Abstract
Chloride ions are essential for proper function of the photosynthetic oxygen-evolving complex (OEC) of Photosystem II (PS II). Although proposed to be directly ligated to the Mn cluster of the OEC, the specific structural and mechanistic roles of chloride remain unresolved. This study utilizes X-ray absorption spectroscopy (XAS) to characterize the Mn–Cl interaction in inorganic compounds that contain structural motifs similar to those proposed for the OEC. Three sets of model compounds are examined; they possess core structures MnIV3O4X (X = Cl, F, or OH) that contain a di-μ-oxo and two mono-μ-oxo bridges or MnIV2O2X (X = Cl, F, OH, OAc) that contain a di-μ-oxo bridge. Each set of compounds is examined for changes in the XAS spectra that are attributable to the replacement of a terminal OH or F ligand, or bridging OAc ligand, by a terminal Cl ligand. The X-ray absorption near edge structure (XANES) shows changes in the spectra on replacement of OH, OAc, or F by Cl ligands that are indicative of the overall charge of the metal atom and are consistent with the electronegativity of the ligand atom. Fourier transforms (FTs) of the extended X-ray absorption fine structure (EXAFS) spectra reveal a feature that is present only in compounds where chloride is directly ligated to Mn. These FT features were simulated using various calculated Mn–X interactions (X = O, N, Cl, F), and the best fits were found when a Mn–Cl interaction at a 2.2–2.3 Å bond distance was included. There are very few high-valent Mn halide complexes that have been synthesized, and it is important to make such a comparative study of the XANES and EXAFS spectra because they have the potential for providing information about the possible presence or absence of halide ligation to the Mn cluster in PS II.
Keywords: Chloride cofactor, K-edge absorption, Manganese EXAFS, Photosystem II
Introduction
The oxygen-evolving complex (OEC) of Photosystem II (PS II) is a tetranuclear manganese cluster that catalyzes the oxidation of water to dioxygen in a multistep process that requires Ca and Cl for proper function of the catalytic center. The OEC is driven by light to cycle through five different stages, called S-states (S0–S4), collecting four oxidizing equivalents before releasing dioxygen [1]. Recent X-ray diffraction measurements have led to a structure of the PS II complex from thermophilic cyanobacteria [2, 3], but at the present resolution it is not possible to discern the presence or absence of Cl ligation to Mn in the OEC. X-ray absorption spectroscopy (XAS) is a useful tool to directly probe the metal cluster within the PS II membrane protein and has provided a basis for a working structural model of the OEC [4, 5]. As has been the case with other protein-bound metal centers, the success of gaining structural parameters from the extended X-ray absorption fine structure (EXAFS) spectra has motivated the synthesis of structurally relevant model complexes. XAS studies of Mn inorganic compounds with known structures that mimic motifs of the proposed OEC structure have already proved beneficial in interpreting the PS II data (for reviews see [6, 7]). However, few studies have addressed the possibility of whether a Mn–halide interaction can be detected by XAS in multinuclear Mn complexes; thus the set of compounds in this study are chosen to clarify this specific aspect of the structure of the Mn OEC.
It has been known for some time that chloride is an essential cofactor in the proper function of the OEC, but its specific structural and mechanistic roles remain controversial [8, 9, 10]. There is one functional Cl− per PS II unit [11], and it is unclear whether it is directly ligated to Mn in any of the five S-states of the oxidation cycle [12]. Studies of the OEC steady-state kinetics with functional substitutions using Br− or NO3− indicate that Cl− is closely associated with the OEC core rather than with the protein matrix and is necessary for the S2 → S3 and S3 → S0 transitions of the water oxidation cycle [10, 11, 13, 14, 15, 16].
The Mn XAS of three sets of inorganic model compounds in this study document the spectroscopic effect of direct chloride ligation to Mn. All of the compounds have manganese atoms in the +4 oxidation state and, as shown in Fig. 1, have di-μ-oxo-bridged core structures (Mn2O2 or Mn3O4); the trinuclear compounds (Mn3O4) also have two mono-μ-oxo bridges. These core structures provide Mn–Mn distances similar to those reported for the OEC Mn cluster: 2.7 and 3.3 Å. The structurally homologous sets of compounds have different Mn ligands: for MnIV3O4X (X = Cl, F, OH) and for MnIV2O2X (X = Cl, F, OH, OAc), where OAc is a bridging ligand and the rest are terminal ligands. By systematically examining each set of compounds, one can gain insight from the replacement of terminal OH and F ligands or bridging OAc ligand by terminal Cl ligands. Mn X-ray absorption near edge structure (XANES) and EXAFS are sensitive to these ligand substitutions and can show whether the changes are significant enough to be applied to systems where the structure is not known so as to detect Mn–Cl ligation. The XANES results show that the trends in K-edge energy are consistent with the electronegativities of the ligands, but do not provide more than a qualitative understanding. The EXAFS results show that replacement of bridging OAc or terminal OH and F by Cl in structurally homologous sets is easily detectable and that this might provide a method for detecting such replacements in systems of unknown structures such as the OEC.
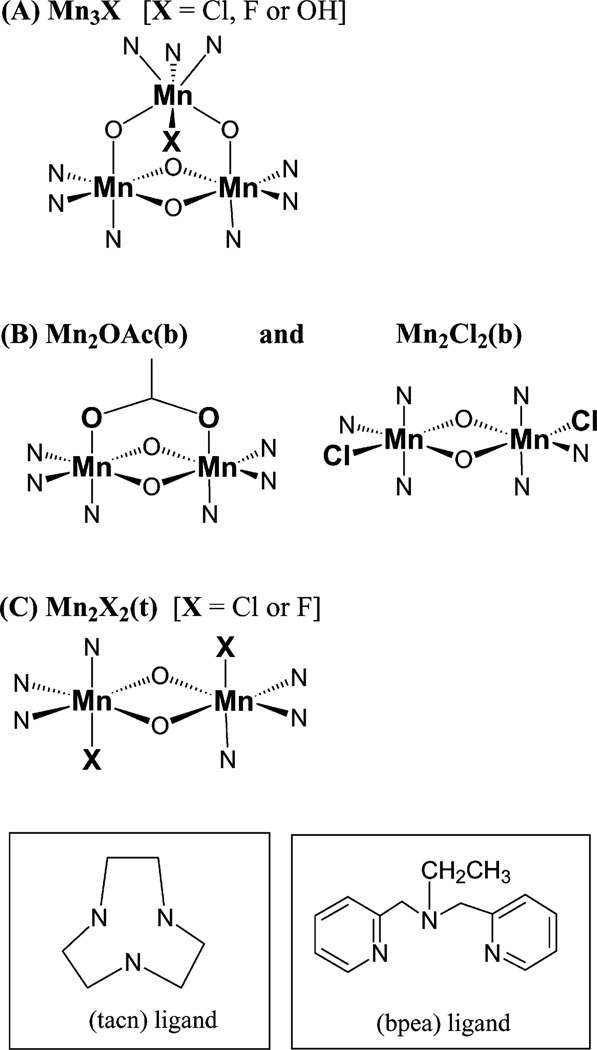
Fig. 1.
