Abstract
Background
The oral cavity is a complex ecosystem where human chemical compounds coexist with a particular microbiota. However, shifts in the normal composition of this microbiota may result in the onset of oral ailments, such as periodontitis and dental caries. In addition, it is known that the microbial colonization of the oral cavity is mediated by protein-protein interactions (PPIs) between the host and microorganisms. Nevertheless, this kind of PPIs is still largely undisclosed. To elucidate these interactions, we have created a computational prediction method that allows us to obtain a first model of the Human-Microbial oral interactome.
Results
We collected high-quality experimental PPIs from five major human databases. The obtained PPIs were used to create our positive dataset and, indirectly, our negative dataset. The positive and negative datasets were merged and used for training and validation of a naïve Bayes classifier. For the final prediction model, we used an ensemble methodology combining five distinct PPI prediction techniques, namely: literature mining, primary protein sequences, orthologous profiles, biological process similarity, and domain interactions. Performance evaluation of our method revealed an area under the ROC-curve (AUC) value greater than 0.926, supporting our primary hypothesis, as no single set of features reached an AUC greater than 0.877. After subjecting our dataset to the prediction model, the classified result was filtered for very high confidence PPIs (probability ≥ 1-10−7), leading to a set of 46,579 PPIs to be further explored.
Conclusions
We believe this dataset holds not only important pathways involved in the onset of infectious oral diseases, but also potential drug-targets and biomarkers. The dataset used for training and validation, the predictions obtained and the network final network are available at http://bioinformatics.ua.pt/software/oralint.
Keywords: Protein-protein interactions, Oral interactome, Bayesian classification
Background
The majority of gene products that crowd a living cell interact, at least transiently, with other protein molecules. Virtually all cellular events, such as signal transduction, intracellular transport, DNA replication, transcription, translation, splicing, secretion, cell cycle control and intermediary metabolism, are mediated by protein-protein interactions (PPIs) [1]. The same applies to host-pathogen systems, where PPIs are essential in the establishment of infection [2]. The binding domains of interacting proteins reveal high structural and physical-chemical affinity with an associated degree of conservation. This is further evidenced by the fact that close protein homologs frequently interact in the same way [3-7]. With this in mind, we can expect understanding of the human interactome to provide insight into physiopathological mechanisms [8].
Numerous experimental techniques have been explored to attain the human interactome: two-hybrid screening [9,10], affinity purification mass spectrometry [11], DNA microarrays [12], protein microarrays [13-15], synthetic lethality [16], phage display [17], X-ray crystallography and nuclear magnetic resonance spectroscopy [18], fluorescence resonance energy transfer [19], surface plasmon resonance [20], atomic force microscopy [21], and electron microscopy [22]. These methods have major drawbacks that render them non-applicable in large-scale PPI prediction, namely the amount of time, associated cost and minimal protein interaction network coverage per run. Additionally, high-throughput approaches are also often associated with low-specificity and large numbers of both false negatives and false positives [23]. Moreover, these techniques were developed to detect intra-species PPIs, which renders them sub-optimal in inter-species PPI identification. Still, experimental methods remain the only viable methodology to validate PPIs.
As an alternative to experimental methods, a wide range of computational approaches for the prediction of intra-species PPIs have been proposed. Computational methods can be categorized according to the types of information they analyze. One common approach consists of using text mining to extract known PPIs from the biomedical literature [24]. Additionally, there are methods based on genomic data (gene neighborhood [25-28], gene fusion [29,30], phylogenetic profiles [31-33], codon usage similarity [34]), protein structure (homology-based method [35], threading-based method [36]), domain information (single domain pairs [37-41], multi-domain pairs [42,43]), protein sequence [44-56], and Gene Ontology (GO) [57] annotation semantic similarity ([58-61]). In contrast, computational efforts to predict inter-species PPIs have been very limited. Dyer et al.[2] combined domain information with a maximum likelihood estimator algorithm [37], while Davis et al.[62] adapted an approach following the threading-based method [36]. To provide a better prediction, Tastan et al.[63] applied a method combining multiple data sources, and used a random forest classifier to predict interactions between HIV-1 virus and human proteins. Despite these advances, the interactomes of several species are still far from complete. Nonetheless, the results of some of these studies provide great working knowledge of the characteristics of protein and gene interaction networks. For instance, the topological characteristics of protein interaction networks (PINs) have been proven to reflect the functionality of the interacting genes. This was demonstrated in yeast, where essential genes were more likely to be well connected and globally centered in the PIN [64,65].
Here we present a computational model to predict inter-species PPIs within the human oral cavity, an environment particularly prone to bacterial colonization. This is mostly due to the fact that human, microbial and environmental factors interact in a dynamic equilibrium within the human oral cavity [66]. Determination of the salivary interactome will clarify the role of saliva in oral biology and enable the identification of disease biomarkers. The presence of blood exudate proteins and exfoliated epithelial cells in saliva suggest it may be an alternative to blood as a diagnostic fluid in many instances. Additionally, if we consider the systemic nature of saliva, the ease and low cost associated with its handling, and the minimal risk linked to its collection for both medical staff and the patient, the reason for studying the oral cavity becomes clear [67].
As a result of this work, analysis of the resulting PPI network revealed some interesting features. Some of the PPIs involving the Rothia mucilaginosa microorganism are very specific and relevant. Moreover, our method not only predicted new PPIs between periodontal pathogens and the host, but also PPIs between different periodontal pathogens, suggesting a synergistic course of action.
Results
We conducted a series of pre-test analyses to assess the performance of our model. Then, we proceeded to test our approach on high-quality experimental protein-protein interaction (PPI) data collected from five databases. The selected databases exclusively contain manually curated PPI data.
Computational model for predicting the human-microbial interactome
Figure 1 summarizes the procedure used to achieve the model of the human-microbial interactome. The starting point of this work is a set of 4,707 proteins identified by proteomic studies as being present in the oral cavity and available on the OralCard database [66,67].
Figure 1.
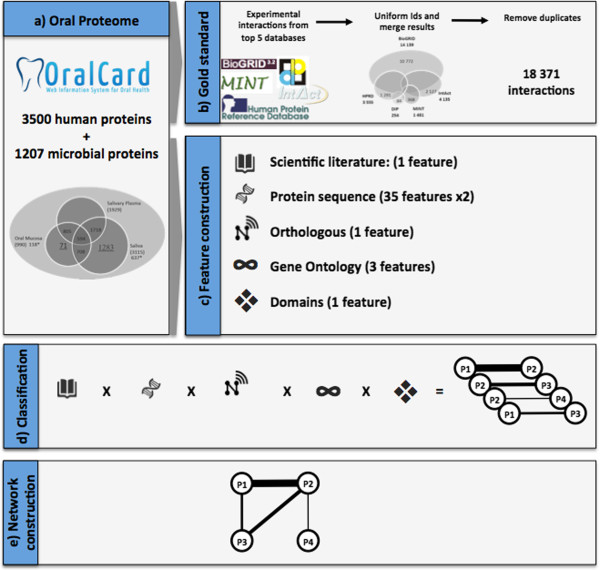
Workflow applied on the construction of the Human-microbial oral interactome”. It also contains footnote information: “a) the proteins identified on the oral proteome are obtained from the Oralcard database; b) the gold standard used for training and validation is obtained by combining the five most relevant curated protein interaction databases; c) for each protein interacting pair five clusters of features are constructed; d) the previously trained classifier is applied to each pair of interaction; and e) finally the interactome network is obtained by combining the individual pairs of proteins.
Since there is no well-established gold standard for PPIs, we collected data from five databases containing high-quality experimentally determined interactions as described further on in Methods. Extracted PPIs from the five databases were merged, creating our gold standard of positive interactions. The gold standard of negative interactions was obtained by randomly pairing the protein list on the premise that all protein pairs produced must differ from those on the positive dataset. A total of 18,371 positive and a similar number of negative pairs were obtained.
Simultaneously, for each possible pair of proteins, we constructed five clusters of features based on: (1) literature; (2) primary protein sequence information; (3) orthologous profiles; (4) biological process similarity, and; (5) enriched conserved domain pairs. This was performed by accessing public databases, extracting, and then processing the collected data.
The gold standard dataset was used to train a Naïve Bayes classifier and to perform further validations on the final model. The classifier was then applied to the set of all possible pairs of protein interactions. Finally, by aggregating all individual pairs of predicted interactions, the final network was obtained.
