp 207–214.
- 70.Polston JE, Hladky LL, Akad F, Wintermantel WM. First report of Cucurbit yellow stunting disorder virus in cucurbits in Florida. Plant Dis. 2008;92:1251. doi: 10.1094/PDIS-92-8-1251B. [DOI] [PubMed] [Google Scholar]
- 71.Ramirez P, Hernandez E, Mora F, Abraitis R, Hammond R. Limited geographic distribution of beet pseudo-yellows virus Costa Rican cucurbits. J Plant Pathol. 2008;90:331–335. [Google Scholar]
- 72.Rubio L, Abou-Jawdah Y, Lin H-X, Falk BW. Geographically distant isolates of the crinivirus Cucurbit yellow stunting disorder virus show very low genetic diversity in the coat protein gene. J Gen Virol. 2001;82:929–933. doi: 10.1099/0022-1317-82-4-929. [DOI] [PubMed] [Google Scholar]
- 73.Rubio L, Soong J, Kao J, Falk BW. Geographic distribution and molecular variation of isolates of three whitefly-borne Closteroviruses of cucurbits: lettuce infectious yellows virus, Cucurbit yellow stunting disorder virus, and Beet pseudo-yellows virus. Phytopathology. 1999;89:707–711. doi: 10.1094/PHYTO.1999.89.8.707. [DOI] [PubMed] [Google Scholar]
- 74.Rubio L, Yeh H–H, Tian T, Falk BW. A heterogeneous population of defective RNAs is associated with lettuce infectious yellows virus. Virology. 2000;271:205–212. doi: 10.1006/viro.2000.0318. [DOI] [PubMed] [Google Scholar]
- 75.Salem NM, Chen AYS, Tzanetakis IE, Mongkolsiriwattana C, Ng JCK. Further complexity of the genus Crinivirus revealed by the complete genome sequence of Lettuce chlorosis virus (LCV) and the similar temporal accumulation of LCV genomic RNAs 1 and 2. Virology. 2009;390:45–55. doi: 10.1016/j.virol.2009.04.025. [DOI] [PubMed] [Google Scholar]
- 76.Simmons AM, Kousik CS, Levi A. Combining reflective mulch and host plant resistance for sweetpotato whitefly (Hemiptera: Aleyrodidae) management in watermelon. Crop Prot. 2010;29:898–902. doi: 10.1016/j.cropro.2010.04.003. [DOI] [Google Scholar]
- 77.Steel E, Barker I, Danks C, Coates D, Boonham N. A tumefaciens-mediated transient expression as a tool for antigen production for cucurbit yellow stunting disorder virus. J Virol Methods. 2010;163:222–228. doi: 10.1016/j.jviromet.2009.09.024. [DOI] [PubMed] [Google Scholar]
- 78.Stewart LR, Medina V, Sudarshana MR, Falk BW. Lettuce infectious yellows virus-encoded P26 induces plasmalemma deposit cytopathology. Virology. 2009;388:212–220. doi: 10.1016/j.virol.2009.03.016. [DOI] [PubMed] [Google Scholar]
- 79.Stewart LR, Medina V, Tian T, Turina M, Falk BW, Ng JCK. A mutation in the Lettuce infectious yellows virus minor coat protein disrupts whitefly transmission but not in planta systemic movement. J Virol. 2010;84:12165–12173. doi: 10.1128/JVI.01192-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Sweiss M, Anfoka G, Abou-Jawdah Y. Molecular characterization of Jordanian isolates of Cucurbit yellow stunting disorder virus. J Phytopathol. 2007;155:557–562. doi: 10.1111/j.1439-0434.2007.01280.x. [DOI] [Google Scholar]
- 81.Tian T, Rubio L, Yeh HH, Crawford B, Falk BW. Lettuce infectious yellows virus: in vitro acquisition analysis using partially purified virions and the whitefly Bemisia tabaci. J Gen Virol. 1999;80:1111–1117. doi: 10.1099/0022-1317-80-5-1111. [DOI] [PubMed] [Google Scholar]
- 82.Tzanetakis IE, Halgren AB, Keller KE, Hokanson SC, Maas JL, McCarthy PL, Martin RR. Identification and detection of a virus associated with strawberry pallidosis disease. Plant Dis. 2004;88:383–390. doi: 10.1094/PDIS.2004.88.4.383. [DOI] [PubMed] [Google Scholar]
- 83.Tzanetakis IE, Martin RR. Complete nucleotide sequence of a strawberry isolate of Beet pseudoyellows virus. Virus Genes. 2004;28:239–246. doi: 10.1023/B:VIRU.0000025771.48128.f8. [DOI] [PubMed] [Google Scholar]