Schematic of core structures of trinuclear and binuclear Mn compounds examined: (A) [Mn3O4X(bpea)3](ClO4)3 where X = Cl, F or OH; (B) [Mn2O2Cl2(bpea)2](ClO4)2 and [Mn2O2(O2CCH3) (bpea)2](ClO4)3; and (C) proposed structure for [Mn2O2X2 (tacn)2](BPh4)2, where X = Cl or F. Compounds are labeled according to the abbreviations used in the text. Bridging ligands to Mn are shown in the core structure, while terminal ligands are shown separately in the inset boxes
Materials and methods
Sample preparation
The model compounds used in this study were prepared and characterized in different laboratories ([17, 18, 19], Armstrong WH, Pal S, Mukhopadhyay S, Mok HJ, Wright DW, manuscript in preparation; relevant structural parameters are included in the Tables) {XRD data are unavailable for the compounds described in [17], but the compounds were characterized by elemental analyses (C, H, N, Cl, Mn) and magnetic susceptibility measurements of the MnIV centers}. There is one set of trinuclear Mn compounds: [MnIV3O4X(bpea)3](ClO4)3, where X = Cl, F, or OH; and two sets of binuclear Mn compounds: [MnIV2O2X2(tacn)2](BPh4)2, where X = Cl or F, and [MnIV2O2Cl2(bpea)2](ClO4)2 grouped with [MnIV2O2(O2CCH3)(bpea)2](ClO4)3. All compounds contain Mn in the +4 oxidation state, and a schematic of the core structures is presented in Fig. 1. For convenience in the text discussion, the compounds are labeled according to the number of Mn atoms and the differentiating ligand of interest as follows: Mn3Cl, Mn3F, and Mn3OH for the Mn trinuclear compounds; Mn2Cl2(b), Mn2OAc(b) for the Mn binuclear compounds containing (bpea) ligands, and Mn2Cl2(t), Mn2F2(t) for those containing (tacn) ligands. Samples for XAS measurements were prepared by carefully mixing each compound with boron nitride in a 1:10 ratio and then packing this powder mixture into 0.5-mm thick aluminum sample holders (4 mm×20 mm) sealed with Mylar windows.
X-ray absorption spectroscopy
XAS data were collected at beam line 2–3 of the Stanford Synchrotron Radiation Laboratory, operating at 3.0 GeV with beam currents between 70 and 100 mA. A monochromator outfitted with Si < 220 > crystals, detuned to 50% to reject higher harmonics, was used to provide radiation in the 6.4–7.1 keV energy range. The sample temperature was maintained at 10 ± 2 K with a liquid helium cryostat (Oxford Instruments, Concord, Mass.). Data were collected as fluorescence excitation spectra using a Lytle detector [20, 21] filled with argon gas and placed orthogonal to the incident beam. The fluorescence signal was divided by the incident photon flux detected at the first ionization chamber (filled with N2 gas) to obtain the absorption. Scans were collected from 6520 to 7100 eV, with step sizes of 0.2 eV in the XANES region (6535–6575 eV) and 0.5 Å−1 in the EXAFS region (k = 2–12 Å−1). Energy calibration was achieved by reference to the position of the maximum pre-edge XAS feature from KMnO4 (6543.3 eV, FWHM ≤ 1.7 eV) taken simultaneously. Two to eight scans were averaged for each sample. Refer to [22] for further details of XAS data collection.
XAS data analysis
The method of data preparation for XANES and EXAFS analysis has been previously documented [23, 24, 25] and is briefly summarized here. For each sample, all measured absorption scans were calibrated and then combined to improve the signal-to-noise ratio. The pre-edge background was fit to a straight line and subtracted from the spectra. The data were then divided by the Mn free-atom absorption and normalized to unity at the absorption edge by extrapolating a quadratic line fit from the post-edge region to intersect with the rising edge. The K-edge energy position was deduced from the inflection point energy (IPE) of the main absorption edge as determined from the first zero crossing of its second derivative. Analytical differentiation of the absorption edge utilized a third-order polynomial fit over a 5 eV region (± 2.5 eV from each data point), a range typically used for PS II XAS data.
For analysis of the EXAFS spectrum, an initial background removal was performed by fitting a quadratic or polynomial line to the post-edge region and subtracting it from the spectra. As described in Cinco et al. [23], the resulting spectra were then converted into a photoelectron wave vector “k-space” and weighted by k3. A spline function was subtracted prior to performing a Fourier transform (FT). The FT contains peaks appearing at apparent distances R′, representing scatterers at an average distance R from Mn. Owing to the phase shift, the apparent distance R′ is generally less than the actual distance R by 0.2–0.5 Å.
The FT peaks at R′ > 1 Å were isolated both individually (when a clear separation is possible) or in groups, and were back-transformed for fitting to single-scattering amplitude and phase functions calculated using the program FEFF [26, 27]. The crystal structure coordinates of a representative compound from each core set of compounds were used to calculate the FEFF functions used in the least-squares curve fitting of the isolated FT peaks. The fit extracts four parameters: R, N, σ2 and E0, where N represents the number of scatterers (per Mn) at distance R and σ2 is the Debye–Waller factor. No firm theoretical basis exists to guide the choice of E0; therefore it was also treated as a variable parameter and was constrained to be equal for all shells in the fit to reduce the number of free parameters in the fits. If a compound had available structural data from XRD measurements, the known coordination numbers were used as fixed parameters (N) in the EXAFS fits while all other parameters were allowed to vary. For substantiation of this approach, we also allowed all parameters to vary within physically reasonable upper and lower limits, and these results are included in a section of Supplementary Tables (available online). The quality of the fit was inspected visually for its overall “envelope” shape and was judged quantitatively by the sum of residuals between the simulation and the experimental data. This normalized error Φ is calculated from the number of data points N, the experimental and calculated EXAFS data χ(ki), and a normalization factor si as follows:
| (1) |
| (2) |
Results
Mn XANES
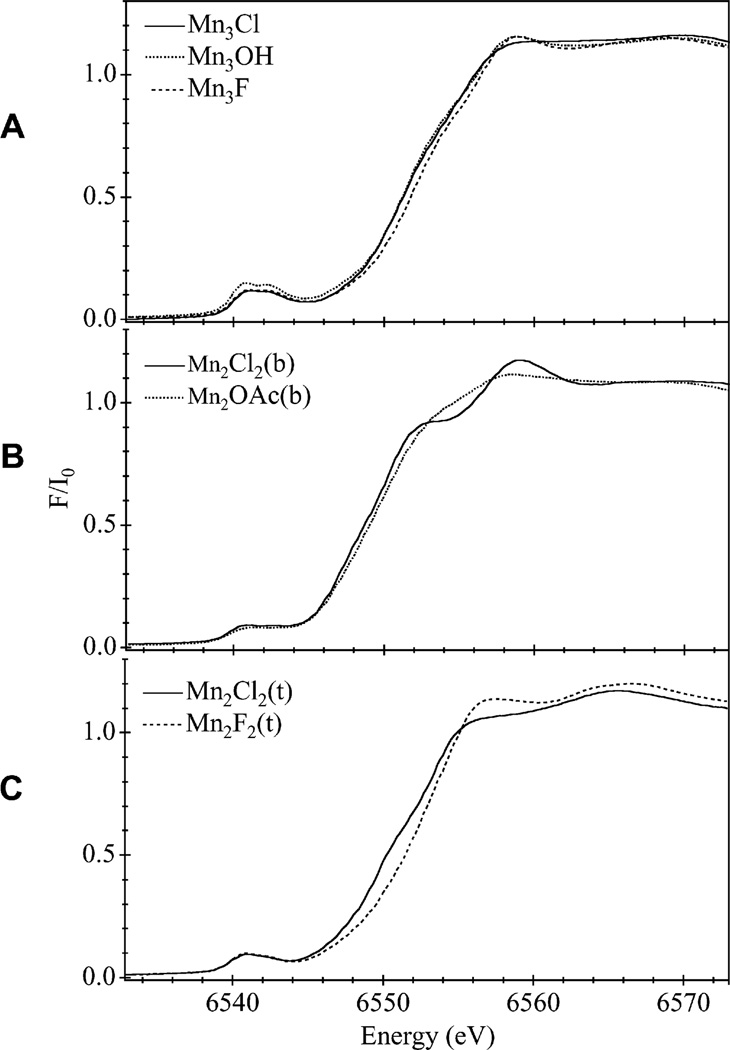
The Mn K-edge absorption spectra of the three sets of compounds are presented in Fig. 2, and their respective second derivatives are shown in Fig. 3. The trinuclear Mn compounds (Fig. 2A) exhibit a higher intensity in the pre-edge regions (~6540 eV) than the binuclear Mn compounds. This forbidden transition [1s → 3d] is noticeably more intense for the Mn3OH compound than for the other trinuclear Mn compounds. A distortion in the core structure of this compound (observed in the XRD) [19] and the reduction in the symmetry of the molecule presumably enhance the “allowedness” of the otherwise forbidden transition. Relative differences in the XANES edge region are better observed in the second derivative of the absorption band; Fig. 3A shows that all three trinuclear compounds have remarkably similar band shapes, albeit different “zero-crossing” positions. The IPE or Mn K-edge energy of the three compounds appears to correlate with the electronegativity of the differing Cl, F, and O ligands. The more electronegative the ligand (δF > δO > δCl), the higher the energy necessary for Mn absorption.