Evaluating the reconstruction of the human interactome
In this section, we evaluate the performance of the proposed method when applied to the set of human proteins from the gold standard. We performed a 5-fold cross-validation to assess the combined and individual contributions of the clusters of features. Table 1 shows the results for the performance of each individual cluster while Table 2 presents the contribution of each cluster to the final classifier by iteratively removing each cluster.
Table 1.
Analysis of the prediction performance of individual features
| Feature | AUC | CA | F1-score | Precision | Recall |
|---|---|---|---|---|---|
| + Literature |
0.781 |
0.722 |
0.723 |
0.721 |
0.726 |
| + Sequence |
0.877 |
0.784 |
0.790 |
0.768 |
0.813 |
| + GO |
0.817 |
0.742 |
0.748 |
0.735 |
0.760 |
| + COGs |
0.663 |
0.652 |
0.537 |
0.806 |
0.402 |
| + DDIs |
0.620 |
0.617 |
0.424 |
0.861 |
0.281 |
| Final Model | 0.926 | 0.850 | 0.851 | 0.848 | 0.854 |
For each line the metrics are obtained by considering only that cluster of features on the classifier. AUC, area under the receiver operating characteristic (ROC) curve; CA, classification accuracy.
Table 2.
Analysis of the contribution to the overall performance of individual cluster of features
| Feature | AUC | CA | F1-score | Precision | Recall |
|---|---|---|---|---|---|
| - Literature |
0.919 |
0.841 |
0.841 |
0.841 |
0.841 |
| - Sequence |
0.891 |
0.794 |
0.774 |
0.855 |
0.708 |
| - GO |
0.916 |
0.838 |
0.839 |
0.835 |
0.842 |
| - COGs |
0.923 |
0.846 |
0.847 |
0.842 |
0.852 |
| - DDIs |
0.911 |
0.831 |
0.834 |
0.819 |
0.850 |
| Final Model | 0.926 | 0.850 | 0.851 | 0.848 | 0.854 |
For each line the metrics are obtained by removing that cluster of features from the classifier. AUC, area under the receiver operating characteristic (ROC) curve; CA, classification accuracy.
The best performance is achieved through the ensemble of the five clusters, returning an area under the receiver operating characteristic (ROC) curve (AUC) of 0.926, a precision of 0.848 and a recall of 0.854. This result is above the performance of any individual feature and can only be achieved with the participation of all, meaning that all features are required and have a complementary contribution.
The Sequence is simultaneously the feature with the best overall performance (AUC = 0.877) and the one that causes the most negative impact when removed from the classifier, making the AUC drop to 0.891. It also has a very interesting recall of 0.813, partially due to the fact that all protein sequences are recognized and therefore the feature has full coverage.
In contrast, the clusters of orthologous groups (COGs, with AUC = 0.663) and domain-domain interactions (DDI, with AUC = 0.620) have the lowest individual AUCs, mainly due to the low coverage of their features. Despite that, they benefit from a considerably high precision that contributes positively to the final classifier. This is especially true for the COGs which, when removed, cause the major drop in precision.
The Literature and the Gene Ontology (GO) features, while not outstanding in any particular metric, have consistent performance on almost all metrics. Nevertheless, they make a very relevant contribution to the final classifier while the removal of the Literature causes a drop of the AUC to 0.919 and the GOs to 0.916.
Global characterization of the human-microbial interactome
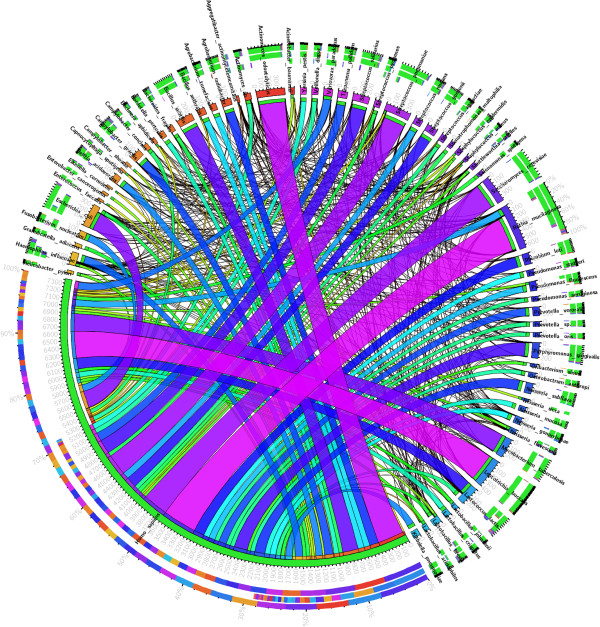
The classifier returned a set of 1.9 million possible interactions with a probability higher than 0.5. This corresponds to an average degree of 404 interactions per protein, which is much above the range of 3 to 30 documented in previous studies [68]. Additionally, there are reports of yeast two-hybrid screenings, the most commonly used high-throughput experimental method, reaching false-positive rates of 70%. With this in mind, and in order to minimize the presence of false-positives in our predicted interactome, we filtered our prediction results to consider only very high confidence PPIs (probability ≥ 1-10−7). We neglected the recall for the sake of precision. As can be observed in Figure 2, the cutoff of 1-10−7 is the lowest probability value where an increment does not imply a decrease in the number of interactions. This cut-off resulted in a total of 46,579 PPIs, with 37,407 being between human proteins, 6,394 between human and microbial proteins, and 2,778 between microbial proteins. The average number of protein interactions per protein after the cutoff was 8. Figure 3 is a visual representation of the interactions between the various organisms found in the oral cavity and the human host. Intra-species interactions are not shown. The thickness of the ribbons between each organism is correlated with the number of PPIs between both organisms, meaning that the organisms sharing highest number of PPIs with the human are Rothia mucilaginosa, Leptotrichia buccalis, and Actinomyces odontolyticus (strain independent).
Figure 2.
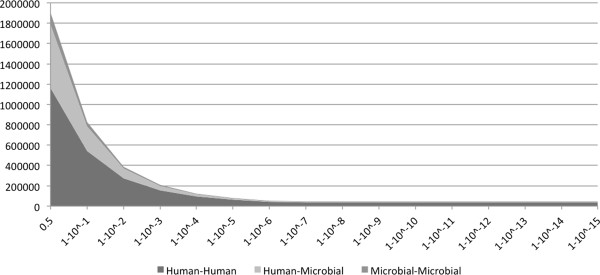
Plot with the relation of the number of interactions (y-axis) by classifier probability (x-axis).
Figure 3.
Representation of the Human-microbial inter-species protein interactions. Each section represents an organism. The ribbons connecting any two sections symbolize the PPIs between two organisms. The thickness of each ribbon correlates with the number of PPIs between both organisms.
With the exception of Homo sapiens with 3,030 proteins, the most represented organisms in the human oral cavity are Rothia mucilaginosa (strain DY-18) (Stomatococcus mucilaginosus), Actinomyces odontolyticus (strain ATCC 17982), and Streptococcus salivarius (strain SK126), with 68, 54, and 26 proteins, respectively. These organisms are opportunistic pathogens known to be associated with periodontitis [69] and caries [70].
The most frequent biological processes are related to host-microbial interactions: GO:0044281 (small molecule metabolic process, involved in 173 PPIs), GO:0019048 (viral interaction with host, involved in 161 PPIs), and GO:0045087 (innate immune response, involved in 145 PPIs).
We also identified the top three human hub-proteins present in our data: epidermal growth factor receptor (EGFR) (UniProt AC P00533, involved in 3247 PPIs), fibronectin (UniProt AC P02751, involved in 3143 PPIs), and cullin-associated NEDD8-dissociated protein 1 (CAND1) (UniProt AC Q86VP6, involved in 2911 PPIs). In terms of non-human original hub-proteins, the most common are a serine/threonine protein kinase from Leptotrichia buccalis (UniProt AC C7NEK0, involved in 258 PPIs), a kinase domain protein from Parviromonas micra ATCC 33270 (UniProt AC A8SM03, involved in 194 PPIs), and Ras-related protein SEC4 from Saccharomyces cerevisiae (UniProt AC P07560, involved in 163 PPIs).
Discussion
Functional analysis of the human-microbial interactome
Unsurprisingly, the most frequent GO biological processes in our final PPI dataset are associated with host-pathogen interactions. The preeminence of innate immune response and viral interaction with host as the most frequent biological processes are self-explanatory. However, the association between small molecule metabolism and host-microbial interactions is not so direct.
