Abstract
Foldamers provide an attractive medium to test the mechanisms by which biological macromolecules fold into complex three-dimensional structures, and ultimately to design novel protein-like architectures with properties unprecedented in nature. Here, we describe a large cage-like structure formed from an amphiphilic arylamide foldamer crystallized from aqueous solution. Forty eight copies of the foldamer assemble into a 5 nm cage-like structure, an omnitruncated octahedron filled with well-ordered ice-like water molecules. The assembly is stabilised by a mix of arylamide stacking interaction, hydrogen bonding and hydrophobic forces. The omnitruncated octahedra tessellate to form a cubic crystal. These findings may provide an important step towards the design of nanostructured particles resembling spherical viruses.
Keywords: de novo design, truncated octahedron, omnitruncated octahedron, foldamer, antimicrobial peptide mimic, supramolecular synthon, organic framework, crystal engineering
Nature uses a limited set of amino acids to build proteins, which take on a myriad of shapes with defined secondary, tertiary, and quaternary structures. In recent years, chemists have shown that this ability to fold into complex structures is not unique to natural biopolymers, and they have begun building “foldamers” comprised of defined sequences of non-natural building blocks that assemble into increasingly complex secondary structures, and even protein-like folds1-3. Moreover, foldamers have been designed that are responsive to ligand-binding, temperature, or that bind to native biologically important proteins, membranes, and oligosaccharides4-11. In this paper, we describe the structural assembly of an amphiphilic arylamide foldamer, 1 (Fig. 1a), which assembles into a 48-mer cage-like structure.
Figure 1. Crystal Structure of triarylamide compound 1.

a, The triarylamide compound 1. Two monomers in the asymmetric unit have different ligand coordination. b, Monomer 1 contains two acetates bound to each Cd(II) ion (grey). A hydrated Na+ ion (purple sphere) binds between the two Cd(II) sites, maintaining electrical neutrality. c, Monomer 2 has two acetates bound to one Cd(II), a single acetate bound to the second Cd(II) and no sodium ion.
Compound 1 was originally designed as a mimic of antimicrobial peptides12. It is a triarylamide comprising two 1,3-diaminobenzene units linked by a 4,6-dicarboxy-substituted pyrimidine and two terminal guanidine-containing amides. Pendant thioether substituents within the diaminobenzene units help to rigidify the structure and also provide points of attachment for positively charged aminoethyl sidechains. Together, these groups create a facially amphiphilic, positively charged structure, previously shown to be essential for their high antibacterial activity in vitro and in animal models12.
Here we show the crystallographic structure of 1 and confirm its amphiphilic structure, which is required for binding to bilayers. More importantly, the foldamer, which was crystallized from aqueous solution in the absence of membranes, associates to form a honeycomb geometry with each cubicle as a truncated octahedron. The assembly can be understood in terms of physicochemical principles, including the hydrophobic effect, aromatic stacking, hydrogen bonding, and ion-binding, and could advance the nano-scale engineering of complex molecular assemblies.
Results
Crystal structure of 1 shows the designed amphiphilicity
NMR studies showed that the triarylamide assembled into a higher order oligomer in aqueous solution (Supplementary Fig. S1 and S2). To obtain a high-resolution structure of 1, it was crystallized from aqueous solution by the hanging drop method in the presence of 0.05 M cadmium acetate and sodium sulfate (1.0 M). Two monomers with closely related structures form the asymmetric unit (Figs. 1b and 1c). Each monomer shows the expected amphiphilic structure, but the arylamide backbone is stabilised differently in the presence than in the absence of Cd2+ 13. In the crystal, Cd2+ ions displace the arylamide protons on the amide units connecting the phenyl and pyrimidyl rings. The foldamers also interact weakly with Cd2+ in aqueous solution (Supplementary Fig. 3). Each Cd2+ interacts with the pyrimidyl nitrogen and thioether – replacing the hydrogen-bonded interactions used in the original design with metal-ligand interactions. The neighboring aminoethyl group serves as a fourth ligand, and acetates or (in one case) a water molecule complete the ligand environment. However there are no ligand metal ion interactions between sites within a single molecule or between molecules in the crystal lattice. Thus, the metal ions appear to promote crystallization, not by bridging between sites as in assembly systems formed between polycoordinate metal ions and polydentate ligands14, but rather by subtly changing the physical and geometric properties of the molecule.
