Abstract
To further our studies of protein sorting and biogenesis of the lysosome-like vacuole in yeast, we have isolated spontaneous mutations in 11 new VPL complementation groups, as well as additional alleles of the eight previously described VPL genes. These mutants were identified by selecting for cells that mislocalize vacuolar proteins to the cell surface. Morphological examination of the vpl mutants indicated that most contain vacuoles of normal appearance; however, some of the mutants generally lack a large vacuole, and instead accumulate smaller organelles. Of the 19 VPL complementation groups, 12 were found to be identical to 12 of 33 VPT complementation groups identified in a separate study. Moreover, the end1 mutant and all of the previously reported pep mutants, with the exception of pep4, were found to exhibit a profound vacuolar protein sorting defect, and complementation tests between the PEP, VPL VPT and END1 groups demonstrated that there are extensive overlaps between these groups. Collectively, mutants in these four collections define 49 complementation groups required to deliver or retain soluble vacuolar enzymes, including carboxypeptidase Y (CPY) and proteinase A. We have also isolated 462 new mutants that lack normal levels of vacuolar CPY activity. Among these latter mutants, only pep4 mutants were found to be specifically defective in vacuolar zymogen activation. We conclude that there is a large number of gene products required for sorting or retention of vacuolar proteins in yeast, and only a single gene, PEP4, that is essential for activation of CPY and other vacuolar zymogens.
Full text
PDF
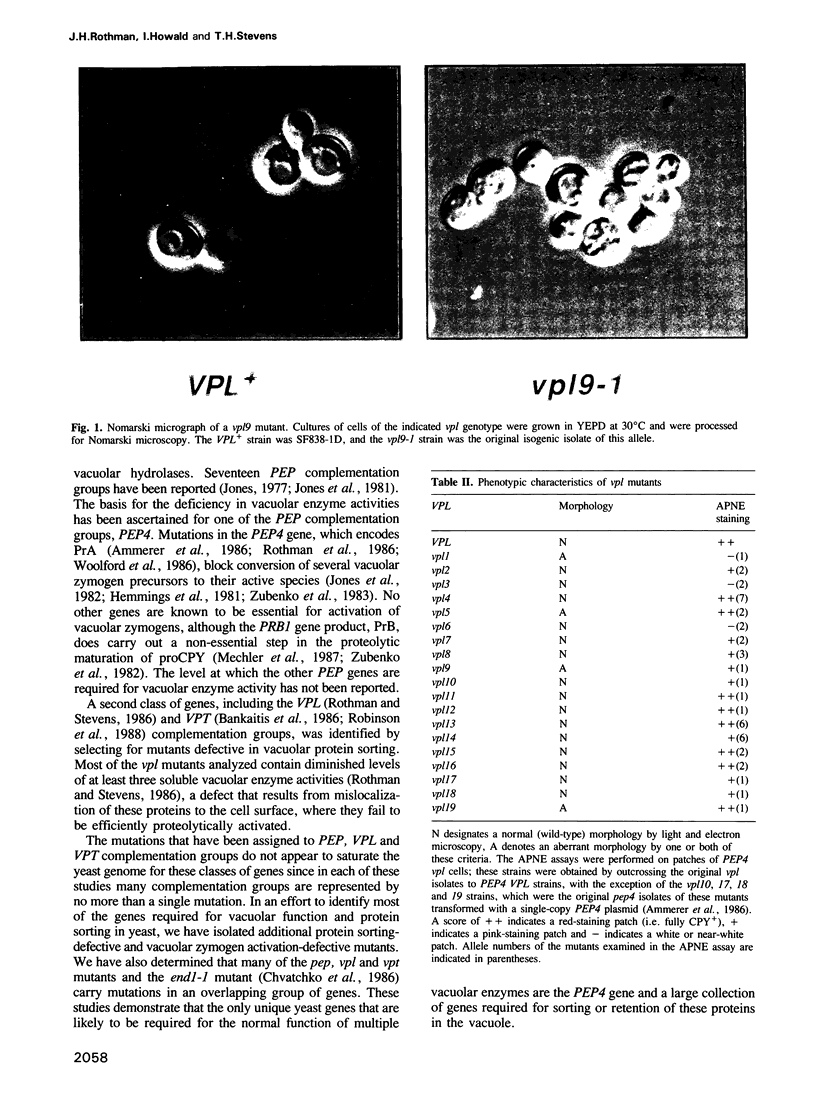







Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Ammerer G., Hunter C. P., Rothman J. H., Saari G. C., Valls L. A., Stevens T. H. PEP4 gene of Saccharomyces cerevisiae encodes proteinase A, a vacuolar enzyme required for processing of vacuolar precursors. Mol Cell Biol. 1986 Jul;6(7):2490–2499. doi: 10.1128/mcb.6.7.2490. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bankaitis V. A., Johnson L. M., Emr S. D. Isolation of yeast mutants defective in protein targeting to the vacuole. Proc Natl Acad Sci U S A. 1986 Dec;83(23):9075–9079. doi: 10.1073/pnas.83.23.9075. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Banta L. M., Robinson J. S., Klionsky D. J., Emr S. D. Organelle assembly in yeast: characterization of yeast mutants defective in vacuolar biogenesis and protein sorting. J Cell Biol. 1988 Oct;107(4):1369–1383. doi: 10.1083/jcb.107.4.1369. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brown W. J., Goodhouse J., Farquhar M. G. Mannose-6-phosphate receptors for lysosomal enzymes cycle between the Golgi complex and endosomes. J Cell Biol. 1986 Oct;103(4):1235–1247. doi: 10.1083/jcb.103.4.1235. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Byers B., Goetsch L. Behavior of spindles and spindle plaques in the cell cycle and conjugation of Saccharomyces cerevisiae. J Bacteriol. 1975 Oct;124(1):511–523. doi: 10.1128/jb.124.1.511-523.1975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Böhni P. C., Deshaies R. J., Schekman R. W. SEC11 is required for signal peptide processing and yeast cell growth. J Cell Biol. 1988 Apr;106(4):1035–1042. doi: 10.1083/jcb.106.4.1035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chamberlain J. P. Fluorographic detection of radioactivity in polyacrylamide gels with the water-soluble fluor, sodium salicylate. Anal Biochem. 1979 Sep 15;98(1):132–135. doi: 10.1016/0003-2697(79)90716-4. [DOI] [PubMed] [Google Scholar]