When faced with an infection, the body will respond by initiating two major cellular signaling pathways with opposing functions: the nuclear factor (NF)-kB and glucocorticoid-mediated signal transduction cascades. While the NF-kB pathway promotes the immune response and inflammation, the glucocorticoid-mediated signal transduction cascade suppresses it. In order to explain the association between small molecule metabolism and host-pathogen interactions we must focus on the NF-kB cascade, as it is known to mediate the transcriptional activation of several cytokines (cell-signaling molecules) involved in immunity [71]. Tumor necrosis factor (TNF)-α and TNF-β, two of these cytokines, play key roles in immune regulation and inflammation [72]. However, these cytokines are mainly responsible for the metabolic instabilities that occur during the infection, as they increase the metabolism of triglycerides inducing hyperlipidemia (escalation of blood lipid levels), stimulate lipolysis (degradation of lipids), accelerate glycogen breakdown and glucose consumption and uptake, and increase the serum levels of hormones that regulate glucose metabolism. These metabolic changes possibly explain the great number of “small molecule metabolic process” biological processes.
Analysis of hub proteins
The top three hub proteins identified share a common trait: these are exploited by pathogens in an attempt to gain entry to the host and survive inside it.
EGFR is a transmembrane protein mainly produced in the salivary glands and the kidneys [73]. Its association with microbial invasion has already been reported for Salmonella typhimurium[74], Candida albicans[75], Reovirus[76], and Vaccinia virus[77]. Apparently, all these pathogens initiate cellular invasion, at least to some extent, by binding to EGFR. This suggests the possibility that several other pathogens are using the EGFR to start host colonization, as supported by Buret et al. [78].
Similarly to EGFR, fibronectin appears to also play the role of a “microbial-anchor”. This glycoprotein is found bound to the β1 integrins in the cell surface, and is generally seen as a key protein for bacterial adhesion within the oral cavity [79,80].
The CAND1 protein, formerly TIP120A, was found to interact with most of the proteins in the Cullin family [81]. The Cullin protein family plays a key role in the ubiquitination of cellular proteins, i.e. performing a post-translational modification in order to label the target protein with ubiquitin molecules. This labeling frequently results in the commitment of the ubiquitin-linked protein to proteasomal degradation [82]. Consequently, CAND1 was suggested to function as a global regulator of cullin-containing ubiquitin ligases [81,83]. Being one of the top hub-proteins, we investigated the relationship between the ubiquitination pathway and pathogen colonization of the host cells. As expected, we found that certain bacteria corrupt the ubiquitination machinery as a means of regulating their virulence factors, or to trigger internalization of bacteria into host cells [84]. Such a mechanism improves the survival and replication chances of bacteria inside the host.
Study of the microbiome role in periodontitis
When the data analysis is focused on a particular disease such as periodontal disease four main features can be observed: i) Rothia mucilaginosa, a microorganism present in the normal human oral microbiome but considered an opportunistic pathogen [85], is the species with the most interactions, with some of them revealing important and specific interactions; ii) new interactions are predicted between periodonto-pathogens and the host, and; iii) interactions between periodonto-pathogens are also predicted, most likely explaining a synergistic course of action, as has been previously proposed [86].
Regarding the first observation, the analysis of the sub-network pertaining to Rothia mucilaginosa shares the characteristics previously described for the hub proteins with 37/638 interactions with the EFGR protein, 40/638 interactions with fibronectin and 34/638 interactions with CAND1. Furthermore, this sub-network presents two predicted interactions which have not been described before: R. mucilaginosa proteins D2NSF5 and C6R5R8, which are predicted to interact with human immunoglobulin chains (P01719 and P01781), and could be related to the immune response specific for this species, explaining why these interactions are worth investigating.
If we consider the bacteria most associated with periodontal disease, our model predicts few interactions between A. actinomycemcomitans, P. gingivalis, and the host proteins. As mentioned before, this is due to the fact that these organisms are not well represented in the original protein data set. However, besides the interactions predicted between these bacteria and the human hub proteins described above, in the case of Porphyromonas gingivalis it is possible to identify at least two potentially interesting new types of interactions between bacterial ribosomal proteins and a major histocompatibility complex protein (P30461). Furthermore, we also identified a possible interaction between the bacterial enolase (Q7MTV8) and a host aquaporin which could interfere with the homeostasis mechanisms of the host. Additionally, when we consider the interactions of P. gingivalis with other bacteria, we find that the same enolase might interact with outer membrane proteins of Haemophilus influenza and Pasteurella multocida. The role of bacterial enolase as a multitask protein involved not only in carbohydrate metabolism but also in virulence has been recognized recently [87].
This suggests that previously unknown and important PPIs for oral colonization and biofilm formation may be present in this dataset. Finally the fact there are possible interactions between P.gingivalis proteases and those of other periodonto-pathogens such as Kingella oralis and Treponema denticola is interesting. This may even shed some light on the synergistic aspects of oral biofilm in periodontal disease [86].
Conclusion
The continuous yield of large-scale data mainly from microarrays and yeast two-hybrid studies has made the study of PPIs very appealing. The main issue associated with PPI study is the high prevalence of false positives and negatives in experimental PPI data. Being the only “reliable” source of PPIs, inaccurate experimental PPI data will contaminate training datasets and therefore compromise the performance of computational PPI prediction methods. For this reason, we believe that an improvement in the quality of experimental PPI data will greatly impact the performance of new computational PPI prediction approaches. While this is not the case at present, we must consider how to avoid the effects of false positives and false negatives in the final PPI prediction model.
We proposed a probabilistic Bayesian-based method to integrate several data sources, to obtain more robust and reliable PPI predictions. By applying naïve Bayes, we automatically up-weigh the most informative features and down-weigh the less informative ones, allowing for automatic error-correction.
Our individual feature analysis results show a great relevance of the selected features. When applied on a naïve Bayes classifier, the individual features synergize, boosting the AUC up to 0.926. This suggests that the reliability of prediction improves with the increase of significant features, meaning that the ensemble final model actually reduces the disadvantages of the individual methods.
Cytoscape was successfully used to validate the network when tested with real pathway examples, discovering new potentially interesting interactions in oral biology, both between the host and the periodontal pathogens and between different periodontal pathogens.
We believe our work may be applied in several scientific areas, and even in other PPI related studies. An example is biomedical PPI screening, to assess if interactions of particular interest might occur and what the related interaction probability is. Another example is pharmacologic research, as a well-established PPI network can provide insights on potential drug targets, but also new uses for existent in-market drugs. Finally, and based on the fact that protein interaction networks are dynamic [88], our work can support researchers in identifying evolutionary patterns.
Methods
Oral proteome
As a starting point for our study we used 4,707 proteins, 3500 from Human and 1207 from microbial, available on the OralCard database [66,67].
These proteins were identified via proteomic analysis of the saliva, frequently by using 2D electrophoresis/mass spectrometry or 2D liquid chromatography/mass spectrometry. By the end of 2012 the salivary proteome was determined to contain 3500 proteins from human origin and 1207 from microbial sources.
Predictive dataset construction
The use of positive (interacting pairs of proteins) and negative (non-interacting pairs of proteins) examples is required for training and assessing the performance of the classifier. All data used in the construction of the positive data set (PDS) and the negative data set (NDS) was downloaded in March 2013.
Positive dataset
We collected experimental oral protein-protein interaction (PPI) data from five databases: 14,139 PPIs from BIOGRID [89], 254 PPIs from DIP [90], 3,555 PPIs from HPRD [91], 4,135 PPIs from IntAct [92], and 1,481 PPIs from MINT [93], totaling 23,564 protein interactions (Figure 4).
Figure 4.
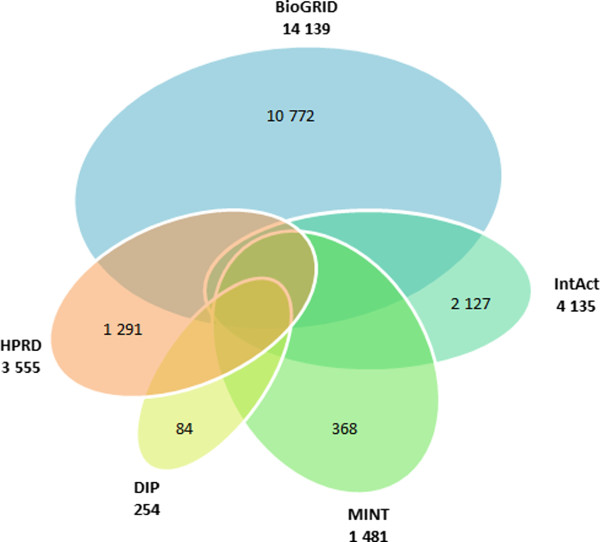
Venn diagram representing the intersections between the five high-quality experimentally determined protein-protein interaction databases.
