Table 1. Data-collection and processing statistics for B. megaterium porphobilinogen deaminase.
Values in parentheses are for the outer resolution shell.
| Data set | Crystal 1 | Crystal 2 |
|---|---|---|
| Beamline | I03, DLS | BM14, ESRF |
| Wavelength (Å) | 0.976 | 1.072 |
| Space group | P212121 | P212121 |
| Unit-cell parameters (Å) | ||
| a | 53.32 | 53.01 |
| b | 65.78 | 65.12 |
| c | 97.21 | 96.78 |
| Mosaic spread (°) | 0.26 | 1.01 |
| Resolution (Å) | 48.60–1.46 (1.53–1.46) | 31.33–1.60 (1.69–1.60) |
| R merge † (%) | 6.1 (55.9) | 8.8 (149.3) |
| CC1/2 ‡ (%) | 99.8 (86.7) | 99.8 (33.1) |
| R meas § (%) | 6.7 (61.3) | 9.5 (168.4) |
| Completeness (%) | 100.0 (100.0) | 95.6 (88.7) |
| Average I/σ(I) | 14.4 (3.0) | 10.1 (0.9) |
| Multiplicity | 6.2 (6.0) | 6.4 (4.5) |
| No. of observed reflections | 378329 (52575) | 273058 (25347) |
| No. of unique reflections | 60772 (8748) | 42866 (5649) |
| Wilson plot B factor (Å2) | 15.8 | 17.0 |
| R factor (%) | 14.1 | 16.1 |
| Free R factor (%) | 18.6 | 24.2 |
| R.m.s.d., bond lengths (Å) | 0.024 | 0.019 |
| R.m.s.d., bond angles (°) | 2.44 | 2.14 |
| No. of reflections in working set | 57626 | 40552 |
| Mean protein B factor (Å2) | 24.6 | 28.2 |
R
merge = 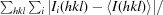
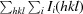 .
.
CC1/2 is the half-set correlation coefficient as described by Karplus & Diederichs (2012 ▶).
R
meas = 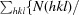
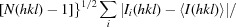
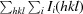 , where 〈I(hkl)〉 is the mean intensity of the N(hkl) observations Ii(hkl) of each unique reflection hkl after scaling.
, where 〈I(hkl)〉 is the mean intensity of the N(hkl) observations Ii(hkl) of each unique reflection hkl after scaling.
