Figure 1. In-ice selection for polymerase ribozyme activity.
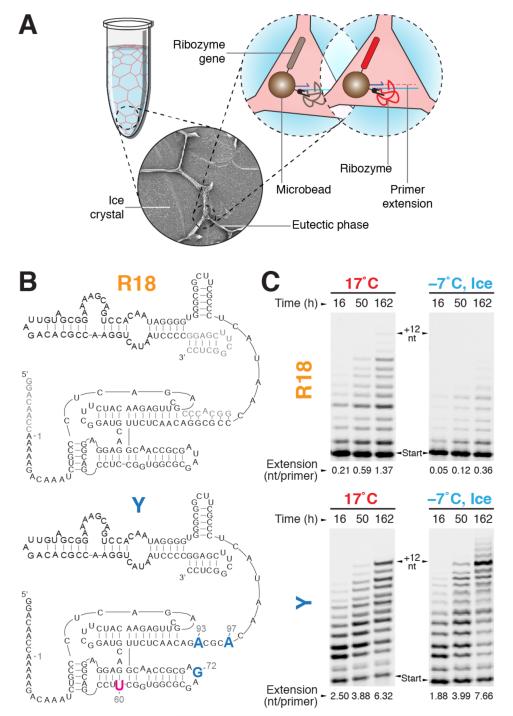
a, Short scheme of in-ice compartmentalised bead-tagging (CBT) selection (full scheme in Supplementary Fig. S1a). Solutes and microbeads are concentrated into the channels of the liquid eutectic phase (pale red) surrounding ice crystals (pale blue). Inset: scanning electron micrograph of ice selection (diameter, 0.15 mm). Ice-active ribozymes (red) extend primers linked to the same bead as themselves and their encoding gene, enabling recovery by flow cytometry. b, Secondary structures of the wild-type R18 ribozyme construct (residues not mutagenised in the starting library are shown in grey) and ribozyme Y, with mutations derived from the in-ice selected ribozyme C8 in blue. c, Polymerase activities of R18 and the evolved ribozyme Y (denaturing PAGE of extension time-courses (primer A /template I)) at 17°C and in ice at −7°C. The average number of nucleotides added per primer is indicated below each lane.