Fig. 2.
Normalized Mn K-edge XANES spectra
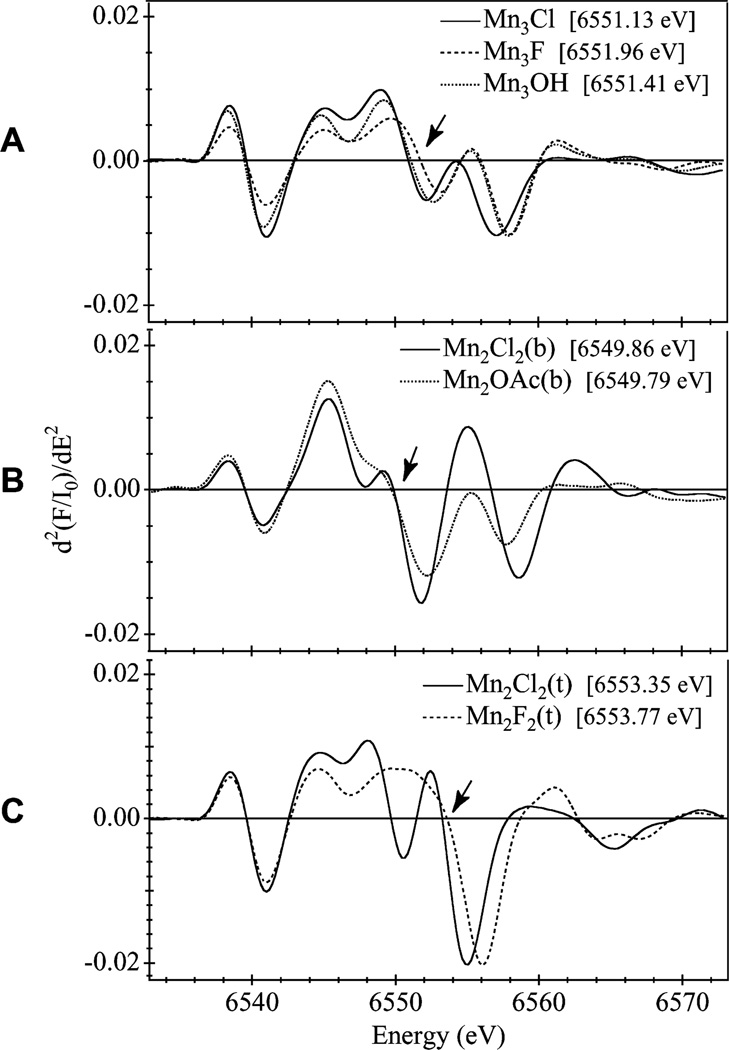
Fig. 3.
Second derivatives of the XANES region shown in Fig. 2 calculated using a 5 eV differentiation interval. The IPE listed for each compound was determined from the zero-crossing position of the main absorption feature, as indicated by arrows
The set of four binuclear Mn compounds share the same Mn2O2 core structure and so are grouped according to the terminal ligands (bpea and tacn). The second derivatives of the XANES (Fig. 3B, C) show that within each group there is general similarity in the absorption band shape and K-edge energy. The aromatic bpea terminal ligands influence the overall Mn environment towards lower K-edge energy values. Like the trinuclear Mn compounds, electronegativity plays a role in the Mn edge energy of each binuclear Mn compound. In the tacn pair (Fig. 3C), the fluoride-containing compound Mn2F2(t) has a higher IPE than the less electronegative Mn2Cl2(t), while in the bpea set the difference in IPE values for the chloride and acetate ligands is very small (0.07 eV). Although oxygen is more electronegative than chloride (δO = 3.44, δCl = 3.16), the contrast is not so great as oxygen compared to fluoride (δF = 3.98). Additionally, differences in the symmetry of the Mn2OAc(b) core structure imposed by the rigidity of the acetate bridge may interfere with an unambiguous determination of the electronegative contribution from the Cl and O ligands. Although qualitative correlations can be derived from XANES while comparing complexes of similar core structure, this example demonstrates the difficulty of reducing the complexity of these absorption edges to a single numerical value when there are significant changes in the surrounding structure.
Mn EXAFS
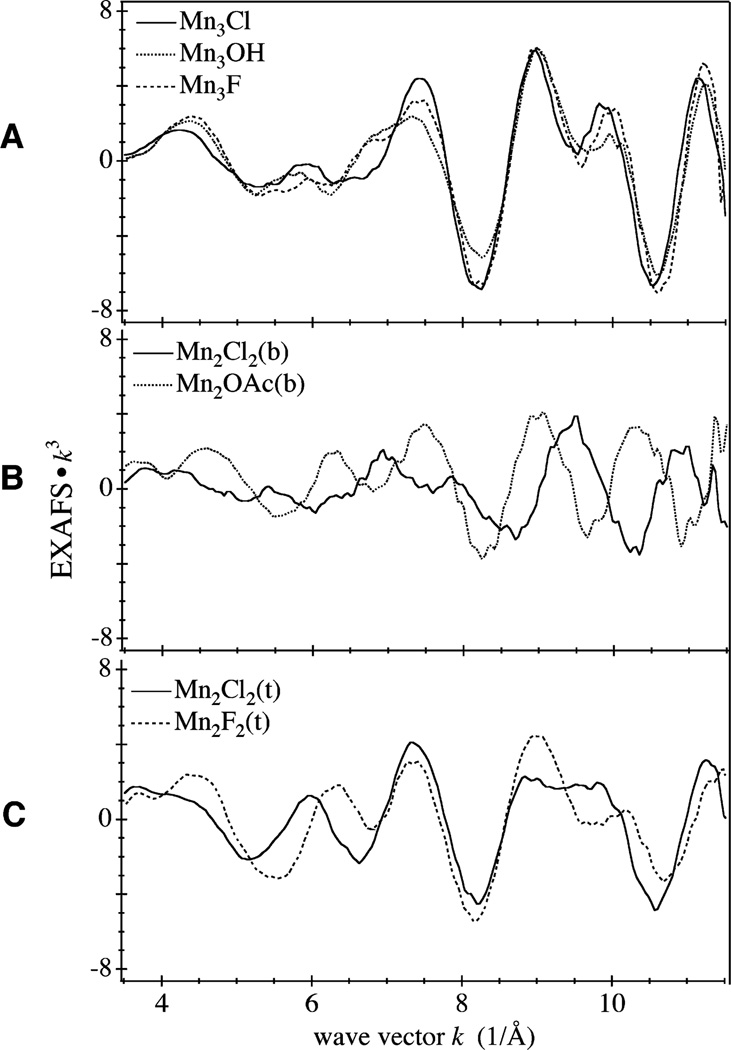
Figure 4 shows the k3-weighted EXAFS spectrum of each compound; the FT of the EXAFS spectrum of each compound is shown in Fig. 5. The k-space spectra for each set of compounds (Fig. 4) show that replacement of OAc, OH, or F by Cl gives rise to significant changes in the spectra, both in the amplitude and the frequency of the EXAFS modulations. The replacement of the bridging OAc by terminal Cl ligands produces the most dramatic change in the k-space spectrum. This is expected because the Mn–Mn distance in these two compounds is noticeably different: 2.76 Å in Mn2Cl2(b) and 2.58 Å in Mn2OAc(b) [18, 19]. In addition, the Mn–Cl distance (2.27 Å) in Mn2Cl2(b) is longer than the average Mn–O distance of the acetate bridge in Mn2OAc(b) (1.94 Å).
