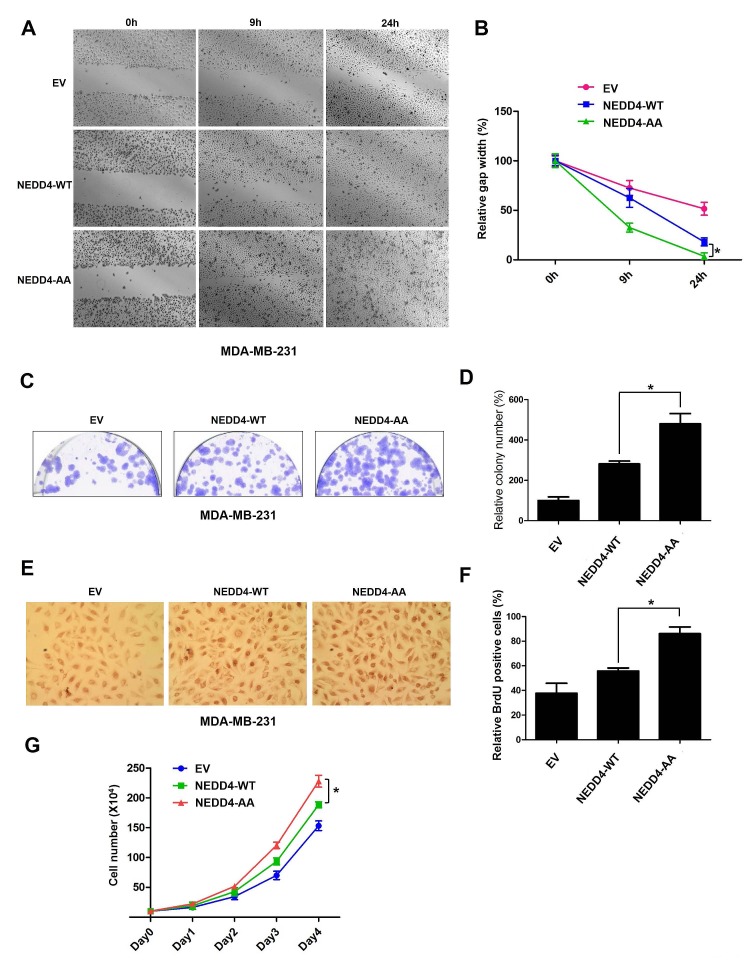
Figure 5. Loss of SCFβ-TRCP-mediated degradation of NEDD4 promoted cancer cell proliferation and migration.
(A-B) Scratch assays were performed with MDA-MB-231 cells that were infected with EV, HA-wild-type (WT)-NEDD4 or HA-S347A/S348A (AA)-NEDD4 encoding retroviral vectors followed by 3 days of puromycin (1 μg/ml) selection to eliminate non-infected cells. The generated various MDA-MB-231 cell lines were seeded on a 6-well plate and scratched on the surface with a pipette tip. Relative values were set at 1 for the gap width at the time of the scratch. Representative photographs at time points 0, 9 and 24 hours after the scratch (A). Measurements were done in duplicate in 3 separate experiments, and data were depicted as average gap width (B). The error bars represented mean ± SD (n = 3), * p<0.05 (Student's t-test), compared with cells expressing WT-NEDD4. (C-D) Colony formation assays were performed with the MDA-MB-231 cell lines generated in (A). After 4 days, the colonies were stained with crystal violet and counted (C). The number of surviving colonies were calculated as the average of triplicates (D). The error bars represented mean ± SD (n = 3), * p<0.05 (Student's t-test), compared with cells expressing WT-NEDD4. (E-F) BrdU labeling analysis was performed with MDA-MB-231 cell lines generated in (A). Cells were incubated with BrdU and uridine for 48 hours and representative photographs were taken (E). Percentage of BrdU positive cells was illustrated in (F). The error bars represented mean ± SD (n = 3), * p<0.05 (Student's t-test), compared with cells expressing WT-NEDD4. (G) MDA-MB-231 cells stably expressing control EV, HA-WT-NEDD4 or HA-S347A/S348A-NEDD4 were seeded and analyzed for cell proliferation capacities. The data shown are derived from three independent experiments, and the error bars represented mean ± SD. (* p<0.05, compared with cells expressing WT-NEDD4; Student's t-test).