Abstract
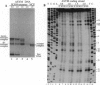
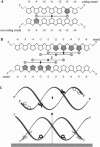
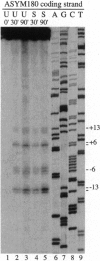
A base-pair resolution method for determining nucleosome position in vitro has been developed to com- plement existing, less accurate methods. Cysteaminyl EDTA was tethered to a recombinant histone octamer via a mutant histone H4 with serine 47 replaced by cysteine. When assembled into nucleosome core particles, the DNA could be cut site specifically by hydroxyl radical-catalyzed chain scission by using the Fenton reaction. Strand cleavage occurs mainly at a single nucleotide close to the dyad axis of the core particle, and assignment of this location via the symmetry of the nucleosome allows base-pair resolution mapping of the histone octamer position on the DNA. The positions of the histone octamer and H3H4 tetramer were mapped on a 146-bp Lytechinus variegatus 5S rRNA sequence and a twofold-symmetric derivative. The weakness of translational determinants of nucleosome positioning relative to the overall affinity of the histone proteins for this DNA is clearly demonstrated. The predominant location of both histone octamer and H3H4 tetramer assembled on the 5S rDNA is off center. Shifting the nucleosome core particle position along DNA within a conserved rotational phase could be induced under physiologically relevant conditions. Since nucleosome shifting has important consequences for chromatin structure and gene regulation, an approach to the thermodynamic characterization of this movement is proposed. This mapping method is potentially adaptable for determining nucleosome position in chromatin in vivo.
Full text
PDF




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Archer T. K., Cordingley M. G., Wolford R. G., Hager G. L. Transcription factor access is mediated by accurately positioned nucleosomes on the mouse mammary tumor virus promoter. Mol Cell Biol. 1991 Feb;11(2):688–698. doi: 10.1128/mcb.11.2.688. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Archer T. K., Lefebvre P., Wolford R. G., Hager G. L. Transcription factor loading on the MMTV promoter: a bimodal mechanism for promoter activation. Science. 1992 Mar 20;255(5051):1573–1576. doi: 10.1126/science.1347958. [DOI] [PubMed] [Google Scholar]
- Arents G., Moudrianakis E. N. Topography of the histone octamer surface: repeating structural motifs utilized in the docking of nucleosomal DNA. Proc Natl Acad Sci U S A. 1993 Nov 15;90(22):10489–10493. doi: 10.1073/pnas.90.22.10489. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cooper J. P., Roth S. Y., Simpson R. T. The global transcriptional regulators, SSN6 and TUP1, play distinct roles in the establishment of a repressive chromatin structure. Genes Dev. 1994 Jun 15;8(12):1400–1410. doi: 10.1101/gad.8.12.1400. [DOI] [PubMed] [Google Scholar]
- Dong F., Hansen J. C., van Holde K. E. DNA and protein determinants of nucleosome positioning on sea urchin 5S rRNA gene sequences in vitro. Proc Natl Acad Sci U S A. 1990 Aug;87(15):5724–5728. doi: 10.1073/pnas.87.15.5724. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dong F., van Holde K. E. Nucleosome positioning is determined by the (H3-H4)2 tetramer. Proc Natl Acad Sci U S A. 1991 Dec 1;88(23):10596–10600. doi: 10.1073/pnas.88.23.10596. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Drew H. R. Can one measure the free energy of binding of the histone octamer to different DNA sequences by salt-dependent reconstitution? J Mol Biol. 1991 Jun 5;219(3):391–392. doi: 10.1016/0022-2836(91)90179-a. [DOI] [PubMed] [Google Scholar]
- Germond J. E., Bellard M., Oudet P., Chambon P. Stability of nucleosomes in native and reconstituted chromatins. Nucleic Acids Res. 1976 Nov;3(11):3173–3192. doi: 10.1093/nar/3.11.3173. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hansen J. C., van Holde K. E., Lohr D. The mechanism of nucleosome assembly onto oligomers of the sea urchin 5 S DNA positioning sequence. J Biol Chem. 1991 Mar 5;266(7):4276–4282. [PubMed] [Google Scholar]
- Hayes J. J., Bashkin J., Tullius T. D., Wolffe A. P. The histone core exerts a dominant constraint on the structure of DNA in a nucleosome. Biochemistry. 1991 Aug 27;30(34):8434–8440. doi: 10.1021/bi00098a022. [DOI] [PubMed] [Google Scholar]