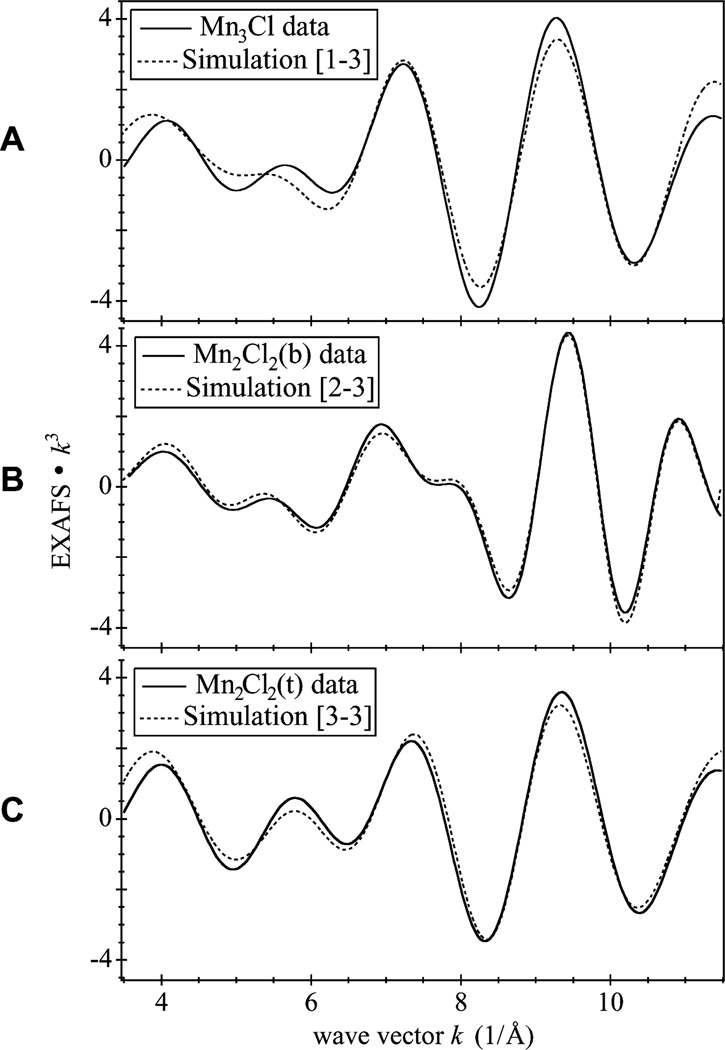
Fig. 4.
Background-subtracted k-space Mn EXAFS weighed by k3
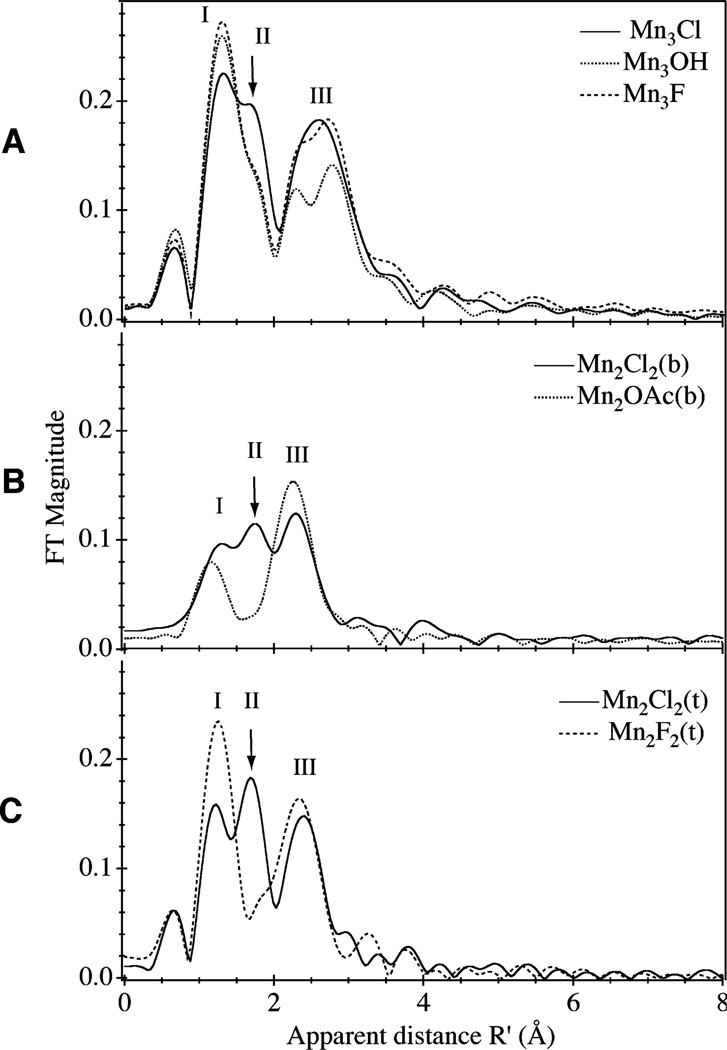
Fig. 5.
Fourier transform spectra of k3-weighted Mn EXAFS shown in Fig. 4
The changes seen in the k-space spectrum are more easily observable in the FTs (Fig. 5) of the EXAFS spectra shown in Fig. 4. The FT peak labeled I (in Fig. 5) contains the backscattering from bridging O, and terminal O and N, ligands at distances of ~1.8–2 Å to the Mn. Fourier peak III contains backscattering from the bridged Mn atoms in these multinuclear complexes. In Fig. 5A, peak III is split into two, indicating the presence of a 2.7 Å Mn–Mn distance from di-μ-oxobridged Mn atoms and also a 3.3 Å Mn–Mn distance from the mono-μ-oxo-bridged Mn atoms. Most importantly from the perspective of this study, the FTs show that only the chloride-containing compounds have a unique feature near apparent distance R′ = 1.7 Å (peak II) that is absent or attenuated in the other compounds. Isolating this Fourier peak II and fitting its back-transformed k-space spectrum (i.e., Fourier filtering) with calculated Mn–X pairs of known identity and bond length reveals the characteristics of the interacting ligand. Because it is not possible to fully separate this FT peak from the neighboring interactions, the R′ = 1.7 Å peak II in each case is filtered together with the strongest neighboring interactions, as a group, for the fitting analysis. The following method was used for the fitting analysis: the coordination numbers (N) were fixed at integral values derived from the structure when it is known and all other parameters were allowed to vary between physically reasonable constraints. For validation of this approach, fits were also carried out where all parameters were allowed to vary. The resulting fits behaved similarly regardless of whether N was fixed and the results from these simulations are included in the Supplementary Tables section. The fits of the chloride-containing compounds are described below.