All the interacting protein pairs were identified by their UniProtKB [94] Accession IDs for normalization purposes. In some instances it was necessary to convert the database own identifiers to UniProtKB Accession IDs. The BioGRID database represents interacting protein pairs using their own identifiers and Entrez Gene IDs. To match them to UniProtKB AccessionIDs we extracted the Gene IDs from the protein pairs and downloaded the list of respective gene products in the UniProtKB Accession ID format. UniProtKB allows direct mapping from the MINT and DIP databases to another identifier. A list of PPI pairs from both databases was uploaded to the UniProtKB mapping feature, resulting in two different sets of UniProtKB Accession ID pairs. HPRD uses its own identification system coupled with NCBI Reference Sequence Accession IDs (RefSeq) to classify PPI pairs. All the RefSeq Protein IDs were converted to UniProtKB Accession IDs and paired accordingly. IntAct PPI pairs are identified with UniProtKB Accession IDs and were directly extracted.
PPI pairs from the five databases were merged and repeated entries were removed. From a total of 23,564 PPIs, 5,193 duplicated entries were removed, resulting in a PDS of 18,371 protein pairs.
Negative dataset
The selection of negative examples to integrate the negative data was based on two methods described in the literature [95]. These methods consist of randomly selecting protein pairs that are not present in a veto list containing all PPIs from the positive data set. The use of this strategy was considered acceptable because the probability of committing an error while picking a random pair is low:
where N is the number of proteins and K is the average degree for the final PPI network. In this study the N is 4,707 and for PPIs the typical value of K is between 6 and 16.
With this strategy we generated a NDS of a size similar to that of the PDS (18,348 “negative” protein pairs), and combined it with the PDS to obtain a training data set with 36,719 PPIs.
Feature construction
In this section we describe the procedure for construction of the five clusters of features. The final results are summarized in Table 3.
Table 3.
Relative coverage of protein-protein interactions present in the training and test data by individual feature clusters
| |
Training data |
Classification |
||
|---|---|---|---|---|
| #Interactions | % of total | #Interactions | % of total | |
| Literature |
22,720 |
61.9% |
4,698,390 |
69.9% |
| Sequence |
35,379 |
96.4% |
6,703,945 |
99.8% |
| GO |
23,769 |
64.8% |
5,130,103 |
76.4% |
| COGs |
9,636 |
26.3% |
1,324,230 |
19.7% |
| DDIs |
5,994 |
16.3% |
516,609 |
7.7% |
| Total | 36,698 | 100.0% | 6,716,792 | 100.0% |
GO, gene ontology; COGs, clusters of orthologous groups; DDIs, domain-domain interactions.
Literature
The literature-based protein-protein interaction scores were calculated by the method described in van Haagen et al.[96]. This method is based on comparing the semantic contexts in which two proteins are mentioned in the published literature. The rationale for the method is that two proteins occurring in similar contexts will have a higher similarity score and are therefore more likely to interact. The semantic context for a given protein is defined by the concepts, from a pre-defined vocabulary, that are frequently mentioned in the same articles with that protein, and is represented by a vector containing a weight for each concept. These weights are based on the co-occurrence statistics, and measure the degree of association between the protein and each concept. Following Jelier et al.[97], we use the symmetric uncertainty coefficient U (Xi, Yj) – where Xi is in this case the protein of interest and Yj is any other concept in the vocabulary – as the weights used for creating the concept profiles:
Where H (X) is the entropy for X and H (X, Y) is the joint entropy for X and Y, calculated based on document frequency counts.
We used a corpus of nearly one million abstracts, obtained by searching Pubmed with 17,402 names and synonyms extracted from UniProtKB for 4,707 proteins in the dataset, after removing nonsensical names such as “uncharacterized protein”. To identify the concepts mentioned in the texts we used Gimli [98], a machine-learning tool for gene and protein name recognition, together with dictionary matching to recognize other concepts from ten different semantic types including chemical entities, anatomical terms, diseases, pathways and GO terms. The dictionaries used contain around 1,3 million distinct names for around 400 thousand concepts. Based on the concept annotation of this corpus, we were able to calculate concept profiles for 22,720 protein pairs from the training dataset and 4,698,390 protein pairs for the classification dataset.
Primary protein sequence information
Several studies have been carried out where detection of protein-protein interaction is derived from information directly extracted from the amino-acid sequences [44-56]. The results indicate that the sequence information alone is sufficient to detect PPIs with reasonable accuracy [87] but may be improved if combined with other strategies.
Taking into account the primary protein sequence information, the following features have been considered in this work: occurrence of the 20 amino-acids in the protein sequence, protein atomic composition, molecular weight and atomic weight, forming a vector of 27 features. The interacting protein pair (X, Y) is represented by concatenating the corresponding features vectors Fx and Fy, represented by (Fx, Fy).
We were able to obtain the sequence profile for 35,379 proteins pairs from the training dataset and 6,703,945 protein pairs for the classification dataset.
Orthologous profiles
By definition, clusters of orthologous groups (COGs) are sets of orthologous genes or orthologous groups of paralogs from three or more phylogenetic trees. In essence, this means that two proteins from different lineages belonging to the same COG are orthologous. Orthologs are genes in different species that evolved from a common ancestor by speciation (i.e. convergent evolution). In contrast, paralogs are genes related by duplication within a genome [99].
Lee et al. [100] aimed to expand the interactomes of various organisms by applying orthology-based methods in inter-species PPI prediction. They expanded orthologous pairs of 18 eukaryotic organisms and merged them with experimental PPI datasets, allowing the inference of PPIs for various species.
In this work we used the Search Tool for the Retrieval of Interacting Genes/Proteins (STRING) [101] database to obtain COGs and their respective combined scores. The combined score is computed by integrating the likelihoods from the different types of evidence, correcting for the probability of randomly observing an interaction [101]. This enhances the predictive performance of the method, as a combined score is only computed when more than one of the data sources in STRING supports a given association.
We were able to obtain the orthologous profile for 9,636 protein pairs from the training dataset and 1,324,230 proteins pairs for the classification dataset.
Biological process similarity
Previous studies have explored the use of GO annotation similarity between two proteins as a PPI predictor [59,102-105]. We downloaded biological process information from the GO Consortium [57] in March 2013 and calculated the depth of the GO terms (nodes) in the Directed Acyclic Graph (DAG), and the total number of proteins comprised between the smallest shared biological process (SSBP) for each pair of proteins and the following three branches. Since the depth of the GO terms in the DAG is implied in the total number of proteins, post-test odds analysis was performed solely on this feature to avoid redundancy. Such an approach was based on the general hypothesis that it is progressively more likely for the proteins comprised within a biological process to interact, if the total number of proteins involved in that process is progressively smaller.
We were able to obtain the gene ontology profile for 23,769 protein pairs from the training dataset and 5,130,103 protein pairs for the classification dataset.
Enriched conserved domain pairs
The Database of Protein Domain Interactions [106] (DOMINE) contains binary domain-domain interaction (DDI) data compiled from a collection of 15 databases and DDI prediction methods. Additionally, DOMINE provides a quality measure of the DDI confidence, as well as a binary classification of whether the domains are part of the same GO biological process. Here, we assume that whenever two given proteins possess one or more interacting domains between them, those proteins will interact. We adopted this DDI data collection as individual features in our approach. Since DOMINE provides DDI information from several sources, we tallied the number of sources that identified a DDI. This strategy confers higher reliability on DDI pairs with higher scores (closer to 15, the maximum number of DDI sources).
We were able to obtain the domain profile for 5,994 protein pairs from the training dataset and 516,609 protein pairs for the classification dataset.
Data classification and validation
The proposed approach was developed, tested, optimized and performed using Orange, an open-source bioinformatics tool featuring Python scripting and a visual and programmatic interface. We used the naïve Bayes [107] classifier to predict PPIs in our data. The naïve Bayes classifier calculates the conditional probability of each attribute Ai given the class label C, from the training data. The Bayes rule is then applied to calculate the probability of C given the specific instance of A1 ,…, An, and then assessing the class with the greatest posterior probability, ensuing classification [108].
