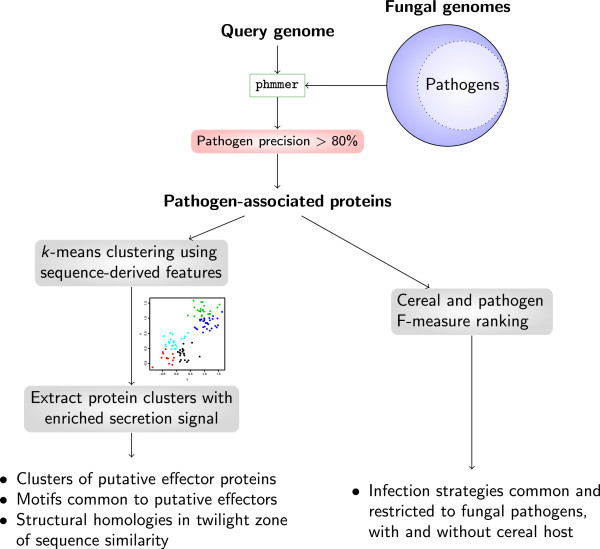
Figure 1.
The comparative pipeline for predicting candidate virulence genes and effectors. Our comparative analysis pipeline predicts proteins in a query pathogen genome which have HMM sequence similarity hits predominantly in fungal pathogens. For a protein to be associated with pathogenicity, the list of phmmer hits must include at least 80% hits to proteins from pathogen species (including the query genome itself). These pathogen-associated proteins are then analyzed in terms of two criteria: (1) their degree of conservation across other fungal pathogens with and without a cereal host and (2) their potential to act as effectors outside the fungal cell. Proteins which are highly conserved across a diverse range of other pathogens as identified by our F-measure ranking are prime candidates for virulence-related proteins. To identify putative effector candidates in an unbiased way, an unsupervised clustering technique based on 35 sequence-derived protein features is used to look for protein clusters with an enriched secretion signal. Further investigation of the clusters with regards to predicted sequence motifs, novel secretion signals, amino acid composition and functional annotation is conducted to find and characterise novel putative effector families.