The FT profile of the Mn3Cl compound (Fig. 5A) shows a shoulder near R′ = 1.7 Å, which was isolated along with the first peak. Table 1 contains the parameters used in simulating the Fourier-filtered EXAFS wave with calculated interaction shells (Mn–O, Mn–N, and Mn–Cl) to represent the nearest atoms to Mn. The graphical results of the best simulations are plotted along with the isolated data in Fig. 6. In each fit, bridging oxygens are grouped apart from the terminal ligands. An initial two-shell simulation of the Mn3Cl compound (Table 1, fit 1-1) is improved by adding a third interaction shell of Mn–O or Mn–Cl with N = 0.3 (Fig. 6). The Mn–Cl interaction greatly improves the two-shell simulation (Δ[Φ] = −36%), whereas a Mn–O interaction shell at a similar distance is only marginally better (Δ[Φ] = −7%). The Mn–Cl distance gained from the fit (2.30 Å) is close to that observed by XRD (2.24 Å).
Table 1.
Parameters obtained from simulations of isolated EXAFS Fourier transform peaks (I, II) of Mn3Cl. Coordination numbers (N) have been fixed according to published XRD data [19, 20] and interatomic distances obtained by XRD are included for comparison. Δ[Φ] shows the relative improvement to the normalized error sum Φ from fit 1-1. The isolated FT data and resulting best fit are shown in Fig. 6
| Fit label | Atomic interaction |
R (Å) | N | σ2(Å2) | Φ×103 | Δ[Φ] | XRDR (Å) |
|---|---|---|---|---|---|---|---|
| 1-1 | Mn–O | 1.79 | 2.7 | 0.0037 | 1.023 | 1.798 | |
| Mn–N | 2.11 | 3.0 | 0.0031 | 2.075 | |||
| 1–2 | Mn–O | 1.79 | 2.7 | 0.0034 | 0.952 | −7% | 1.798 |
| Mn–N | 2.10 | 3.0 | 0.0038 | 2.075 | |||
| Mn–O | 2.15 | 0.3 | 0.0010 | ||||
| 1–3 | Mn–O | 1.79 | 2.7 | 0.0028 | 0.652 | −36% | 1.798 |
| Mn–N | 2.08 | 3.0 | 0.0042 | 2.075 | |||
| Mn–Cl | 2.30 | 0.3 | 0.0010 | 2.242 |
Fig. 6.
Fourier-filtered k-space EXAFS oscillation corresponding to FT peaks I and II of Mn3Cl (shown in Fig. 5A), FT peaks I, II, and III of Mn2Cl2(b) (shown in Fig. 5B), and FT peaks I and II of Mn2Cl2(t) (shown in Fig. 5C). In each case, the isolated data are compared with the simulation that includes a Mn–Cl interaction shell. The atomic interaction shells used and the resulting parameters from each fit are detailed in Tables 1, 2, and 3 respectively
Although not included in this study, parallel fits of the Mn3F and Mn3OH compounds were systematically carried out to document the effect of using a Mn–Cl interaction shell when there is no chloride present in the compound. This exercise showed that the initial two-shell simulation worsens as expected when a Mn–Cl interaction is included in place of a Mn–F or Mn–O at approximately the same distance; however, if the Mn–Cl shell is allowed to reach longer interatomic distances (> 2.2 Å), a better overall fit can be obtained (Supplementary Tables S4 and S5). Although it may be difficult to identify Cl unequivocally from any one complex, it is clear from the comparison of the FTs between the Mn3F, Mn3OH, and Mn3Cl complexes that the FT for the Cl-containing complex is distinct. This is the kind of comparison between Cl-containing and Cl-free PS II that we envision for the future.
The EXAFS simulations of the binuclear Mn compounds lead to similar conclusions to Mn3Cl. Both show strong peaks near R′ = 1.7 Å which can be isolated along with the nearest FT features. In Mn2Cl2(b), the middle peak feature cannot be cleanly separated from the longer distance interactions so the entire three-peak profile has been filtered and analyzed as a group (Table 2, Fig. 6). Because this FT isolate includes all three peaks and expands to R′≈3 Å, longer distance interactions like Mn–C must be included in the simulation. A good fit is already obtained starting with a four-shell simulation (Fit 2-1) representing all core and terminal ligands except for Cl. As in Mn3Cl, this initial fit was improved upon adding another short-range interaction at a fixed coordination N = 1.0. Table 2 shows that the simulation including Mn–Cl (Δ[Φ] = −50%) is a better choice than Mn–O (Δ[Φ] = −36%). The Mn–Cl distance obtained from the fit is 2.21 Å and is comparable to that measured through XRD (2.273 Å). Figure 6B shows the best fit to the data.
Table 2.
Parameters obtained from simulations of isolated EXAFS Fourier transform peaks (I, II, III) of Mn2Cl2(b). Coordination numbers (N) have been fixed according to published XRD data [18] and interatomic distances obtained by XRD are included for comparison. Δ[Φ] shows the relative improvement to the normalized error sum Φ from fit 2-1. The isolated FT data and resulting simulated best fit are shown in Fig. 6
| Fit label | Atomic interaction |
R (Å) | N | σ2(Å2) | Φ×103 | Δ[Φ] | XRDR (Å) |
|---|---|---|---|---|---|---|---|
| 2-1 | Mn–O | 1.79 | 2.0 | 0.0048 | 0.176 | 1.811 | |
| Mn–N | 2.07 | 3.0 | 0.0055 | 2.033 | |||
| Mn–Mn | 2.75 | 1.0 | 0.0029 | 2.756 | |||
| Mn–C | 3.00 | 7.0 | 0.0070 | 2.914 | |||
| 2-2 | Mn–O | 1.78 | 2.0 | 0.0052 | 0.112 | −36% | 1.811 |
| Mn–N | 2.03 | 3.0 | 0.0178 | 2.033 | |||
| Mn–Mn | 2.75 | 1.0 | 0.0026 | 2.756 | |||
| Mn–C | 2.99 | 7.0 | 0.0072 | 2.914 | |||
| Mn–O | 2.08 | 1.0 | 0.0022 | ||||
| 2–3 | Mn–O | 1.78 | 2.0 | 0.0049 | 0.088 | −50% | 1.811 |
| Mn–N | 2.10 | 3.0 | 0.0127 | 2.033 | |||
| Mn–Mn | 2.74 | 1.0 | 0.0027 | 2.756 | |||
| Mn–C | 2.98 | 7.0 | 0.0100 | 2.914 | |||
| Mn–Cl | 2.21 | 1.0 | 0.0090 | 2.273 |
In the case of Mn2Cl2(t), there is no available XRD structure. Assuming from the compositional analysis that it has a core structure similar to Mn2Cl2(b), the coordination values in each fit have been fixed analogously. FT peaks I and II (Fig. 5C) are isolated together and the resulting fit parameters are reported in Table 3, while the best simulation and data are plotted in Fig. 6. The error sum Φ is once again lowest in the simulation that includes a Mn–Cl interaction shell over other likely Mn–O or Mn–N shells. The resulting Mn–Cl bond distance (R = 2.27 Å) is comparable to those of the other two compounds. Relaxing all fit parameters to find the lowest mathematical minimum arrives at the same general result, namely that the Mn–Cl vector is significantly superior to the alternative Mn–O at the same distance (R≈2.2 Å).