The receiver operating characteristic (ROC) curve, which is the plot of the true positive (TP) rate with the false positive (FP) rate, depicting the relative trade-off between both rates [109] was used to evaluate the method’s performance. When comparing classifiers with very similar ROC curves, it may be necessary to estimate a single scalar value to represent the expected performance. One of the most common methods is calculation of the area under the ROC curve (AUC) [110], which we used to compare the naïve Bayes classifier. Therefore, we assessed the individual contributions of each feature in terms of classification accuracy (CA), area under curve (AUC), F1-score, precision and recall.
Interactome analysis
We used Cytoscape to visualize and validate the obtained PPI network. The PPIs were classified as “HUMAN-HUMAN”, if the interacting proteins were only of human origin, as “MICRO-MICRO”, if the interacting proteins were only of microbial origin, or as “HUMAN-MICRO”.
We imported the network data to Cytoscape defining the two proteins in the same interacting protein pair as Source Interaction (protein one) and Target Interaction (protein two). The chosen Interaction Type was the above-mentioned organism-organism classification. A file containing node attributes was also imported, containing microorganism and biological process information extracted from the UniProt database pertaining to each individual protein in the network.
Availability
All data required to analyze the results and re-run this experiment are available for download at http://bioinformatics.ua.pt/software/oralint. This includes the unique list of UniProt AC for the proteins in the oral cavity, the gold standard of interactions, the dataset used for training and validation, the predictions obtained, and the Cytoscape project file with the network.
Competing interests
The authors declare that they have no competing interests.
Authors’ contributions
EDC participated in the design of the study, constructed the positive and negative datasets, performed the analysis of hub proteins, characterized and analysed the human-microbial interactome, and drafted the manuscript. JPA conceived the study, participated in its design, performed feature construction and selection, parameterized the classifier, and helped to draft the manuscript. SM performed the text-mining analysis and helped to draft the manuscript. CP carried out primary protein sequence analysis and helped to draft the manuscript. NR and MJC analysed the role of the microbiome in periodontitis and helped to draft the manuscript. MB and JLO participated in the design and conception of the study, coordinated it, and helped to draft the manuscript. All authors read and approved the final manuscript.
Contributor Information
Edgar D Coelho, Email: eduarte@ua.pt.
Joel P Arrais, Email: jpa@dei.uc.pt.
Sérgio Matos, Email: aleixomatos@ua.pt.
Carlos Pereira, Email: cpereira@dei.uc.pt.
Nuno Rosa, Email: nrosa@crb.ucp.pt.
Maria José Correia, Email: mcorreia@crb.ucp.pt.
Marlene Barros, Email: mbarros@crb.ucp.pt.
José Luís Oliveira, Email: jlo@ua.pt.
Acknowledgements
This work has received support from the RD-CONNECT project (EC contract number 305444). Edgar D. Coelho is funded by Fundação para a Ciência e Tecnologia, FCT, under Grant SFRH/BD/86343/2012.
References
- Phizicky EM, Fields S. Protein-protein interactions: methods for detection and analysis. Microbiol Rev. 1995;59:94–123. doi: 10.1128/mr.59.1.94-123.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dyer MD, Murali TM, Sobral BW. Computational prediction of host-pathogen protein–protein interactions. Bioinformatics. 2007;23:i159–i166. doi: 10.1093/bioinformatics/btm208. [DOI] [PubMed] [Google Scholar]
- Littler SJ, Hubbard SJ. Conservation of orientation and sequence in protein domain–domain interactions. J Mol Biol. 2005;345:1265–1279. doi: 10.1016/j.jmb.2004.11.011. [DOI] [PubMed] [Google Scholar]
- Valdar WS, Thornton JM. Protein-protein interfaces: analysis of amino acid conservation in homodimers. Proteins. 2001;42:108–124. doi: 10.1002/1097-0134(20010101)42:1<108::AID-PROT110>3.0.CO;2-O. [DOI] [PubMed] [Google Scholar]
- Aloy P, Ceulemans H, Stark A, Russell RB. The relationship between sequence and interaction divergence in proteins. J Mol Biol. 2003;332:989–998. doi: 10.1016/j.jmb.2003.07.006. [DOI] [PubMed] [Google Scholar]
- Teichmann SA. The constraints protein-protein interactions place on sequence divergence. J Mol Biol. 2002;324:399–407. doi: 10.1016/S0022-2836(02)01144-0. [DOI] [PubMed] [Google Scholar]
- Panchenko AR, Wolf YI, Panchenko LA, Madej T. Evolutionary plasticity of protein families: coupling between sequence and structure variation. Proteins. 2005;61:535–544. doi: 10.1002/prot.20644. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rual J-F, Venkatesan K, Hao T, Hirozane-Kishikawa T, Dricot A, Li N, Berriz GF, Gibbons FD, Dreze M, Ayivi-Guedehoussou N, Klitgord N, Simon C, Boxem M, Milstein S, Rosenberg J, Goldberg DS, Zhang LV, Wong SL, Franklin G, Li S, Boxem M, Milstein S, Rosenberg J, Goldberg DS, Zhang LV, Wong SL, Franklin G, Li S, Albala JS, Lim J. et al. Towards a proteome-scale map of the human protein-protein interaction network. Nature. 2005;437:1173–1178. doi: 10.1038/nature04209. [DOI] [PubMed] [Google Scholar]
- Uetz P, Giot L, Cagney G, Mansfield TA, Judson RS, Knight JR, Lockshon D, Narayan V, Srinivasan M, Pochart P, Qureshi-Emili A, Li Y, Godwin B, Conover D, Kalbfleisch T, Vijayadamodar G, Yang M, Johnston M, Fields S, Rothberg JM. A comprehensive analysis of protein-protein interactions in Saccharomyces cerevisiae. Nature. 2000;403:623–627. doi: 10.1038/35001009. [DOI] [PubMed] [Google Scholar]
- Ito T, Chiba T, Ozawa R, Yoshida M, Hattori M, Sakaki Y. A comprehensive two-hybrid analysis to explore the yeast protein interactome. Proceedings of the National Academy of Sciences. 2001;98:4569–4574. doi: 10.1073/pnas.061034498. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rigaut G, Shevchenko A, Rutz B, Wilm M, Mann M, Séraphin B. A generic protein purification method for protein complex characterization and proteome exploration. Nat Biotechnol. 1999;17:1030–1032. doi: 10.1038/13732. [DOI] [PubMed] [Google Scholar]
- Eisen MB, Spellman PT, Brown PO, Botstein D. Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci U S A. 1998;95:14863–14868. doi: 10.1073/pnas.95.25.14863. [DOI] [PMC free article] [PubMed] [Google Scholar]
- MacBeath G, Schreiber SL. Printing proteins as microarrays for high-throughput function determination. Science. 2000;289:1760–1763. doi: 10.1126/science.289.5485.1760. [DOI] [PubMed] [Google Scholar]
- Zhu H, Bilgin M, Bangham R, Hall D, Casamayor A, Bertone P, Lan N, Jansen R, Bidlingmaier S, Houfek T, Mitchell T, Miller P, Dean RA, Gerstein M, Snyder M. Global analysis of protein activities using proteome chips. Science. 2001;293:2101–2105. doi: 10.1126/science.1062191. [DOI] [PubMed] [Google Scholar]
- Jones RB, Gordus A, Krall JA, MacBeath G. A quantitative protein interaction network for the ErbB receptors using protein microarrays. Nature. 2006;439:168–174. doi: 10.1038/nature04177. [DOI] [PubMed] [Google Scholar]
- Ye P, Peyser BD, Pan X, Boeke JD, Spencer FA, Bader JS. Gene function prediction from congruent synthetic lethal interactions in yeast. Mol Syst Biol. 2005;1(2005):0026. doi: 10.1038/msb4100034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith GP. Filamentous fusion phage: novel expression vectors that display cloned antigens on the virion surface. Science. 1985;228:1315–1317. doi: 10.1126/science.4001944. [DOI] [PubMed] [Google Scholar]