The structure of the triarylamide monomer in the crystallographic lattice displays the amphiphilic structure anticipated in the design. The trifluoromethyl groups and the nonpolar portions of the aryl backbone segregate from the strongly polar amine and guanidine sidechains. The overall arrangement is consistent with that determined by solid-state NMR of the triarylamide bound to phospholipid bilayers in the absence of Cd2+ 15. It is unlikely that the metal ion plays a significant role in the biological activity of the molecule, because other variants of this triarylamide that lack the thioether and/or the ethylamine ligating groups have high antimicrobial activity7,11.
A complex assembly stabilized by diverse interactions
The arylamide crystallizes in a space group of high symmetry, P 4̄ 3 n, which has not yet been seen for an organic framework structure. The unit cell (Fig. 2-3) contains 24 copies of the asymmetric dimer, arranged in the shape of an omnitruncated octahedron, a special case of an Archimedean solid, the truncated octahedron, which has eight hexagonal and six square faces. Each arylamide dimer associates with 113 water molecules, which are primarily located within the cores of the truncated octahedra, for a total of 2712 water molecules per unit cell.
Figure 2. Omnitruncated octahedron formed by 48 copies of the triarylamide.

a, The structure is viewed down the square faces, which have two-fold symmetry. The hexagonal faces have crystallographic 3-fold symmetry as shown. The inset shows the packing between arylamide neighbors interacting at edges between hexagonal faces. b, The structure is viewed down the hexagonal face, which has three-fold crystallographic symmetry. The inset shows that the individual arylamides interact with their aryl groups stacked in an offset manner, the adjacent terminal amides in a tight hydrogen bond, and the trifluoromethyl groups in close proximity.
Figure 3. Interactions of small molecules and water with the triarylamide.
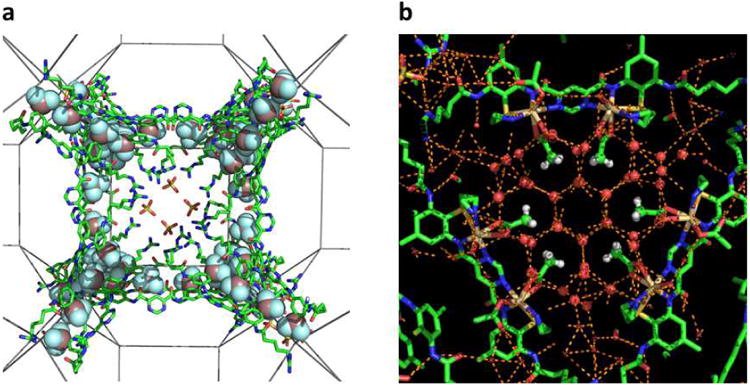
a, Along the square face, sulfate ions are shown interacting with the Arg-like sidechain of the arylamides. Trifluorometyl groups are also shown in space-filling representation; b, along the square face, the acetates are shown with their methyl groups (protons included) surrounded by a clathrate of water molecules.
The 48 triarylamides lie with their backbones roughly parallel to the edges of the eight hexagons (Fig. 2). Each arylamide engages in two distinct types of interactions, which together uniquely define and stabilise the overall assembly. Arylamides that lie along the edges of neighboring hexagons interact in what we term a “trifluoromethyl zipper” interaction, in which the water-repelling trifluoromethyl groups intimately interdigitate along a non-exact two-fold axis of symmetry. This trifluoromethyl zipper arrangement is reminiscent of the leucine zippers16 that stabilize coiled coils in water-soluble proteins, as well as the alanine coils found frequently in membrane proteins17. While engaging in trifluoromethyl zipper interactions, the arylamides also engage in a second type of interaction between arylamides that lie within a single hexameric ring. We designate this interaction the arylamide elbow motif; the terminal phenyl rings of the arylamides in this motif stack in a face-to-face interaction with the centers of the rings offset as often seen in aromatic stacking18. The dimer is further stabilised by tight hydrogen bonding between the terminal amide groups, and their trifluoromethyl groups also cluster together. The hydrogen-bonded interaction between meta-substituted amides leads to a 120° angle between the two arylamides, which is repeated to create the hexameric rings.