- 84.Tzanetakis IE, Wintermantel WM, Cortez AA, Barnes JE, Barrett SM, Bolda MP, Martin RR. Epidemiology of Strawberry pallidosis-associated virus and Occurrence of Pallidosis Disease in North America. Plant Dis. 2006;90:1343–1346. doi: 10.1094/PD-90-1343. [DOI] [PubMed] [Google Scholar]
- 85.Tzanetakis IE, Wintermantel WM, Martin RR. First report of Beet pseudo yellows virus in strawberry in the United States: a second crinivirus able to cause pallidosis disease. Plant Dis. 2003;87:1398. doi: 10.1094/PDIS.2003.87.11.1398C. [DOI] [PubMed] [Google Scholar]
- 86.Tzanetakis IE, Wintermantel WM, Poudel B, Zhou J. Diodia vein chlorosis virus is a group-1 crinivirus. Arch Virol. 2011;156:2033–2037. doi: 10.1007/s00705-011-1055-3. [DOI] [PubMed] [Google Scholar]
- 87.Wang J, Stewart LR, Kiss Z, Falk BW. Lettuce infectious yellows virus (LIYV) RNA 1-encoded P34 is an RNA-binding protein and exhibits perinuclear localization. Virology. 2010;403:67–77. doi: 10.1016/j.virol.2010.04.006. [DOI] [PubMed] [Google Scholar]
- 88.Wang J, Yeh H-H, Falk BW. cis preferential replication of Lettuce infectious yellows virus (LIYV) RNA 1: the initial step in the asynchronous replication of the LIYV genomic RNAs. Virology. 2009;386:217–223. doi: 10.1016/j.virol.2009.01.004. [DOI] [PubMed] [Google Scholar]
- 89.Wintermantel WM. Pumpkin (Cucurbita maxima and C. pepo), a new host of Beet pseudo yellows virus in California. Plant Dis. 2004;88:82. doi: 10.1094/PDIS.2004.88.1.82C. [DOI] [PubMed] [Google Scholar]
- 90.Wintermantel WM, Hladky LL. Methods for detection and differentiation of existing and new crinivirus species through multiplex and degenerate primer RT-PCR. J Virol Methods. 2010;170:106–114. doi: 10.1016/j.jviromet.2010.09.008. [DOI] [PubMed] [Google Scholar]
- 91.Wintermantel WM, Hladky LL, Cortez AA, Natwick ET. A new expanded host range of Cucurbit yellow stunting disorder virus includes three agricultural crops. Plant Dis. 2009;93:685–690. doi: 10.1094/PDIS-93-7-0685. [DOI] [PubMed] [Google Scholar]
- 92.Wintermantel WM, Hladky LL, Gulati-Sakhuja A, Li R, Liu HY, Tzanetakis IE. The complete nucleotide sequence and genome organization of tomato infectious chlorosis virus: a distinct crinivirus most closely related to lettuce infectious yellows virus. Arch Virol. 2009;154:1335–1341. doi: 10.1007/s00705-009-0432-7. [DOI] [PubMed] [Google Scholar]
- 93.Wisler GC, Duffus JE. Transmission properties of whitefly-borne Criniviruses and their impact on virus epidemiology. In: Duffus JE, Harris KF, Smith OP, editors. Virus–insect–plant interactions. San Diego: Academic Press; 2001. p. 297. [Google Scholar]
- 94.Wisler GC, Duffus JE, Liu HY, Li RH. Ecology and epidemiology of whitefly-transmitted Closteroviruses. Plant Dis. 1998;82:270–280. doi: 10.1094/PDIS.1998.82.3.270. [DOI] [PubMed] [Google Scholar]
- 95.Yakoubi S, Desbiez C, Fakhfakh H, Wipf-Scheibel C, Marrakchi M, Lecoq H. Occurence of Cucurbit yellow stunting disorder virus and Cucumber vein yellowing virus in Tunisia. J Plant Pathol. 2007;89:417–420. [Google Scholar]
- 96.Yeh H-H, Tian T, Medina V, Falk BW. Green fluorescent protein expression from recombinant Lettuce infectious yellows virus-defective RNAs originating from RNA 2. Virology. 2001;289:54–62. doi: 10.1006/viro.2001.1110. [DOI] [PubMed] [Google Scholar]
- 97.Yeh H-H, Tian T, Rubio L, Crawford B, Falk BW. Asynchronous accumulation of Lettuce infectious yellows virus RNAs 1 and 2 and identification of an RNA 1 trans enhancer of RNA 2 accumulation. J Virol. 2000;74:5762–5768. doi: 10.1128/JVI.74.13.5762-5768.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Zeng R, Dai FM, Chen WJ, Lu JP. First report of Cucurbit chlorotic yellows virus infecting melon in China. Plant Dis. 2011;95:354. doi: 10.1094/PDIS-08-10-0613. [DOI] [PubMed] [Google Scholar]