- Tong AHY, Evangelista M, Parsons AB, Xu H, Bader GD, Pagé N, Robinson M, Raghibizadeh S, Hogue CWV, Bussey H, Andrews B, Tyers M, Boone C. Systematic genetic analysis with ordered arrays of yeast deletion mutants. Science. 2001;294:2364–2368. doi: 10.1126/science.1065810. [DOI] [PubMed] [Google Scholar]
- Yan Y, Marriott G. Analysis of protein interactions using fluorescence technologies. Curr Opin Chem Biol. 2003;7:635–640. doi: 10.1016/j.cbpa.2003.08.017. [DOI] [PubMed] [Google Scholar]
- Cooper MA. Label-free screening of bio-molecular interactions. Anal Bioanal Chem. 2003;377:834–842. doi: 10.1007/s00216-003-2111-y. [DOI] [PubMed] [Google Scholar]
- Yang Y, Wang H, Erie DA. Quantitative characterization of biomolecular assemblies and interactions using atomic force microscopy. Methods. 2003;29:175–187. doi: 10.1016/S1046-2023(02)00308-0. [DOI] [PubMed] [Google Scholar]
- Baumeister W, Grimm R, Walz J. Electron tomography of molecules and cells. Trends Cell Biol. 1999;9:81–85. doi: 10.1016/S0962-8924(98)01423-8. [DOI] [PubMed] [Google Scholar]
- Xia JF, Wang SL, Lei YK. Computational methods for the prediction of protein-protein interactions. Protein Pept Lett. 2010;17:1069–1078. doi: 10.2174/092986610791760405. [DOI] [PubMed] [Google Scholar]
- Jaeger S, Gaudan S, Leser U, Rebholz-Schuhmann D. Integrating protein-protein interactions and text mining for protein function prediction. BMC Bioinformatics. 2008;9(Suppl 8):S2. doi: 10.1186/1471-2105-9-S8-S2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tamames J, Casari G, Ouzounis C, Valencia A. Conserved clusters of functionally related genes in two bacterial genomes. J Mol Evol. 1997;44:66–73. doi: 10.1007/PL00006122. [DOI] [PubMed] [Google Scholar]
- Dandekar T, Snel B, Huynen M, Bork P. Conservation of gene order: a fingerprint of proteins that physically interact. Trends Biochem Sci. 1998;23:324–328. doi: 10.1016/S0968-0004(98)01274-2. [DOI] [PubMed] [Google Scholar]
- Overbeek R, Fonstein M, D'Souza M, Pusch GD, Maltsev N. The use of gene clusters to infer functional coupling. Proc Natl Acad Sci U S A. 1999;96:2896–2901. doi: 10.1073/pnas.96.6.2896. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blumenthal T. Gene clusters and polycistronic transcription in eukaryotes. Bioessays. 1998;20:480–487. doi: 10.1002/(SICI)1521-1878(199806)20:6<480::AID-BIES6>3.0.CO;2-Q. [DOI] [PubMed] [Google Scholar]
- Enright AJ, Iliopoulos I, Kyrpides NC, Ouzounis CA. Protein interaction maps for complete genomes based on gene fusion events. Nature. 1999;402:86–90. doi: 10.1038/47056. [DOI] [PubMed] [Google Scholar]
- Marcotte EM, Pellegrini M, Ng HL, Rice DW, Yeates TO, Eisenberg D. Detecting protein function and protein-protein interactions from genome sequences. Science. 1999;285:751–753. doi: 10.1126/science.285.5428.751. [DOI] [PubMed] [Google Scholar]
- Ouzounis C, Kyrpides N. The emergence of major cellular processes in evolution. FEBS Lett. 1996;390:119–123. doi: 10.1016/0014-5793(96)00631-X. [DOI] [PubMed] [Google Scholar]
- Pellegrini M, Marcotte EM, Thompson MJ, Eisenberg D, Yeates TO. Assigning protein functions by comparative genome analysis: protein phylogenetic profiles. Proc Natl Acad Sci U S A. 1999;96:4285–4288. doi: 10.1073/pnas.96.8.4285. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barker D, Pagel M. Predicting functional gene links from phylogenetic-statistical analyses of whole genomes. PLoS Comput Biol. 2005;1:e3. doi: 10.1371/journal.pcbi.0010003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Najafabadi HS, Salavati R. Sequence-based prediction of protein-protein interactions by means of codon usage. Genome Biol. 2008;9:R87. doi: 10.1186/gb-2008-9-5-r87. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aloy P, Russell RB. Interrogating protein interaction networks through structural biology. Proc Natl Acad Sci. 2002;99:5896–5901. doi: 10.1073/pnas.092147999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lu L, Lu H, Skolnick J. MULTIPROSPECTOR: an algorithm for the prediction of protein-protein interactions by multimeric threading. Proteins. 2002;49:350–364. doi: 10.1002/prot.10222. [DOI] [PubMed] [Google Scholar]
- Sprinzak E, Margalit H. Correlated sequence-signatures as markers of protein-protein interaction. J Mol Biol. 2001;311:681–692. doi: 10.1006/jmbi.2001.4920. [DOI] [PubMed] [Google Scholar]
- Deng M, Mehta S, Sun F, Chen T. Inferring domain-domain interactions from protein-protein interactions. Genome Res. 2002;12:1540–1548. doi: 10.1101/gr.153002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen L, Wu LY, Wang Y, Zhang XS. Inferring protein interactions from experimental data by association probabilistic method. Proteins. 2006;62:833–837. doi: 10.1002/prot.20783. [DOI] [PubMed] [Google Scholar]
- Morrison JL, Breitling R, Higham DJ, Gilbert DR. A lock-and-key model for protein-protein interactions. Bioinformatics. 2006;22:2012–2019. doi: 10.1093/bioinformatics/btl338. [DOI] [PubMed] [Google Scholar]
- Huang C, Morcos F, Kanaan SP, Wuchty S, Chen DZ, Izaguirre JA. Predicting protein-protein interactions from protein domains using a set cover approach. IEEE/ACM Trans Comput Biol Bioinform. 2007;4:78–87. doi: 10.1109/TCBB.2007.1001. [DOI] [PubMed] [Google Scholar]
- Chen X-W, Liu M. Prediction of protein–protein interactions using random decision forest framework. Bioinformatics. 2005;21:4394–4400. doi: 10.1093/bioinformatics/bti721. [DOI] [PubMed] [Google Scholar]
- Wang R-S, Wang Y, Wu L-Y, Zhang X-S, Chen L. Analysis on multi-domain cooperation for predicting protein-protein interactions. BMC Bioinformatics. 2007;8:391. doi: 10.1186/1471-2105-8-391. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bock JR, Gough DA. Predicting protein–protein interactions from primary structure. Bioinformatics. 2001;17:455–460. doi: 10.1093/bioinformatics/17.5.455. [DOI] [PubMed] [Google Scholar]
- Bock JR, Gough DA. Whole-proteome interaction mining. Bioinformatics. 2003;19:125–134. doi: 10.1093/bioinformatics/19.1.125. [DOI] [PubMed] [Google Scholar]
- Martin S, Roe D, Faulon J-L. Predicting protein–protein interactions using signature products. Bioinformatics. 2005;21:218–226. doi: 10.1093/bioinformatics/bth483. [DOI] [PubMed] [Google Scholar]
- Ben-Hur A, Noble WS. Kernel methods for predicting protein–protein interactions. Bioinformatics. 2005;21:38–46. doi: 10.1093/bioinformatics/bti1016. [DOI] [PubMed] [Google Scholar]
- Pitre S, Dehne F, Chan A, Cheetham J, Duong A, Emili A, Gebbia M, Greenblatt J, Jessulat M, Krogan N, Luo X, Golshani A. PIPE: a protein-protein interaction prediction engine based on the re-occurring short polypeptide sequences between known interacting protein pairs. BMC Bioinformatics. 2006;7:365. doi: 10.1186/1471-2105-7-365. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nanni L, Lumini A. An ensemble of K-local hyperplanes for predicting protein–protein interactions. Bioinformatics. 2006;22:1207–1210. doi: 10.1093/bioinformatics/btl055. [DOI] [PubMed] [Google Scholar]
- Nanni L. Hyperplanes for predicting protein–protein interactions. Neurocomputing. 2005;69:257–263. doi: 10.1016/j.neucom.2005.05.007. [DOI] [Google Scholar]