The simultaneous interactions of the triarylamides in both the trifuoromethyl zipper and the arylamide elbow motif create the 48-mer assembly seen in the crystal (Fig. 2). The arylamide elbows create the hexamers, while the trifluoromethyl zippers couple adjacent hexamers to form the overall three-dimensional structure. Interestingly, this arrangement also leads to extensive clustering of the trifluoromethyl groups along the vertices, creating fluorocarbon cores (Fig. 3a).
Electrostatic and hydrogen-bonded interactions between sulfate ions and the arginine-like guanidine sidechains feature prominently in each four-sided face of the truncated octahedral (Fig. 3). A total of four sulfates are seen in each face, two coming from each unit cell of the crystal lattice (Fig. 3a). Each sulfate receives a total of six hydrogen bonds from three guanidine-containing sidechains that engage the anion in a bidentate interaction. The hexagonal faces also show a rich array of molecular interactions; the methyl groups of the acetates (counterions of the Cd2+ ions) project towards the center of the hexagon (Fig. 3b). Each acetate ion is surrounded by a clathrate of water molecules, which join near the center to create a nearly ideal ice-like hexagon of water molecules.
The individual truncated octahedra found in the unit cell tessellate in three dimensions to form a cube. Intersecting square and hexagonal channels run through the length of the crystal (Fig. 4). Arylamides that do not form a trifluoromethyl zipper within an individual unit cell, form an equivalent zipper motif with other monomers from adjacent unit cells. Also, the packing of the truncated octahedra contribute to the cluster of sulfate/guanidine interactions along the square channels, as well as the clustering of trifluoromethyl groups at the vertices and along the hexagonal channels. Thus, the same interactions that stabilise individual truncated octahedra also contribute to their packing into a crystal lattice.
Figure 4. Packing of the unit cells in the crystal structure.

Three-dimensional space is compactly packed by the unit cells, viewed down the square (a) and hexagonal (b) channels.
Assembly of compound 1 in aqueous solution
To determine how the compound behaves in solution, we also performed NMR and UV-visible spectroscopy experiments in a buffer similar to the crystallization solution. Proton assignments for the peaks resonating between 7.0-10.0 ppm are obtained by 1D and 2D NMR (Supplementary Fig. 1). NMR titration of the compound with Cd2+ ions (Supplementary Fig. 2) shows that upon addition of metal ion the peaks were broadened and in some cases appear at new positions in the spectrum. At sub-stoichiometric Cd2+ concentrations, multiple peaks are observed, indicating that different forms of 1 with 0, 1, and 2 equivalents of Cd2+ bound were in slow to intermediate exchange on the time scale of the differences in chemical shift between the various forms (micro to milliseconds). After a stoichiometric amount (two equivalents per foldamer) of Cd2+ is reached the peaks begin sharpening as the distribution becomes more homogeneous. Little change is observed after 5.0 equivalents (40 mM) were added. These data show that the foldamer binds Cd2+ stoichiometrically at this concentration, and that it existed in the Cd2+-bound state under conditions of crystallization. Also, UV-visible spectroscopy confirmed that Cd2+ binds to the foldamer in a 2:1 (metal ion/foldamer) stoichiometry, as in the crystal (Supplementary Fig. 3).
To examine the self-association of the foldamer we examined the concentration dependence of its proton and 19F NMR spectra, while holding the Cd2+ constant at saturating concentrations. As the concentration was increased to 2 mM the proton NMR spectra showed small changes, while at 8.2 mM the peaks shifted further and two new peaks appeared in the spectrum (Fig. 5a). The new peaks are assigned to amides and guanidine protons from 1, as was confirmed by hydrogen-deuterium exchange (Fig. 5b). The fact that, at the highest concentration, the amides were observable at pH 7.5 in H2O shows that their exchange with bulk solvent is slowed by the formation of an oligomer. The concentration dependent 19F NMR spectra also showed biphasic changes (Fig. 5c). In this case, larger changes in chemical shift were observed between 0.1 and 2 mM. A second process occurs at higher concentrations (between 2 and 10 mM), causing the peaks to shift in the reverse direction and to broaden. While a quantitative analysis is complicated by the many equilibria, these data clearly show that 1 associates in aqueous solution, and that the mean association state increases as the concentration increases.