- Chvatchko Y., Howald I., Riezman H. Two yeast mutants defective in endocytosis are defective in pheromone response. Cell. 1986 Aug 1;46(3):355–364. doi: 10.1016/0092-8674(86)90656-2. [DOI] [PubMed] [Google Scholar]
- Deshaies R. J., Schekman R. A yeast mutant defective at an early stage in import of secretory protein precursors into the endoplasmic reticulum. J Cell Biol. 1987 Aug;105(2):633–645. doi: 10.1083/jcb.105.2.633. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Geuze H. J., Slot J. W., Strous G. J., Hasilik A., von Figura K. Possible pathways for lysosomal enzyme delivery. J Cell Biol. 1985 Dec;101(6):2253–2262. doi: 10.1083/jcb.101.6.2253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Griffiths G., Hoflack B., Simons K., Mellman I., Kornfeld S. The mannose 6-phosphate receptor and the biogenesis of lysosomes. Cell. 1988 Feb 12;52(3):329–341. doi: 10.1016/s0092-8674(88)80026-6. [DOI] [PubMed] [Google Scholar]
- Hasilik A., Tanner W. Biosynthesis of the vacuolar yeast glycoprotein carboxypeptidase Y. Conversion of precursor into the enzyme. Eur J Biochem. 1978 Apr 17;85(2):599–608. doi: 10.1111/j.1432-1033.1978.tb12275.x. [DOI] [PubMed] [Google Scholar]
- Hemmings B. A., Zubenko G. S., Hasilik A., Jones E. W. Mutant defective in processing of an enzyme located in the lysosome-like vacuole of Saccharomyces cerevisiae. Proc Natl Acad Sci U S A. 1981 Jan;78(1):435–439. doi: 10.1073/pnas.78.1.435. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jenness D. D., Spatrick P. Down regulation of the alpha-factor pheromone receptor in S. cerevisiae. Cell. 1986 Aug 1;46(3):345–353. doi: 10.1016/0092-8674(86)90655-0. [DOI] [PubMed] [Google Scholar]
- Johnson L. M., Bankaitis V. A., Emr S. D. Distinct sequence determinants direct intracellular sorting and modification of a yeast vacuolar protease. Cell. 1987 Mar 13;48(5):875–885. doi: 10.1016/0092-8674(87)90084-5. [DOI] [PubMed] [Google Scholar]
- Jones E. W. Proteinase mutants of Saccharomyces cerevisiae. Genetics. 1977 Jan;85(1):23–33. doi: 10.1093/genetics/85.1.23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jones E. W. The synthesis and function of proteases in Saccharomyces: genetic approaches. Annu Rev Genet. 1984;18:233–270. doi: 10.1146/annurev.ge.18.120184.001313. [DOI] [PubMed] [Google Scholar]
- Jones E. W., Zubenko G. S., Parker R. R. PEP4 gene function is required for expression of several vacuolar hydrolases in Saccharomyces cerevisiae. Genetics. 1982 Dec;102(4):665–677. doi: 10.1093/genetics/102.4.665. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klionsky D. J., Banta L. M., Emr S. D. Intracellular sorting and processing of a yeast vacuolar hydrolase: proteinase A propeptide contains vacuolar targeting information. Mol Cell Biol. 1988 May;8(5):2105–2116. doi: 10.1128/mcb.8.5.2105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kuranda M. J., Robbins P. W. Cloning and heterologous expression of glycosidase genes from Saccharomyces cerevisiae. Proc Natl Acad Sci U S A. 1987 May;84(9):2585–2589. doi: 10.1073/pnas.84.9.2585. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lippincott-Schwartz J., Fambrough D. M. Cycling of the integral membrane glycoprotein, LEP100, between plasma membrane and lysosomes: kinetic and morphological analysis. Cell. 1987 Jun 5;49(5):669–677. doi: 10.1016/0092-8674(87)90543-5. [DOI] [PubMed] [Google Scholar]