Table 3.
Parameters obtained from simulations of isolated EXAFS Fourier transform peaks (I, II) of Mn2Cl2(t). Coordination numbers (N) have been fixed according to its compositional similarity with Mn2Cl2(b). Δ[Φ] shows the relative improvement to the normalized error sum Φ from fit 3-1. The isolated FT data and resulting simulated best fit are shown in Fig. 6
| Fit label | Atomic interaction |
R (Å) | N | σ2(Å2) | Φ×103 | Δ[Φ] |
|---|---|---|---|---|---|---|
| 3-1 | Mn–O | 1.79 | 2.0 | 0.0029 | 1.159 | |
| Mn–N | 2.07 | 3.0 | 0.0038 | |||
| 3-2 | Mn–O | 1.79 | 2.0 | 0.0018 | 0.718 | −38% |
| Mn–N | 2.03 | 3.0 | 0.0055 | |||
| Mn–O | 2.14 | 1.0 | 0.0010 | |||
| 3-3 | Mn–O | 1.77 | 2.0 | 0.0010 | 0.262 | −77% |
| Mn–N | 2.00 | 3.0 | 0.0056 | |||
| Mn–Cl | 2.27 | 1.0 | 0.0035 |
Discussion
Mn XANES
The XANES region is rich with information and has thus far eluded complete theoretical analysis. One of the experimental measures typically reported for K-edge spectra is the ionization energy. Taken as the first zero-crossing observed in the calculated second-derivative function of the main absorption edge, this so-called “edge energy” or inflection point energy (IPE), although widely reported, is by itself an incomplete representation of the K-edge absorption. Although it is a qualitatively adequate practice to compare the IPE values of two related compounds, it is only a starting point; the structure-rich profile of the second-derivative absorption spectrum should also be taken into consideration.
This study addresses the influence of the ligand environment on the observed Mn XAS, especially in relation to chloride ligation. This is an issue that is directly relevant to the proposed water oxidation mechanism of PS II [28, 29] and the unresolved role of the chloride co-factor. Because XANES records a 1s → 4p absorption transition (K-edge), the Mn absorption edge is sensitive to both oxidation state and surrounding ligands [30]. The compounds chosen for this study have manganese in the +4 oxidation state and have been separated into three sets according to the number of Mn atoms and the nature of the terminal ligand group. With the Mn oxidation state kept constant, the effect of the surrounding ligands on the Mn absorption edge can be systematically evaluated.
Although all of the compounds in this study have the same formal Mn4+ oxidation state, the variations seen in the Mn XANES show that the K-edge is sensitive to changes in the effective charge of the Mn induced by the electron-donating character of the ligating atoms [31]. This effect is best observed for two structurally similar Mn compounds with different halide ligands. The greater electronegativity of the F ligand draws electrons away from the Mn4+ core and therefore shifts the K-edge absorption towards a higher energy relative to that for a Cl ligand. The difference in IPE between Mn2Cl2(t) and Mn2F2(t) is 0.85 eV (Fig. 3C), while in the set of trinuclear Mn compounds (Mn3Cl vs. Mn3F) it is 0.5 eV. Although different in magnitude, the similarity in behavior distinctly places the compounds containing fluoride at a higher IPE than those containing chloride ligands. This trend is consistent with previous observations made on tetranuclear Mn cubane compounds containing a single Cl or F ligand directly ligated to three of the four Mn atoms [23]. In that case, the Mn K-edge energy (determined from IPE taken at 6 eV) was 2.8 eV higher for the fluoride-containing cubane relative to the non-halide cubane. The higher electronegativity of fluoride clearly affects the Mn absorption profile in a consistent manner.
In contrast, the comparison between Cl and O ligands is not so clearly ordered according to electronegativity. As noted in the Results section, the structural changes imposed by an acetate bridge in the case of the Mn2OAc(b), and hydrogen bonding in Mn3OH, affect the K-edge energy; thus, the IPE values cannot be directly compared with those of their chloride-containing analogs. Such subtle structural changes in the entire surrounding ligand environment of the Mn have a complex effect on the observed XANES. The differences in the K-edge profiles (accentuated in the second derivative shown in Fig. 3) between the two sets of binuclear Mn compounds show that the terminal ligand environment (tacn vs. bpea) has a significant effect on the Mn XANES despite the similarity in the core structure and halide ligand. Aromatic ligands have unoccupied π* orbitals which can have strong π back-bonding interactions with the Mn d-orbitals. The shape of the absorption K-edge is remarkably different for Mn2Cl2(t) and Mn2Cl2(b), and the edge energy for the binuclear Mn compounds containing the aromatic bpea ligands is generally lower than that of the tacn compounds. All of these structural factors impede a direct comparison of the IPE values between compounds containing chloride in place of oxygen ligands.
Lastly, the XANES region can be affected not only by the covalency but also by the symmetry of the metal center [29, 30]. Less susceptible to changes in the valence environment of the metal, the “forbidden” 1s → 3d pre-edge (XANESMn3OH pre-edge is noticeably higher than that of the other compounds. This change in the symmetry of the metal center is confirmed by XRD measurements [19], which show that there is hydrogen bonding between the OH group and the oxygens bridging the Mn, resulting in a distortion of the core structure.
Further study of model compounds is needed to fully understand the nature of the substructures seen in absorption K-edges and to solidify correlations between differences in chemical structure and observed spectra. Although an indicator of Mn–ligand interactions, IPE values cannot be used in isolation without further consideration of the chemical structure or other environmental factors.
Mn EXAFS
The sets of compounds chosen for this study address the effect of the absence or presence of Cl on the observed Mn XAS and the potential impact for studying the structural role of the chloride cofactor in PS II. Within each set of compounds, the chloride-containing compound is distinguished by a peak in the FT profile (Fig. 5) near apparent distance R′ = 1.7 Å that is absent or greatly decreased in the non-chloride-containing compounds. The XRD measurements of Mn2Cl2(b) and Mn3Cl show that the Mn–Cl bond is longer than the Mn–F and Mn–OH bonds in Mn2F2(t), Mn3F, and Mn3OH ([18, 19], Armstrong WH, Pal S, Mukhopadhyay S, Mok HJ, Wright DW, manuscript in preparation). This longer bond distance separates the Mn–Cl interaction from the other short bond distances (Mn–O and Mn–N), resulting in a distinct shoulder or peak in the FT profile. Fourier filtering of these features and fitting in k-space with simulated phase and amplitude functions of known interatomic interactions and distance allows us to assign these features to Mn–Cl interactions.
Because crystallographic structures are available for Mn3Cl and for Mn2Cl2(b), the results of the XRD measurements are included in Tables 1 and 2 for comparison with the EXAFS fit simulations. The results from the two methods are in good agreement. In the best fit for each compound, the largest difference in bond distances is 0.067 Å. When all parameters are allowed to vary, the bond distances remain consistent and the resulting coordination numbers arrive within 25% of the expected XRD values. This gives us confidence that the resulting EXAFS fit simulations are accurate for the unknown structure of Mn2Cl2(t) and that the observed peak at R′≈1.7 Å in all three chloride-containing compounds is indeed due to a Mn–Cl interaction.
