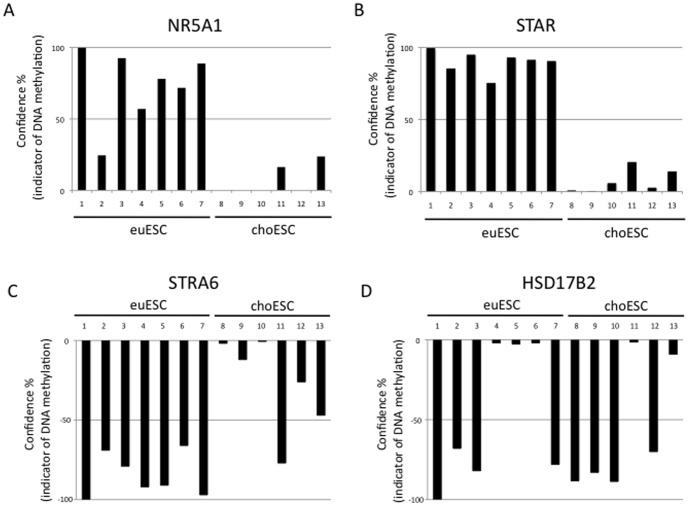
Figure 5. The DNA methylation status as determined by the methylation-sensitive high resolution analyses (MS-HRMA) of NR5A1 (A), STAR (B), STRA6 (C) and HSD17B2 (D) in euESCa and choESC.
Sample No.-wide DNA methylation and transpciptome as euESCa, and sample No. 8, 9, 10 were also analyzed as choESC. Sample No. 1 and 10 were analyzed for bisulfite sequencing as euESCa and choESC, respectively. The confidence value calculated by MS-HRMA indirectly indicates the DNA methylation level. The DNA methylation status of euESCa-1 was shown as 100% identical with regard to the DNA methylation. The confidence values of each sample were calculated in comparison with euESCa-1. In NR5A1 and STAR, the confidence values were lower in choESC than those in euESCa, indicating that the choESC are hypomethylated compared with the euESCa. In STRA6 and HSD17B2, the DNA methylation status of euESCa-1 was shown as −100% (arbitrary defined reverse axis value) identical regarding DNA methylation, indicating a DNA hypomethylation status. In STRA6, the confidence values were higher in choESC than those in euESCa, indicating that the choESC are hypermethylated compared with the euESCa. In HSD17B2, the DNA methylation status varied among individuals with euESCa and choESC. The samples of euESCa and choESC were isolated from seven and six patients, respectively.