- Lippincott-Schwartz J., Fambrough D. M. Lysosomal membrane dynamics: structure and interorganellar movement of a major lysosomal membrane glycoprotein. J Cell Biol. 1986 May;102(5):1593–1605. doi: 10.1083/jcb.102.5.1593. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Malhotra V., Orci L., Glick B. S., Block M. R., Rothman J. E. Role of an N-ethylmaleimide-sensitive transport component in promoting fusion of transport vesicles with cisternae of the Golgi stack. Cell. 1988 Jul 15;54(2):221–227. doi: 10.1016/0092-8674(88)90554-5. [DOI] [PubMed] [Google Scholar]
- Mechler B., Müller H., Wolf D. H. Maturation of vacuolar (lysosomal) enzymes in yeast: proteinase yscA and proteinase yscB are catalysts of the processing and activation event of carboxypeptidase yscY. EMBO J. 1987 Jul;6(7):2157–2163. doi: 10.1002/j.1460-2075.1987.tb02483.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mechler B., Müller M., Müller H., Meussdoerffer F., Wolf D. H. In vivo biosynthesis of the vacuolar proteinases A and B in the yeast Saccharomyces cerevisiae. J Biol Chem. 1982 Oct 10;257(19):11203–11206. [PubMed] [Google Scholar]
- Moehle C. M., Aynardi M. W., Kolodny M. R., Park F. J., Jones E. W. Protease B of Saccharomyces cerevisiae: isolation and regulation of the PRB1 structural gene. Genetics. 1987 Feb;115(2):255–263. doi: 10.1093/genetics/115.2.255. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moehle C. M., Tizard R., Lemmon S. K., Smart J., Jones E. W. Protease B of the lysosomelike vacuole of the yeast Saccharomyces cerevisiae is homologous to the subtilisin family of serine proteases. Mol Cell Biol. 1987 Dec;7(12):4390–4399. doi: 10.1128/mcb.7.12.4390. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pfeffer S. R., Rothman J. E. Biosynthetic protein transport and sorting by the endoplasmic reticulum and Golgi. Annu Rev Biochem. 1987;56:829–852. doi: 10.1146/annurev.bi.56.070187.004145. [DOI] [PubMed] [Google Scholar]
- Riezman H. Endocytosis in yeast: several of the yeast secretory mutants are defective in endocytosis. Cell. 1985 Apr;40(4):1001–1009. doi: 10.1016/0092-8674(85)90360-5. [DOI] [PubMed] [Google Scholar]
- Rine J., Strathern J. N., Hicks J. B., Herskowitz I. A suppressor of mating-type locus mutations in Saccharomyces cerevisiae: evidence for and identification of cryptic mating-type loci. Genetics. 1979 Dec;93(4):877–901. doi: 10.1093/genetics/93.4.877. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Robinson J. S., Klionsky D. J., Banta L. M., Emr S. D. Protein sorting in Saccharomyces cerevisiae: isolation of mutants defective in the delivery and processing of multiple vacuolar hydrolases. Mol Cell Biol. 1988 Nov;8(11):4936–4948. doi: 10.1128/mcb.8.11.4936. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rothman J. H., Hunter C. P., Valls L. A., Stevens T. H. Overproduction-induced mislocalization of a yeast vacuolar protein allows isolation of its structural gene. Proc Natl Acad Sci U S A. 1986 May;83(10):3248–3252. doi: 10.1073/pnas.83.10.3248. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rothman J. H., Stevens T. H. Protein sorting in yeast: mutants defective in vacuole biogenesis mislocalize vacuolar proteins into the late secretory pathway. Cell. 1986 Dec 26;47(6):1041–1051. doi: 10.1016/0092-8674(86)90819-6. [DOI] [PubMed] [Google Scholar]