The phase and amplitude functions used in the fitting procedure were calculated using the coordinates for a compound with the same core structure. Therefore, the binuclear Mn compounds were fit using a slightly different set of functions from those used for the trinuclear Mn compounds. This implicitly assumes that we have some idea of the starting point of the structure that guides the iteration procedure to obtain the best fit of the data. Because this is a situation that is not possible for a metalloprotein system like PS II, we attempted fits of the FT peaks near R′≈1.7 Å using simulated functions that were not selected or separated according to core structure. The fits produced using these simple two-atom (Mn–backscatterer) functions were not so satisfactory overall, but the underlying result was the same: fits containing a Mn–Cl interaction (at ~2.3 Å) were superior to those containing a Mn–O interaction at a similar distance. The characteristic shape of the Mn–Cl back-scatter function cannot be easily simulated by that of Mn–F or Mn–O, which are difficult to distinguish from each other; the fit containing Mn–Cl stands out clearly as the best fit in the chloride-containing compounds regardless of the starting assumption of Mn core structure. The comparison of the individual Mn–O, Mn–F, and Mn–Cl backscatter functions with an interatomic distance of 2.2 Å is shown in Supplementary Fig. S1.
Conclusions
This study demonstrates that XAS can be used to detect Mn–Cl ligation in a set of Mn compounds that possess the same Mn oxidation state and similar core structures. The XANES data show that the K-edge energies of structurally analogous compounds with different halides (Cl versus F) show a detectable decrease in the Mn K-edge energy with decreasing ligand electronegativity. However, the comparison between Cl and O is not conclusive, owing to differences in the structures of Mn2OAc(b) and Mn3OH, as compared to the chloride containing compounds. This finding has implications for the interpretation of the OEC XANES data. First, the Cl/Mn atom ratio in the OEC is 1/4 [11], while in these compounds it is 1/1 or 1/3. Thus, the effect of direct chloride ligation among the Mn–O bonds in the OEC may not be easily detected in the Mn K-edge, although chloride has a somewhat lower electronegativity than oxygen. Second, slight changes among the core structures of the model compounds have an effect on the observed XANES. This means that significant changes in the Mn K-edge energy cannot be attributed solely to changes in the Mn oxidation state, but can also be the result of structural rearrangement due to chloride binding or releasing at different stages of the water oxidation cycle. The evaluation of XANES is strengthened by coupling with other techniques like XRD or XRF (X-ray fluorescence) [32].
In the post-edge region, the k-space view of the EXAFS profile in each set of model compounds shows that the amplitude and frequency is affected by the longer Mn–Mn and Mn–Cl bond distances found in the chloride-containing compounds. In fact, it may be this longer Mn–Cl bond that allows detection in the FT profile where a peak or shoulder appears near R′ = 1.7 Å. Simulations of these features fit best to Mn–Cl interactions with a ~2.3 Å bond distance; in the case of Mn3Cl and Mn2Cl2(b), all structural parameters obtained from the EXAFS data are in good agreement with available results from XRD measurements. This validation gives us confidence that interpretation of the EXAFS data from the unknown Mn2Cl2(t) structure is also accurate. Furthermore, EXAFS measurements are able to detect changes in the OEC structure as it cycles through the five S-states of the water oxidation process. Chloride has been shown to be a necessary cofactor during the S2 → S3 transition [10, 11, 13], but it is unknown whether it is directly ligated to the Mn core of the OEC. Polarized Mn EXAFS measurements of PS II samples poised in the S3-state showed considerable dichroism and significant structural changes compared to the S2-state [33]. The FT of the EXAFS spectrum from the S3-state data clearly shows a Fourier peak near R′≈1.9 Å. Although this apparent distance is longer than that found in the chloride-containing model complexes in this study, the isolated k-space profiles are similar. Simulations of this R′≈1.9 Å feature in PS II revealed that the best fit is a Mn–Cl interaction with a 2.2 Å bond distance, suggestive that chloride may be a direct ligand of Mn during the S3-state. The results from the model compounds presented in this paper support that conclusion.
Supplementary Material
Acknowledgements
The authors would like to thank Drs. Johannes Messinger, Carmen Fernandez, and Karen MacFarlane Holman for help with XAS data collection as well as Dr. Junko Yano for help with manuscript preparation. Funding for this work was provided by the National Institutes of Health grant GM-55302 and by the Director, Office of Science, Office of Basic Energy Sciences, Division of Chemical Sciences, Geosciences, and Biosciences, US Department of Energy under contract DE-AC03-76SF00098. Synchrotron radiation facilities were provided by the Stanford Synchrotron Radiation Laboratory (SSRL), which is operated by the US Department of Energy, Office of Basic Energy Sciences. The SSRL Biotechnology program is supported by the National Institutes of Health, National Center of Research Resources, Biomedical Technology Program, and by the Department of Energy, Office of Biological and Environmental Research.
Abbreviations
- bpea
N,N-bis(2-pyridylmethyl) ethylamine
- EXAFS
extended X-ray absorption fine structure
- FT
Fourier transform
- IPE
inflection point energy
- OEC
oxygen evolving complex
- PS II
Photosystem II
- tacn
1,4,7-triazacyclononane
- XANES
X-ray absorption near edge structure
- XAS
X-ray absorption spectroscopy
- XRD
X-ray diffraction
Footnotes
Electronic Supplementary Material Supplementary material is available in the online version of this article at http://dx.doi.org/10.1007/s00775-003-0520-1
Contributor Information
Shelly A. Pizarro, Melvin Calvin Laboratory, Physical Biosciences Division, Lawrence Berkeley National Laboratory, Berkeley, CA 94720-5230, USA Department of Chemistry, University of California, Berkeley, CA 94720-5230, USA.
Hendrik Visser, Melvin Calvin Laboratory, Physical Biosciences Division, Lawrence Berkeley National Laboratory, Berkeley, CA 94720-5230, USA; Department of Chemistry, University of California, Berkeley, CA 94720-5230, USA.
Roehl M. Cinco, Melvin Calvin Laboratory, Physical Biosciences Division, Lawrence Berkeley National Laboratory, Berkeley, CA 94720-5230, USA Department of Chemistry, University of California, Berkeley, CA 94720-5230, USA.
John H. Robblee, Melvin Calvin Laboratory, Physical Biosciences Division, Lawrence Berkeley National Laboratory, Berkeley, CA 94720-5230, USA Department of Chemistry, University of California, Berkeley, CA 94720-5230, USA.
Samudranil Pal, Department of Chemistry, Eugene F. Merkert Chemistry Center, Boston College, Chestnut Hill, MA 02467-3860, USA.
Sumitra Mukhopadhyay, Department of Chemistry, Eugene F. Merkert Chemistry Center, Boston College, Chestnut Hill, MA 02467-3860, USA.
Henry J. Mok, Department of Chemistry, Eugene F. Merkert Chemistry Center, Boston College, Chestnut Hill, MA 02467-3860, USA
Kenneth Sauer, Melvin Calvin Laboratory, Physical Biosciences Division, Lawrence Berkeley National Laboratory, Berkeley, CA 94720-5230, USA; Department of Chemistry, University of California, Berkeley, CA 94720-5230, USA.
