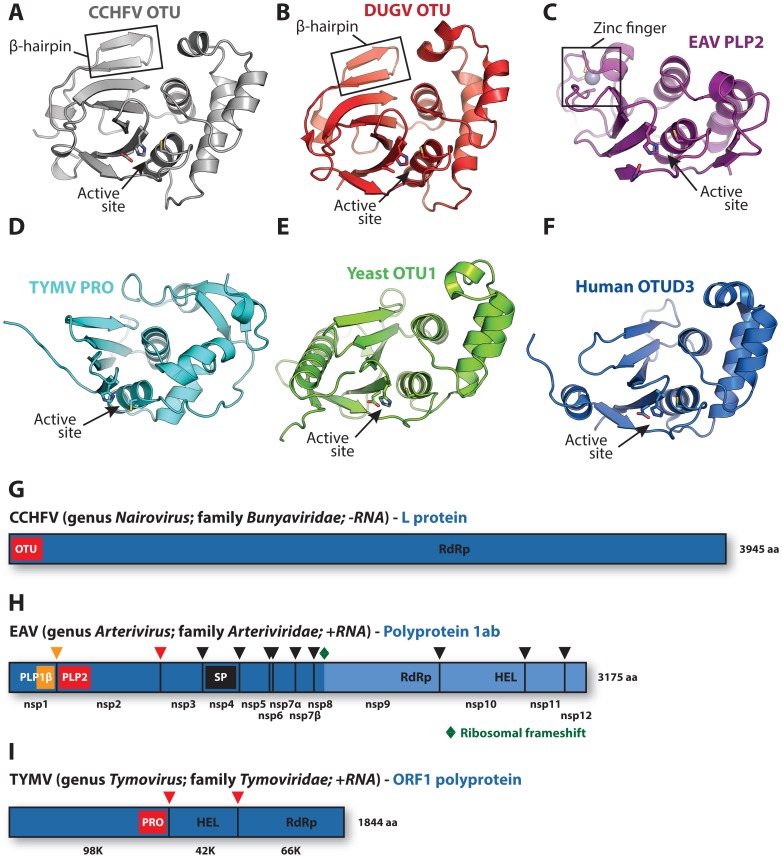
Figure 1. Viral and eukaryotic OTU domain structures and viral protein context.
Crystal structures of (A) CCHFV OTU (3PT2) [23], (B) DUGV OTU (4HXD) [25], (C) EAV PLP2 (4IUM) [26], (D) TYMV PRO (4A5U) [27], [28], (E) yeast OTU1 (3BY4) [57], and (F) human OTUD3 (4BOU) [46]. The β-hairpin motifs of CCHFV OTU and DUGV OTU are indicated in boxes in panels A and B, respectively, and the zinc-finger motif of EAV PLP2 is boxed in panel C. Active sites are indicated with arrows. The CCHFV OTU, DUGV OTU, EAV PLP2, and yeast OTU1 domains were crystallized in complex with Ub, which has been removed for clarity. Structure images were generated using PyMol [60]. (G) Schematic representation of the CCHFV large (L) protein [61], [62]. A similar organization is found in the DUGV L protein, but is not depicted. The OTU domain resides in the N-terminal region of this protein and is not involved in autoproteolytic cleavage events [48]. (H) Schematic representation of the EAV polyprotein 1ab [63]. PLP2 resides in nonstructural protein 2 (nsp2) and is responsible for the cleavage between nsp2 and nsp3 [51]. (I) Schematic representation of the TYMV ORF1 polyprotein [50]. PRO resides in the N-terminal product of this polyprotein and is responsible for two internal cleavages [49], [50]. Key replicative enzymes are indicated in G, H, and I. Colored arrowheads denote cleavage sites for the indicated protease domains. HEL, helicase; PLP, papain-like protease; RdRp, RNA-dependent RNA polymerase; SP, serine protease.