- Hayes J. J., Clark D. J., Wolffe A. P. Histone contributions to the structure of DNA in the nucleosome. Proc Natl Acad Sci U S A. 1991 Aug 1;88(15):6829–6833. doi: 10.1073/pnas.88.15.6829. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee D. Y., Hayes J. J., Pruss D., Wolffe A. P. A positive role for histone acetylation in transcription factor access to nucleosomal DNA. Cell. 1993 Jan 15;72(1):73–84. doi: 10.1016/0092-8674(93)90051-q. [DOI] [PubMed] [Google Scholar]
- Lennox R. W., Cohen L. H. Analysis of histone subtypes and their modified forms by polyacrylamide gel electrophoresis. Methods Enzymol. 1989;170:532–549. doi: 10.1016/0076-6879(89)70063-x. [DOI] [PubMed] [Google Scholar]
- Lu A. L., Steege D. A., Stafford D. W. Nucleotide sequence of a 5S ribosomal RNA gene in the sea urchin Lytechinus variegatus. Nucleic Acids Res. 1980 Apr 25;8(8):1839–1853. doi: 10.1093/nar/8.8.1839. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meersseman G., Pennings S., Bradbury E. M. Chromatosome positioning on assembled long chromatin. Linker histones affect nucleosome placement on 5 S rDNA. J Mol Biol. 1991 Jul 5;220(1):89–100. doi: 10.1016/0022-2836(91)90383-h. [DOI] [PubMed] [Google Scholar]
- Meersseman G., Pennings S., Bradbury E. M. Mobile nucleosomes--a general behavior. EMBO J. 1992 Aug;11(8):2951–2959. doi: 10.1002/j.1460-2075.1992.tb05365.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Piña B., Barettino D., Truss M., Beato M. Structural features of a regulatory nucleosome. J Mol Biol. 1990 Dec 20;216(4):975–990. doi: 10.1016/S0022-2836(99)80015-1. [DOI] [PubMed] [Google Scholar]
- Record M. T., Jr, Ha J. H., Fisher M. A. Analysis of equilibrium and kinetic measurements to determine thermodynamic origins of stability and specificity and mechanism of formation of site-specific complexes between proteins and helical DNA. Methods Enzymol. 1991;208:291–343. doi: 10.1016/0076-6879(91)08018-d. [DOI] [PubMed] [Google Scholar]
- Richard-Foy H., Hager G. L. Sequence-specific positioning of nucleosomes over the steroid-inducible MMTV promoter. EMBO J. 1987 Aug;6(8):2321–2328. doi: 10.1002/j.1460-2075.1987.tb02507.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Richmond T. J., Finch J. T., Rushton B., Rhodes D., Klug A. Structure of the nucleosome core particle at 7 A resolution. Nature. 1984 Oct 11;311(5986):532–537. doi: 10.1038/311532a0. [DOI] [PubMed] [Google Scholar]
- Richmond T. J., Rechsteiner T., Luger K. Studies of nucleosome structure. Cold Spring Harb Symp Quant Biol. 1993;58:265–272. doi: 10.1101/sqb.1993.058.01.031. [DOI] [PubMed] [Google Scholar]
- Richmond T. J., Searles M. A., Simpson R. T. Crystals of a nucleosome core particle containing defined sequence DNA. J Mol Biol. 1988 Jan 5;199(1):161–170. doi: 10.1016/0022-2836(88)90386-5. [DOI] [PubMed] [Google Scholar]
- Roth S. Y., Shimizu M., Johnson L., Grunstein M., Simpson R. T. Stable nucleosome positioning and complete repression by the yeast alpha 2 repressor are disrupted by amino-terminal mutations in histone H4. Genes Dev. 1992 Mar;6(3):411–425. doi: 10.1101/gad.6.3.411. [DOI] [PubMed] [Google Scholar]
- Satchwell S. C., Drew H. R., Travers A. A. Sequence periodicities in chicken nucleosome core DNA. J Mol Biol. 1986 Oct 20;191(4):659–675. doi: 10.1016/0022-2836(86)90452-3. [DOI] [PubMed] [Google Scholar]
- Schaich K. M. Preparation of metal-free solutions for studies of active oxygen species. Methods Enzymol. 1990;186:121–125. doi: 10.1016/0076-6879(90)86099-h. [DOI] [PubMed] [Google Scholar]
- Shrader T. E., Crothers D. M. Artificial nucleosome positioning sequences. Proc Natl Acad Sci U S A. 1989 Oct;86(19):7418–7422. doi: 10.1073/pnas.86.19.7418. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Simpson R. T., Stafford D. W. Structural features of a phased nucleosome core particle. Proc Natl Acad Sci U S A. 1983 Jan;80(1):51–55. doi: 10.1073/pnas.80.1.51. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Studitsky V. M., Clark D. J., Felsenfeld G. A histone octamer can step around a transcribing polymerase without leaving the template. Cell. 1994 Jan 28;76(2):371–382. doi: 10.1016/0092-8674(94)90343-3. [DOI] [PubMed] [Google Scholar]
- Thoma F. Nucleosome positioning. Biochim Biophys Acta. 1992 Feb 28;1130(1):1–19. doi: 10.1016/0167-4781(92)90455-9. [DOI] [PubMed] [Google Scholar]
- Thoma F., Simpson R. T. Local protein-DNA interactions may determine nucleosome positions on yeast plasmids. Nature. 1985 May 16;315(6016):250–252. doi: 10.1038/315250a0. [DOI] [PubMed] [Google Scholar]