Table 1. Data-collection and refinement statistics.
Values in parentheses are for the outer resolution shell.
| Data-processing statistics | |
| Wavelength (Å) | 1.0332 |
| Space group | P1 |
| Unit-cell parameters (Å, °) | a = 48.79, b = 63.84, c = 163.89, α = 100.67, β = 90.13, γ = 103.49 |
| No. of molecules in asymmetric unit | 8 |
| Solvent content (%) | 57 |
| Resolution range (Å) | 80.4–3.20 (3.37–3.20) |
| No. of observations | 114089 |
| No. of unique reflections | 30145 |
| Average multiplicity | 3.8 (3.9) |
| Completeness (%) | 96.8 (98.4) |
| Wilson B factor (Å2) | 61.5 |
| 〈I/σ(I)〉 | 3.4 (1.30) |
| R p.i.m. † | 0.201 (0.547) |
| Refinement statistics | |
| Resolution range (Å) | 80.4–3.20 |
| Total No. of reflections | 30144 |
| No. of working reflections | 28621 |
| No. of test reflections | 1523 |
| R xpct ‡ | 0.233 |
| R free § | 0.266 |
| No. of atoms | |
| Total | 11158 |
| Protein | 10914 |
| Carbohydrate | 244 |
| R.m.s. bond-length deviation (Å) | 0.007 |
| R.m.s. bond-angle deviation (°) | 0.90 |
| Mean B factor (Å2) | |
| Overall | 79.3 |
| Main chain | 74.8 |
| Side chain | 83.8 |
| Carbohydrate | 99.5 |
| R.m.s. backbone B-factor deviation¶ | 2.2 |
| Ramachandran statistics†† (%) | |
| Favoured | 95.4 |
| Allowed | 99.8 |
| Outliers | 0.2 |
R
p.i.m. (the precision-indicating merging R factor) = 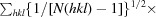
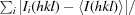
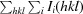 (Weiss, 2001 ▶).
(Weiss, 2001 ▶).
R
xpct = 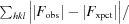
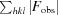 , where |F
obs| and |F
xpct| are the observed structure-factor amplitude and the expectation of the model structure-factor amplitude, respectively (Blanc et al., 2004 ▶).
, where |F
obs| and |F
xpct| are the observed structure-factor amplitude and the expectation of the model structure-factor amplitude, respectively (Blanc et al., 2004 ▶).
R free equals the R xpct of the test set (5% of the data that were removed prior to refinement).
R.m.s. deviation between B factors for bonded main-chain atoms.
As defined by MolProbity (Chen et al., 2010 ▶).
