Figure 5.
Enrichment of Activation Modules in Human Alveolar Macrophages
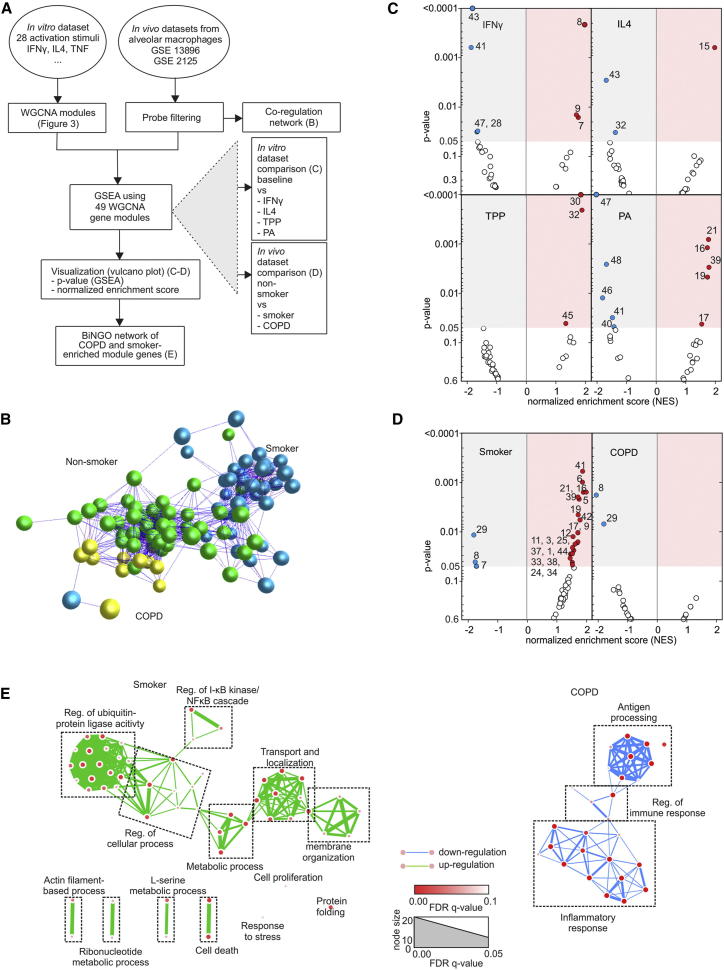
(A) Schema describing the workflow for Figure 5.
(B) Correlation network of human alveolar macrophages (n = 100) from two studies (Shaykhiev et al., 2009; Woodruff et al., 2005) using 374 genes differentially expressed between nonsmokers (n = 39) and smokers (n = 49) or COPD (n = 12) patients (FC > 2.0, FDR adjusted p value < 0.05).
(C) Volcano plots of normalized enrichment scores (NES) and enrichment p values based on GSEA using WGCNA modules defined in Figure 3. Shown are data for the stimuli IFN-γ, IL-4, TPP, and palmitic acid (PA). Red circles show gene sets positively significantly enriched (NES > 1, p value < 0.05); blue circles show gene sets significantly depleted (NES < −1, p value < 0.05).
(D) Volcano plots of normalized enrichment scores (NES) and enrichment p values of the same gene sets applied to data from alveolar macrophages derived from smokers and COPD patients. Representative results of several permutation runs of GSEA.
(E) Network visualization of GOEA of positively enriched modules (p value < 0.01) for smokers (modules 41, 6, 21, 16, 39, 5, 19, and 42) and negatively enriched modules (p value < 0.01) for COPD patients (modules 8, 29) using BiNGO and EnrichmentMap. Red nodes represent enriched GO-terms, node size, and color represent corresponding FDR-adjusted enrichment p values (q values).