Figure 5. NMR studies on the foldamer 1.

a, 1H NMR spectra as a function of the concentration of 1 (298 K, 95% H2O/5% D2O, 30 mM CdSO4, 100 mM HEPES pH 7.5, 600 mM NaOAc). b, Comparison of the same sample recorded in H2O and D2O shows the disappeared exchangeable protons E and F in D2O highlighted by the dash lines. The H2O sample was 8.2 mM compound in 30 mM cadmium sulfate, 100 mM HEPES pH 7.5, 600 mM sodium acetate and the D2O sample was obtained by lyophilizing the H2O sample overnight and then adding D2O. c, 19F NMR spectra as a function of concentration of 1 at 95% H2O/5% D2O, 100 mM HEPES pH 7.5, 1 M NaOAc, 50 mM CdSO4. Assignments for protons A-F are given in Supplementary Figs. 1 and 2. Peaks labelled with a * are impurities in the buffer.
Discussion
Proteins are built up from secondary structures with pronounced facially amphiphilic character, which have also served as building blocks in the design of the first helical bundles composed of peptides synthesised from both α- and β-amino acids. Therefore, it was of considerable interest to determine the structures formed by the present triarylamides. The discovery of a large cage-like structure is interesting, given the complex protein-like interactions that stabilise the assembly.
It is of interest to compare the observed structure of 1 with those more purposefully designed built from proteins19-21 or small molecules14. Organic crystals are often assembled through strong directional intermolecular forces in organic solvents22. One of the most noteworthy fields in crystal engineering is metal-organic frameworks, assembly systems with predetermined directionalities between polycoordinate metal ions and polydentate ligands14. Additionally, certain organic frameworks are built by strong hydrogen bonding23.
The assembly described here uses a diversity of interactions similar to those employed by natural proteins. It is also interesting to consider its construction in terms of “supramolecular synthons”, as in other examples of crystal engineering. The arylamide elbow motif in triarylamidecan be considered a supramolecular synthon, which engages in both hydrogen bonding and aromatic stacking (Fig. 6a, in pink shadow). The two N-H···O=C hydrogen bonds in the arylamide elbow motif have a distance of 2.8 Å and an angle of 157° and 166°, which show that they are strong hydrogen bonds as in other examples of crystal engineering24. The distance between the two phenyl rings is 3.7 Å, as observed in other crystal studies25. Moreover, the pronounced facially amphiphilic character and high symmetry of 1 gives rise to its assembly and crystallization into an omnitruncated octahedron in aqueous environments. The assembly can be conceptually analysed according to Aufbau principles26, although the complexity of its structure and self-association equilibria prior to crystallization render it difficulty to assign a detailed kinetic mechanism of assembly. Hydrogen bonding and aromatic stacking provide geometrically specific interactions that stabilise the aromatic elbow (Fig. 6a), and repetition of this interaction pattern leads to assembly of the hexameric ring (Fig. 6b) that forms the hexagonal face of the truncated octahedron. Two hexagons can further assemble via hydrophobic association of trifluoromethyl groups and hydrogen bonding between the guanidine and carbonyl of neighboring arginine-like groups stabilise (Fig. 6c). The formation of the solvent-inaccesible trifluoromethyl core is further consolidated by assembly of two more hexagons; the resulting saddle-shaped tetramer of hexamers forms a complete vertex for the truncated octahedron (Fig. 6d). Two such tetramers of hexamers associate and create a “double crown” assembly on the square face (Fig. 6e), which serves as a “long-range synthon” module for packing in the late stages of crystallization27. This supramolecular subunit packs in primitive cubic cells and generates a crystalline framework (Fig. 6f).