Karl Wieghardt, Max Planck Institut für Bioorganische Chemie, 45413 Mülheim an der Ruhr, Germany.
William H. Armstrong, Department of Chemistry, Eugene F. Merkert Chemistry Center, Boston College, Chestnut Hill, MA 02467-3860, USA
Vittal K. Yachandra, Melvin Calvin Laboratory, Physical Biosciences Division, Lawrence Berkeley National Laboratory, Berkeley, CA 94720-5230, USA, vkyachandra@lbl.gov, Tel.: +1-510-4864330, Fax: +1-510-4866059
References
- 1.Kok B, Forbush B, McGloin M. Photochem Photobiol. 1970;11:457–475. doi: 10.1111/j.1751-1097.1970.tb06017.x. [DOI] [PubMed] [Google Scholar]
- 2.Zouni A, Witt H-T, Kern J, Fromme P, Krauß N, Saenger W, Orth P. Nature. 2001;409:739–743. doi: 10.1038/35055589. [DOI] [PubMed] [Google Scholar]
- 3.Kamiya N, Shen JR. Proc Natl Acad Sci USA. 2003;100:98–103. doi: 10.1073/pnas.0135651100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Yachandra VK, DeRose VJ, Latimer MJ, Mukerji I, Sauer K, Klein MP. Science. 1993;260:675–679. doi: 10.1126/science.8480177. [DOI] [PubMed] [Google Scholar]
- 5.Yachandra VK, Sauer K, Klein MP. Chem Rev. 1996;96:2927–2950. doi: 10.1021/cr950052k. [DOI] [PubMed] [Google Scholar]
- 6.Pecoraro VL, Hsieh W-Y. In: Metal ions in biological systems: manganese and its role in biological processes. Sigel A, Sigel H, editors. vol 37. New York: Dekker; 2000. pp. 429–504. [PubMed] [Google Scholar]
- 7.Hage R, Kerschner J. Trends Inorg Chem. 1998;5:145–159. [Google Scholar]
- 8.Homann PH. J Bioenerg Biomembr. 1987;19:105–123. doi: 10.1007/BF00762720. [DOI] [PubMed] [Google Scholar]
- 9.Coleman WJ. Photosynth Res. 1990;23:1–27. doi: 10.1007/BF00030059. [DOI] [PubMed] [Google Scholar]
- 10.Wincencjusz H, Yocum CF, van Gorkom HJ. Biochemistry. 1999;38:3719–3725. doi: 10.1021/bi982295n. [DOI] [PubMed] [Google Scholar]
- 11.Lindberg K, Andréasson L-E. Biochemistry. 1996;35:14259–14267. doi: 10.1021/bi961244s. [DOI] [PubMed] [Google Scholar]
- 12.Olesen K, Andréasson L-E. Biochemistry. 2003;42:2025–2035. doi: 10.1021/bi026175y. [DOI] [PubMed] [Google Scholar]
- 13.Wincencjusz H, van Gorkom HJ, Yocum CF. Biochemistry. 1997;36:3663–3670. doi: 10.1021/bi9626719. [DOI] [PubMed] [Google Scholar]
- 14.Sandusky PO, Yocum CF. Biochim Biophys Acta. 1984;766:603–611. [Google Scholar]
- 15.Yocum CF. Biochim Biophys Acta. 1991;1059:1–15. [Google Scholar]
- 16.Ono T, Nakayama H, Gleiter H, Inoue Y, Kawamori A. Arch Biochem Biophys. 1987;256:618–624. doi: 10.1016/0003-9861(87)90619-9. [DOI] [PubMed] [Google Scholar]
- 17.Wieghardt K, Bossek U, Zsolnai L, Huttner G, Blondin G, Girerd J-J, Babonneau FJ. J Chem Soc Chem Commun. 1987:651–653. [Google Scholar]
- 18.Pal S, Olmstead MM, Armstrong WH. Inorg Chem. 1995;34:4708–4715. [Google Scholar]
- 19.Pal S, Chan MK, Armstrong WH. J Am Chem Soc. 1992;114:6398–6406. [Google Scholar]
- 20.Stern EA, Heald SM. Rev Sci Instrum. 1979;50:1579. doi: 10.1063/1.1135763. [DOI] [PubMed] [Google Scholar]
- 21.Lytle FW, Greegor RB, Sandstrom DR, Marques EC, Wong J, Spiro CL, Huffman GP, Huggins FE. Nucl Instrum Methods. 1984;226:542–548. [Google Scholar]
- 22.Roelofs TA, Liang MC, Latimer MJ, Cinco RM, Rompel A, Andrews JC, Sauer K, Yachandra VK, Klein MP. Proc Natl Acad Sci USA. 1996;93:3335–3340. doi: 10.1073/pnas.93.8.3335. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Cinco RM, Rompel A, Visser H, Aromí G, Christou G, Sauer K, Yachandra VK, Klein MP. Inorg Chem. 1999;38:5988–5998. doi: 10.1021/ic991003j. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Latimer MJ, DeRose VJ, Mukerji I, Yachandra VK, Sauer K, Klein MP. Biochemistry. 1995;34:10898–10909. doi: 10.1021/bi00034a024. [DOI] [PubMed] [Google Scholar]
- 25.DeRose VJ, Mukerji I, Latimer MJ, Yachandra VK, Sauer K, Klein MP. J Am Chem Soc. 1994;116:5239–5249. [Google Scholar]
- 26.Rehr JJ, Mustre de Leon J, Zabinsky SI, Albers RC. J Am Chem Soc. 1991;113:5135–5140. [Google Scholar]
- 27.Rehr JJ, Albers RC, Zabinsky SI. Phys Rev Lett. 1992;69:3397–3400. doi: 10.1103/PhysRevLett.69.3397. [DOI] [PubMed] [Google Scholar]
- 28.Messinger J, Robblee JH, Bergmann U, Fernandez C, Glatzel P, Visser H, Cinco RM, McFarlane KL, Bellacchio E, Pizarro S, Cramer SP, Sauer K, Klein MP, Yachandra VK. J Am Chem Soc. 2001;123:7804–7820. doi: 10.1021/ja004307+. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Iuzzolino L, Dittmer J, Dörner W, Meyer-Klaucke W, Dau H. Biochemistry. 1998;37:17112–17119. doi: 10.1021/bi9817360. [DOI] [PubMed] [Google Scholar]
- 30.Visser H, Anxolabéhère-Mallart E, Bergmann U, Glatzel P, Robblee JH, Cramer SP, Girerd J-J, Sauer K, Klein MP, Yachandra VK. J Am Chem Soc. 2001;123:7031–7039. doi: 10.1021/ja004306h. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Kirby JA, Goodin DB, Wydrzynski T, Robertson AS, Klein MP. J Am Chem Soc. 1981;103:5537–5542. [Google Scholar]
- 32.Bergmann U, Glatzel P, Robblee JH, Messinger J, Fernandez C, Cinco R, Visser H, McFarlane K, Bellacchio E, Pizarro S, Sauer K, Yachandra VK, Klein MP, Cox BL, Nealson KH, Cramer SP. J Synchrotron Radiat. 2001;8:199–203. doi: 10.1107/s0909049500016484. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Fernandez C, Cinco RM, Robblee JH, Messinger J, Pizarro SA, Sauer K, Klein MP, Yachandra VK. In: Proceedings of the XIth international congress on photosynthesis: mechanisms and effects. Garab G, editor. vol 2. Dordrecht: Kluwer; 1998. pp. 1399–1402. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.