Figure 1.
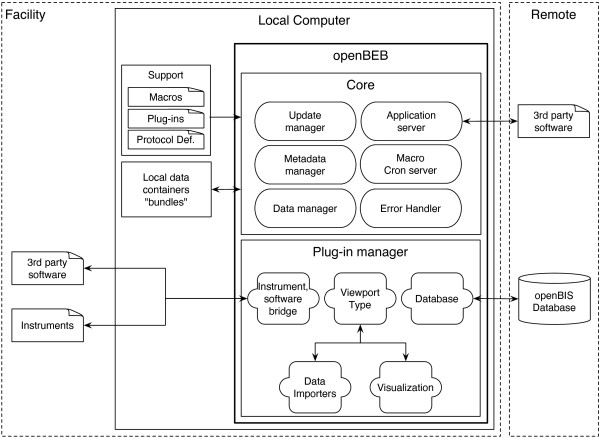
Overview of the openBEB software. For end-users, openBEB acts as a link between experimental biology and database systems. It is designed to facilitate data acquisition, browsing, annotation and synchronization with databases. For developers, it provides an environment for the fast development of data-acquisition and instrument control plug-ins, providing a core framework for data-management and coordination of different modules by a unified macro language. OpenBEB runs locally and consists of a core program and a plug-in manager. Plug-ins are dynamically loaded during start-up. The core program maintains a local bundle, i.e., a container structure in the file system for raw-data, metadata and cache files. All files (except the cache files) are readable by standard programs, can be browsed in the file system, and can be directly accessed by other local programs. The core program can be controlled by a graphical user interface (GUI) and via macros and a TCP-based protocol. The update manager of openBEB automatically updates an application support folder containing metadata definition files and plug-ins. Plug-ins are loaded and coordinated by the plug-in manager. The open architecture of openBEB allows plug-ins to be written to handle different types of data (data-type plug-ins, e.g., for images or simple xy-scatter data). Data-importers or plug-ins for data visualization (called viewports) can be loaded. The plug-in manager also provides a unified interface for instrument control plug-ins facilitating the rapid development of instrument control software. Database synchronization is also achieved using plug-ins; the standard openBEB installation is designed to communicate with the openBIS system [6]. The error handler provides a logging and error-processing tool for the core program as well as for the plug-ins.
