Figure 10.
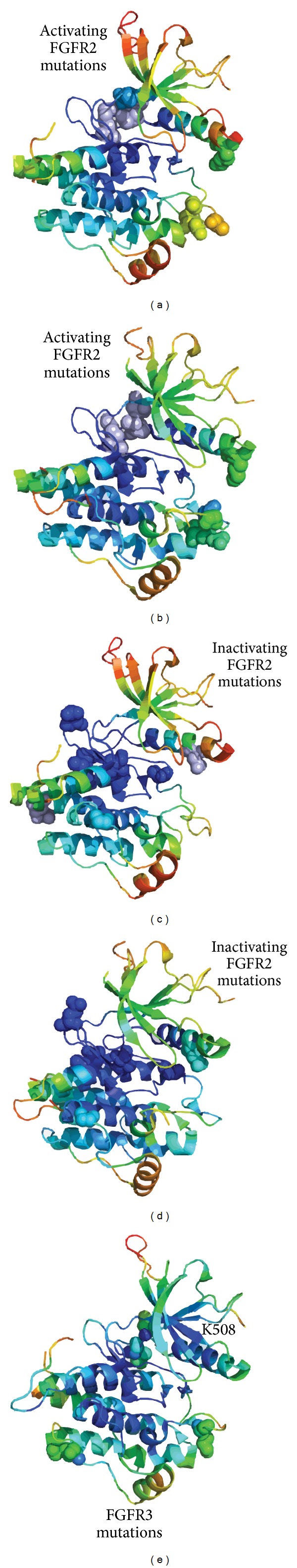
Functional dynamics and structural map of the FGFR2 and FGFR3 cancer mutations. Functional dynamics and conformational mobility map of the EGFR2 activating mutations (I548V, N550K, E565G, N549H, N540K, K526E, K660E, R678G, K641R, and G663E) projected onto the unphosphorylated FGFR2 crystal structure, pdb id 2PSQ (a) and phosphorylated active FGFR2 crystal structure, pdb id 2PVF (b). Dynamic maps of the EGFR2 inactivating mutations (E636K, M640I, I642V, R759Q, E475K, D530N, A648T, and G701S) projected onto the unphosphorylated FGFR2 crystal structure (c) and the phosphorylated active FGFR2 crystal structure (d). Dynamic map of the FGFR3 activating mutations (I538V, N540K, and K650E) and inactivating mutation (K508) projected onto the crystal structure of FGFR3 (pdb id 4k33) (e). A surface-based protein representation is employed, colored (blue-to-red) according to the protein residue motilities (from more rigid-blue regions to more flexible-red regions).
