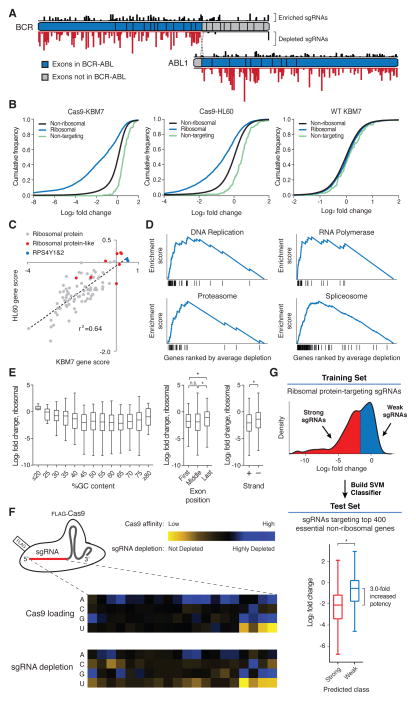
Fig. 3. Negative selection screens using CRISPR/Cas9 reveal rules governing sgRNA efficacy.
(A) Selective depletion of sgRNAs targeting exons of BCR and ABL1 present in the fusion protein. Individual sgRNAs are plotted according to their target sequence position along each gene and the height of each bar indicates the level of depletion observed. Boxes indicate individual exons. (B) Cas9-dependent depletion of sgRNAs targeting ribosomal proteins. Cumulative distribution function plots of log2 fold changes in sgRNA abundance before and after twelve cell doublings in Cas9-KBM7, Cas9-HL60 and WT-KBM7 cells. (C) Requirement of similar sets of ribosomal protein genes for proliferation in the HL60 and KBM7 cells. Gene scores are defined as the median log2 fold change of all sgRNAs targeting a gene. (D) Depleted sgRNAs target genes involved in fundamental biological processes. Gene Set Enrichment Analysis was performed on genes ranked by their combined depletion scores from screens in HL60 and KBM7 cells. Vertical lines underneath the x-axis denote members of the gene set analyzed. (E) Features influencing sgRNA efficacy. Depletion (log2 fold change) of sgRNAs targeting ribosomal protein genes was used as an indicator of sgRNA efficacy. Correlation between log2 fold changes and spacer %GC content (left), exon position targeted (middle) and strand targeted (right) are depicted. (*p<0.05) (F) sgRNA target sequence preferences for Cas9 loading and cleavage efficiency. Position-specific nucleotide preferences for Cas9 loading are determined by counting sgRNAs bound to Cas9 normalized to the number of corresponding genomic integrations. Heatmaps depict sequence-dependent variation in Cas9 loading (top) and ribosomal protein gene-targeting sgRNA depletion (bottom). The color scale represents the median value (of Cas9 affinity or log2 fold-change) for all sgRNAs with the specified nucleotide at the specified position. (G) sgRNA efficacy prediction. Ribosomal protein gene-targeting sgRNAs were designated as ‘weak’ or ‘strong’ based their log2 fold change and used to train a support-vector-machine (SVM) classifier. As an independent test, the SVM was used to predict the efficacy of sgRNAs targeting 400 essential non-ribosomal genes. (*p<0.05)