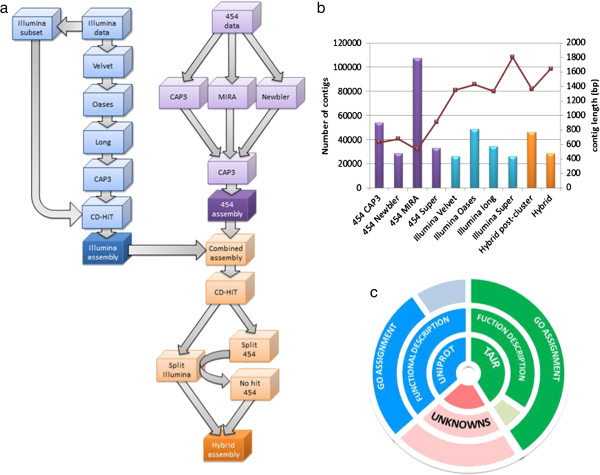
Figure 2.
Transcriptome reconstruction pipeline and results for assembly process at each stage and annotation metrics. a) Flow diagram of de novo reconstruction using 454 pyrosequencing and Illumina data. Illumina data is shown in blue and 454 in purple, the combined data (hybrid assembly) is shown in orange. b) Summary statistics of each stage of de novo transcriptome reconstruction in R.stricta, uing 454 and Illumina sequencing data. Bars show the number of contigs created with the scale on the primary (left) axis and lines show mean contig length on the secondary (right) axis. The x-axis shows the assembly stages. Names pre-fixed with ‘454’ and ‘Illumina’ designate the source of the data for assembly (purple and blue respectively), ‘Hybrid’ means a combination of 454 and Illumina (orange). The remaining x-axis names show the software/stage used, except ‘Super’ which shows merging of the data. C, annotation metrics of R.stricta de novo transcriptome reconstruction, the layered pie chart shows the source of the annotion; TAIR, green; Uniprot, blue; and unannotated (unknowns) pink. The inner layer shows the source of transcripts annotated, the middle layer shows the proportion annotated with a functional description in solid colour and without in faded colour. The outer layer shows the proportion assigned at least one GO term in solid colour and without in faded colour. For unknowns the inner layer is solid colour to show unannotated and the middle and outer are by default faded as they contain no GO or description.