- Shen J, Zhang J, Luo X, Zhu W, Yu K, Chen K, Li Y, Jiang H. Predicting protein–protein interactions based only on sequences information. Proc Natl Acad Sci. 2007;104:4337–4341. doi: 10.1073/pnas.0607879104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guo Y, Yu L, Wen Z, Li M. Using support vector machine combined with auto covariance to predict protein–protein interactions from protein sequences. Nucleic Acids Res. 2008;36:3025–3030. doi: 10.1093/nar/gkn159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xia JF, Han K, Huang DS. Sequence-based prediction of protein-protein interactions by means of rotation forest and autocorrelation descriptor. Protein Pept Lett. 2010;17:137–145. doi: 10.2174/092986610789909403. [DOI] [PubMed] [Google Scholar]
- Rajasekaran S, Merlin JC, Kundeti V, Mi T, Oommen A, Vyas J, Alaniz I, Chung K, Chowdhury F, Deverasatty S, Irvey TM, Lacambacal D, Lara D, Panchangam S, Rathnayake V, Watts P, Schiller MR. A computational tool for identifying minimotifs in protein-protein interactions and improving the accuracy of minimotif predictions. Proteins. 2011;79:153–164. doi: 10.1002/prot.22868. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Knisley D, Knisley J. Predicting protein–protein interactions using graph invariants and a neural network. Comput Biol Chem. 2011;35:108–113. doi: 10.1016/j.compbiolchem.2011.03.003. [DOI] [PubMed] [Google Scholar]
- Zhang Y, Zhang D, Mi G, Ma D, Li G, Guo Y, Li M, Zhu M. Using ensemble methods to deal with imbalanced data in predicting protein-protein interactions. Comput Biol Chem. 2012;36:36–41. doi: 10.1016/j.compbiolchem.2011.12.003. [DOI] [PubMed] [Google Scholar]
- Harris MA, Clark J, Ireland A, Lomax J, Ashburner M, Foulger R, Eilbeck K, Lewis S, Marshall B, Mungall C, Richter J, Rubin GM, Blake JA, Bult C, Dolan M, Drabkin H, Eppig JT, Hill DP, Ni L, Ringwald M, Balakrishnan R, Cherry JM, Christie KR, Costanzo MC, Dwight SS, Engel S, Fisk DG, Hirschman JE, Hong EL, Nash RS. et al. The gene ontology (GO) database and informatics resource. Nucleic Acids Res. 2004;32:D258–D261. doi: 10.1093/nar/gkh036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jain S, Bader GD. An improved method for scoring protein-protein interactions using semantic similarity within the gene ontology. BMC Bioinformatics. 2010;11:562. doi: 10.1186/1471-2105-11-562. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maetschke SR, Simonsen M, Davis MJ, Ragan MA. Gene Ontology-driven inference of protein–protein interactions using inducers. Bioinformatics. 2012;28:69–75. doi: 10.1093/bioinformatics/btr610. [DOI] [PubMed] [Google Scholar]
- Park B, Cui G, Lee H, Huang D-S, Han K. PPISearchEngine: gene ontology-based search for protein–protein interactions. Comput Methods Biomech Biomed Engin. 2012;16:1–8. doi: 10.1080/10255842.2011.631528. [DOI] [PubMed] [Google Scholar]
- Wu X, Zhu L, Guo J, Zhang DY, Lin K. Prediction of yeast protein-protein interaction network: insights from the Gene Ontology and annotations. Nucleic Acids Res. 2006;34:2137–2150. doi: 10.1093/nar/gkl219. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Davis FP, Barkan DT, Eswar N, McKerrow JH, Sali A. Host pathogen protein interactions predicted by comparative modeling. Protein Sci. 2007;16:2585–2596. doi: 10.1110/ps.073228407. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tastan O, Qi Y, Carbonell JG, Klein-Seetharaman J. Prediction of interactions between HIV-1 and human proteins by information integration. Pac Symp Biocomput. 2009. pp. 516–527. [PMC free article] [PubMed]
- Jeong H, Mason SP, Barabasi AL, Oltvai ZN. Lethality and centrality in protein networks. Nature. 2001;411:41–42. doi: 10.1038/35075138. [DOI] [PubMed] [Google Scholar]
- Wuchty S, Almaas E. Peeling the yeast protein network. Proteomics. 2005;5:444–449. doi: 10.1002/pmic.200400962. [DOI] [PubMed] [Google Scholar]
- Arrais JP, Rosa N, Melo J, Coelho ED, Amaral D, Correia MJ, Barros M, Oliveira JL. OralCard: a bioinformatic tool for the study of oral proteome. Arch Oral Biol. 2013;58(7):762–772. doi: 10.1016/j.archoralbio.2012.12.012. [DOI] [PubMed] [Google Scholar]
- Rosa N, Correia MJ, Arrais JP, Lopes P, Melo J, Oliveira JL, Barros M. From the salivary proteome to the OralOme: comprehensive molecular oral biology. Arch Oral Biol. 2012;57(7):853–864. doi: 10.1016/j.archoralbio.2011.12.010. [DOI] [PubMed] [Google Scholar]
- Vecchiola C, Pandey S, Buyya R. High-performance cloud computing: a view of scientific applications. 2009. pp. 4–16. Proceedings of the 10th International Symposium on Pervasive Systems, Algorithms and Networks I-SPAN 2009, IEEE Computer Society.
- Yamane K, Nambu T, Yamanaka T, Mashimo C, Sugimori C, Leung K-P, Fukushima H. Complete genome sequence of rothia mucilaginosa DY-18: a clinical isolate with dense meshwork-like structures from a persistent apical periodontitis lesion. Sequencing. 2010;2010:1–6. [Google Scholar]
- Batty I. Actinomyces odontolyticus, a new species of actinomycete regularly isolated from deep carious dentine. J Pathol Bacteriol. 1958;75:455–459. doi: 10.1002/path.1700750225. [DOI] [PubMed] [Google Scholar]
- McKay LI, Cidlowski JA. Molecular control of immune/inflammatory responses: interactions between nuclear factor-κB and steroid receptor-signaling pathways. Endocrine Rev. 1999;20:435–459. doi: 10.1210/edrv.20.4.0375. [DOI] [PubMed] [Google Scholar]
- McDevitt H, Munson S, Ettinger R, Wu A. Multiple roles for tumor necrosis factor-alpha and lymphotoxin alpha/beta in immunity and autoimmunity. Arthritis Res. 2002;4:S141–S152. doi: 10.1186/ar570. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barnard JA, Beauchamp RD, Russell WE, Dubois RN, Coffey RJ. Epidermal growth factor-related peptides and their relevance to gastrointestinal pathophysiology. Gastroenterology. 1995;108:564–580. doi: 10.1016/0016-5085(95)90087-X. [DOI] [PubMed] [Google Scholar]
- Galan JE, Pace J, Hayman MJ. Involvement of the epidermal growth factor receptor in the invasion of cultured mammalian cells by Salmonella typhimurium. Nature. 1992;357:588–589. doi: 10.1038/357588a0. [DOI] [PubMed] [Google Scholar]
- Zhu W, Phan QT, Boontheung P, Solis NV, Loo JA, Filler SG. EGFR and HER2 receptor kinase signaling mediate epithelial cell invasion by Candida albicans during oropharyngeal infection. Proc Natl Acad Sci U S A. 2012;109:14194–14199. doi: 10.1073/pnas.1117676109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Strong JE, Tang D, Lee PW. Evidence that the epidermal growth factor receptor on host cells confers reovirus infection efficiency. Virology. 1993;197:405–411. doi: 10.1006/viro.1993.1602. [DOI] [PubMed] [Google Scholar]
- Eppstein DA, Vivienne Marsh Y, Schreiber AB, Newman SR, Todaro GJ, Nestor JJ Jr. Epidermal growth factor receptor occupancy inhibits vaccinia virus infection. Nature. 1985;318:663–665. doi: 10.1038/318663a0. [DOI] [PubMed] [Google Scholar]
- Buret A, Gall DG, Olson ME, Hardin JA. The role of the epidermal growth factor receptor in microbial infections of the gastrointestinal tract. Microbes Infect. 1999;1:1139–1144. doi: 10.1016/S1286-4579(99)00201-4. [DOI] [PubMed] [Google Scholar]
- Llena-Puy MC, Montanana-Llorens C, Forner-Navarro L. Fibronectin levels in stimulated whole-saliva and their relationship with cariogenic oral bacteria. Int Dent J. 2000;50:57–59. doi: 10.1111/j.1875-595X.2000.tb00548.x. [DOI] [PubMed] [Google Scholar]
- Henderson B, Nair S, Pallas J, Williams MA. Fibronectin: a multidomain host adhesin targeted by bacterial fibronectin-binding proteins. FEMS Microbiol Rev. 2011;35:147–200. doi: 10.1111/j.1574-6976.2010.00243.x. [DOI] [PubMed] [Google Scholar]
- Min K-W, Hwang J-W, Lee J-S, Park Y, T-a T, Yoon J-B. TIP120A associates with cullins and modulates ubiquitin ligase activity. J. Biol. Chem. 2003;278:15905–15910. doi: 10.1074/jbc.M213070200. [DOI] [PubMed] [Google Scholar]
