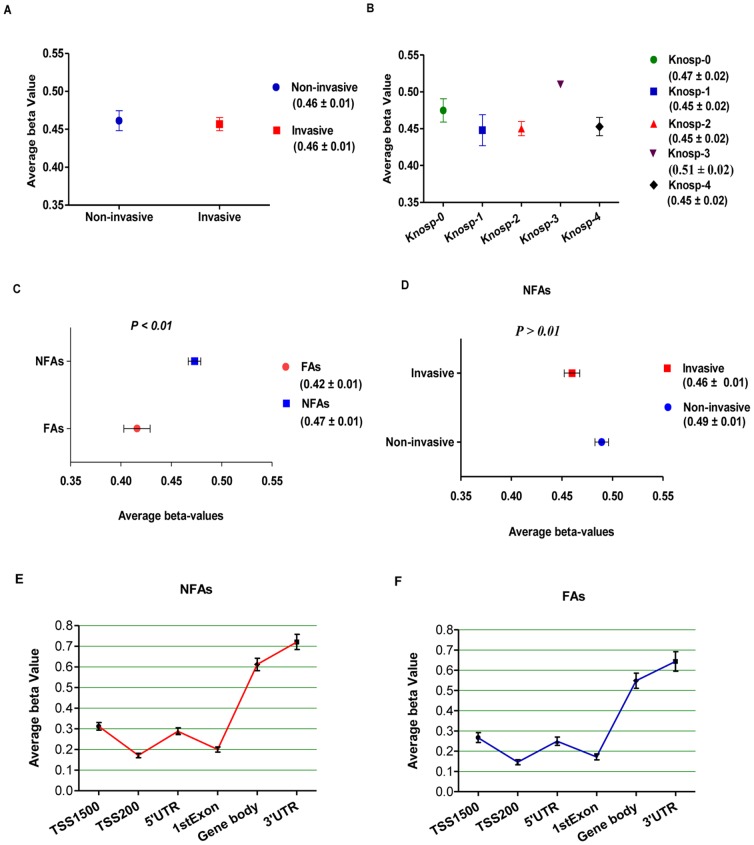
Figure 2. Global CpG DNA methylation profiles in PAs.
A, Comparison of mean DNA methylation levels across 383,718 HM450 probes in noninvasive (Knosp grade 0, 1) and invasive (Knosp grade 2, 3, 4) PAs. B, Mean DNA methylation levels in PA cases across the five different Knosp grades (0–4) among the set of 383,718 HM450 probes. C, Comparison of mean DNA methylation levels across 383,718 HM450 probes in FAs and NFAs. P-value was calculated via simple t-test. D, Comparison of mean DNA methylation levels across 383,718 HM450 probes in only NFAs. P-value was calculated via simple t-test. All of the above DNA methylation beta values are shown as mean ± SEM in different groups. E and F, DNA methylation levels in gene regions (TSS1500, TSS200, 5′UTR, 1stExon, gene body and 3′UTR) for NFAs (panel E) and FAs (Panel F). TSS200: 200 bp within transcription start site; TSS1500: 1500 bp within transcription start site; UTR: untranslated region; 1stExon: the first exon of gene.