- Salminen A., Novick P. J. A ras-like protein is required for a post-Golgi event in yeast secretion. Cell. 1987 May 22;49(4):527–538. doi: 10.1016/0092-8674(87)90455-7. [DOI] [PubMed] [Google Scholar]
- Schekman R. Protein localization and membrane traffic in yeast. Annu Rev Cell Biol. 1985;1:115–143. doi: 10.1146/annurev.cb.01.110185.000555. [DOI] [PubMed] [Google Scholar]
- Scott J. H., Schekman R. Lyticase: endoglucanase and protease activities that act together in yeast cell lysis. J Bacteriol. 1980 May;142(2):414–423. doi: 10.1128/jb.142.2.414-423.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stearns T., Botstein D. Unlinked noncomplementation: isolation of new conditional-lethal mutations in each of the tubulin genes of Saccharomyces cerevisiae. Genetics. 1988 Jun;119(2):249–260. doi: 10.1093/genetics/119.2.249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stevens T. H., Rothman J. H., Payne G. S., Schekman R. Gene dosage-dependent secretion of yeast vacuolar carboxypeptidase Y. J Cell Biol. 1986 May;102(5):1551–1557. doi: 10.1083/jcb.102.5.1551. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stevens T., Esmon B., Schekman R. Early stages in the yeast secretory pathway are required for transport of carboxypeptidase Y to the vacuole. Cell. 1982 Sep;30(2):439–448. doi: 10.1016/0092-8674(82)90241-0. [DOI] [PubMed] [Google Scholar]
- Uchida E., Ohsumi Y., Anraku Y. Purification and properties of H+-translocating, Mg2+-adenosine triphosphatase from vacuolar membranes of Saccharomyces cerevisiae. J Biol Chem. 1985 Jan 25;260(2):1090–1095. [PubMed] [Google Scholar]
- Valls L. A., Hunter C. P., Rothman J. H., Stevens T. H. Protein sorting in yeast: the localization determinant of yeast vacuolar carboxypeptidase Y resides in the propeptide. Cell. 1987 Mar 13;48(5):887–897. doi: 10.1016/0092-8674(87)90085-7. [DOI] [PubMed] [Google Scholar]
- Weisman L. S., Bacallao R., Wickner W. Multiple methods of visualizing the yeast vacuole permit evaluation of its morphology and inheritance during the cell cycle. J Cell Biol. 1987 Oct;105(4):1539–1547. doi: 10.1083/jcb.105.4.1539. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Woolford C. A., Daniels L. B., Park F. J., Jones E. W., Van Arsdell J. N., Innis M. A. The PEP4 gene encodes an aspartyl protease implicated in the posttranslational regulation of Saccharomyces cerevisiae vacuolar hydrolases. Mol Cell Biol. 1986 Jul;6(7):2500–2510. doi: 10.1128/mcb.6.7.2500. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zubenko G. S., Mitchell A. P., Jones E. W. Mapping of the proteinase b structural gene PRB1, in Saccharomyces cerevisiae and identification of nonsense alleles within the locus. Genetics. 1980 Sep;96(1):137–146. doi: 10.1093/genetics/96.1.137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zubenko G. S., Park F. J., Jones E. W. Genetic properties of mutations at the PEP4 locus in Saccharomyces cerevisiae. Genetics. 1982 Dec;102(4):679–690. doi: 10.1093/genetics/102.4.679. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zubenko G. S., Park F. J., Jones E. W. Mutations in PEP4 locus of Saccharomyces cerevisiae block final step in maturation of two vacuolar hydrolases. Proc Natl Acad Sci U S A. 1983 Jan;80(2):510–514. doi: 10.1073/pnas.80.2.510. [DOI] [PMC free article] [PubMed] [Google Scholar]