- Sarikas A, Hartmann T, Pan ZQ. The cullin protein family. Genome Biol. 2011;12:220. doi: 10.1186/gb-2011-12-4-220. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zheng J, Yang X, Harrell JM, Ryzhikov S, Shim E-H, Lykke-Andersen K, Wei N, Sun H, Kobayashi R, Zhang H. CAND1 binds to unneddylated CUL1 and regulates the formation of SCF ubiquitin E3 ligase complex. Mol Cell. 2002;10:1519–1526. doi: 10.1016/S1097-2765(02)00784-0. [DOI] [PubMed] [Google Scholar]
- Munro P, Flatau G, Lemichez E. Bacteria and the ubiquitin pathway. Curr Opin Microbiol. 2007;10:39–46. doi: 10.1016/j.mib.2006.11.002. [DOI] [PubMed] [Google Scholar]
- Curtis H, Dirk G, Rob K, Sahar A, Badger JH, Chinwalla AT, Creasy HH, Earl AM, FitzGerald MG, Fulton RS, Giglio MG, Kymberlie H-P, Lobos EA, Ramana M, Vincent M, Martin JC, Makedonka M, Muzny DM, Sodergren EJ, Versalovic J, Wollam AM, Worley KC, Wortman JR, Young SK, Qiandong Z, Aagaard KM, Abolude OO, Emma A-V, Alm EJ, Lucia A. et al. Structure, function and diversity of the healthy human microbiome. Nature. 2012;486:207–214. doi: 10.1038/nature11234. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Avila-Campos MJ, Velasquez-Melendez G. Prevalence of putative periodontopathogens from periodontal patients and healthy subjects in Sao Paulo, SP, Brazil. Rev Inst Med Trop Sao Paulo. 2002;44:1–5. doi: 10.1590/s0036-46652002000100001. [DOI] [PubMed] [Google Scholar]
- Antikainen J, Kuparinen V, Lahteenmaki K, Korhonen TK. Enolases from Gram-positive bacterial pathogens and commensal lactobacilli share functional similarity in virulence-associated traits. FEMS Immunol Med Microbiol. 2007;51:526–534. doi: 10.1111/j.1574-695X.2007.00330.x. [DOI] [PubMed] [Google Scholar]
- Levy ED, Pereira-Leal JB. Evolution and dynamics of protein interactions and networks. Curr Opin Struct Biol. 2008;18:349–357. doi: 10.1016/j.sbi.2008.03.003. [DOI] [PubMed] [Google Scholar]
- Stark C, Breitkreutz B-J, Reguly T, Boucher L, Breitkreutz A, Tyers M. BioGRID: a general repository for interaction datasets. Nucleic Acids Res. 2006;34:D535–539. doi: 10.1093/nar/gkj109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Salwinski L, Miller CS, Smith AJ, Pettit FK, Bowie JU, Eisenberg D. The database of interacting proteins: 2004 update. Nucleic Acids Res. 2004;32:D449–D451. doi: 10.1093/nar/gkh086. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Keshava Prasad TS, Goel R, Kandasamy K, Keerthikumar S, Kumar S, Mathivanan S, Telikicherla D, Raju R, Shafreen B, Venugopal A, Balakrishnan L, Marimuthu A, Banerjee S, Somanathan DS, Sebastian A, Rani S, Ray S, Harrys Kishore CJ, Kanth S, Ahmed M, Kashyap MK, Mohmood R, Ramachandra YL, Krishna V, Rahiman BA, Mohan S, Ranganathan P, Ramabadran S, Chaerkady R, Pandey A. Human protein reference database–2009 update. Nucleic Acids Res. 2009;37:D767–772. doi: 10.1093/nar/gkn892. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kerrien S, Aranda B, Breuza L, Bridge A, Broackes-Carter F, Chen C, Duesbury M, Dumousseau M, Feuermann M, Hinz U, Jandrasits C, Jimenez RC, Khadake J, Mahadevan U, Masson P, Pedruzzi I, Pfeiffenberger E, Porras P, Raghunath A, Roechert B, Orchard S, Hermjakob H. The IntAct molecular interaction database in 2012. Nucleic Acids Res. 2012;40:D841–D846. doi: 10.1093/nar/gkr1088. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chatr-aryamontri A, Ceol A, Montecchi Palazzi L, Nardelli G, Schneider MV, Castagnoli L, Cesareni G. MINT, the molecular interaction database: 2012 update. Nucleic Acids Res. 2012;40:D857–861. doi: 10.1093/nar/gkr930. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Consortium TU. Reorganizing the protein space at the Universal protein resource (UniProt) Nucleic Acids Res. 2012;40:D71–D75. doi: 10.1093/nar/gkr981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ben-Hur A, Noble WS. Choosing negative examples for the prediction of protein-protein interactions. BMC Bioinformatics. 2006;7(Suppl 1):S2. doi: 10.1186/1471-2105-7-S1-S2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van Haagen HHHBM, Hoen PAC't, Botelho Bovo A, de Morrée A, van Mulligen EM, Chichester C, Kors JA, den Dunnen JT, van Ommen G-JB, van der Maarel SM, Medina Kern V, Mons B, Schuemie MJ. Novel protein-protein interactions inferred from literature context. PLoS One. 2009;4:e7894. doi: 10.1371/journal.pone.0007894. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jelier R, Schuemie MJ, Roes PJ, van Mulligen EM, Kors JA. Literature-based concept profiles for gene annotation: the issue of weighting. Int J Med Inform. 2008;77:354–362. doi: 10.1016/j.ijmedinf.2007.07.004. [DOI] [PubMed] [Google Scholar]
- Campos D, Matos S, Oliveira J. Gimli: open source and high-performance biomedical name recognition. BMC Bioinformatics. 2013;14:54. doi: 10.1186/1471-2105-14-54. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tatusov RL, Koonin EV, Lipman DJ. A genomic perspective on protein families. Science. 1997;278:631–637. doi: 10.1126/science.278.5338.631. [DOI] [PubMed] [Google Scholar]
- Lee S-A, C-h C, Tsai C-H, Lai J-M, Wang F-S, Kao C-Y, Huang C-YF. Ortholog-based protein-protein interaction prediction and its application to inter-species interactions. BMC Bioinformatics. 2008;9(Suppl 12):S11. doi: 10.1186/1471-2105-9-S12-S11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Szklarczyk D, Franceschini A, Kuhn M, Simonovic M, Roth A, Minguez P, Doerks T, Stark M, Muller J, Bork P, Jensen LJ, von Mering C. The STRING database in 2011: functional interaction networks of proteins, globally integrated and scored. Nucleic Acids Res. 2011;39:D561–568. doi: 10.1093/nar/gkq973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lin N, Wu B, Jansen R, Gerstein M, Zhao H. Information assessment on predicting protein-protein interactions. BMC Bioinformatics. 2004;5:154. doi: 10.1186/1471-2105-5-154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miller JP, Lo RS, Ben-Hur A, Desmarais C, Stagljar I, Noble WS, Fields S. Large-scale identification of yeast integral membrane protein interactions. Proc Natl Acad Sci U S A. 2005;102:12123–12128. doi: 10.1073/pnas.0505482102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Patil A, Nakamura H. Filtering high-throughput protein-protein interaction data using a combination of genomic features. BMC Bioinformatics. 2005;6:100. doi: 10.1186/1471-2105-6-100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Qi Y, Bar-Joseph Z, Klein-Seetharaman J. Evaluation of different biological data and computational classification methods for use in protein interaction prediction. Proteins. 2006;63:490–500. doi: 10.1002/prot.20865. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yellaboina S, Tasneem A, Zaykin DV, Raghavachari B, Jothi R. DOMINE: a comprehensive collection of known and predicted domain-domain interactions. Nucleic Acids Res. 2011;39:D730–D735. doi: 10.1093/nar/gkq1229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Duda R, Hart P. Pattern Classification and Scene Analysis. New York: John Wiley & Sons Inc; 1973. [Google Scholar]
- Friedman N, Geiger D, Goldszmidt M. Bayesian Network Classifiers. Mach Learn. 1997;29:131–163. doi: 10.1023/A:1007465528199. [DOI] [Google Scholar]
- Swets JA. Measuring the accuracy of diagnostic systems. Science. 1988;240:1285–1293. doi: 10.1126/science.3287615. [DOI] [PubMed] [Google Scholar]
- Hanley JA, McNeil BJ. The meaning and use of the area under a receiver operating characteristic (ROC) curve. Radiology. 1982;143:29–36. doi: 10.1148/radiology.143.1.7063747. [DOI] [PubMed] [Google Scholar]