Figure 6. Hierarchical assembly of the foldamer crystal.

Hydrogen bonds are shown as yellow dashed lines and fluorine atoms are displayed as spheres. The yellow wires are lattices defined by the symmetry of the crystal. a, The supramolecular synthon (denoted in pink shadow) is characterized by its combined aromatic and hydrogen-bonding interactions. b, The elbow-like synthons induce the formation of a hexagonal ring. c, Two 6-membered rings associate by hydrophobic trifluoromethyl zippers and hydrogen bonds between the guanidine and carbonyl. d, e, Further stabilization by trifluoromethyl zippers prompts the assembly of a saddle-shaped tetramer of hexamers and then the double-crown-shaped long-range synthon module. f, The modules pack in primitive cubic cells to form the framework-like crystal.
In Conclusion, the crystallographic structure of 1 has revealed a visually arresting Archimedean solid. The interactions that stabilize this assembly are readily apparent and should encourage future rational designs of foldamers with well defined tertiary and quaternary structures.
Methods
Crystallography
The arylamide foldamer 1 was synthesised as described previously12, and dissolved at 20 mg/ml in water. Crystals were obtained using the hanging-drop vapor-diffusion method at room temperature, by mixing equal volumes of foldamer solution with reservoir solution containing 0.05 M CdSO4, 0.1 M HEPES pH 7.5 and 1.0 M NaOAc. Crystals were flash frozen with the cryoprotectant Parabar 10312 (Hampton Research), and data to 0.920 Å were collected on the beamlines 24-ID-E and 24-ID-C at Advanced Photon Source, Argonne National Laboratory. The structure was solved by direct methods, as described in the Supplementary Methods.
NMR spectroscopy
Spectra were recorded at 298 K on a Bruker 800 or 900 MHz spectrometer equipped with a cryogenic probe for 1H spectra or a Bruker 300 MHz for 19F spectra. 1H spectra typically were recorded with 256 scans and 31 ppm spectral width and 19F spectra typically were recorded with 256 scans and 100 ppm spectral width. 1H chemical shifts were referenced with respect to the residual water peak at 4.75 ppm and the 19F chemical shifts were calibrated using the external standard trifluoroacetic acid chemical shifts at -76.6 ppm. All spectra were processed and analysed using the program TopSpin 3.0 (Bruker, Karlsruhe, Germany). Prior to Fourier transformation, time domain data were multiplied by sine square bell window functions shifted by 90° and zero-filled once. Additional details are provided in Supplementary Methods.
Supplementary Material
Acknowledgments
S.-Q.Z. thanks N. H. Joh for technical assistance and G. Ulas for helpful discussions. This work was supported by NIH grant GM54616 to W.F.D., and the MRSEC program of NIH through a grant to the LRSM at the University of Pennsylvania. Use of the Advanced Photon Source was supported by the U. S. Department of Energy, Office of Science, Office of Basic Energy Sciences, under Contract No. DE-AC02-06CH11357.
Footnotes
Author Contributions: S.-Q.Z. and W.F.D. designed the experiments. S.-Q.Z. crystallized the foldamer, collected and processed the diffraction data, and performed the UV-vis experiments. V.P. and A.M. solved and refined the crystallographic structure. Y.W. conducted the NMR experiments. W.F.D., S.-Q.Z., Y.W. and A.L. conducted the structural analysis. All authors contributed to preparing figures and writing the manuscript.
Additional Information: Accession codes: Crystal structure for the triarylamide can be accessed from the Cambridge Crystallographic Data Centre (CCDC) with Deposition number 956538. These data can be obtained free of charge from The Cambridge Crystallographic Data Centre via www.ccdc.cam.ac.uk/data_request/cif
Competing financial interests: The authors declare no competing financial interests.
References
- 1.Goodman JL, Molski MA, Qiu J, Schepartz A. Tetrameric beta(3)-peptide bundles. Chembiochem: a European journal of chemical biology. 2008;9:1576–1578. doi: 10.1002/cbic.200800039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Petersson EJ, Schepartz A. Toward beta-amino acid proteins: design, synthesis, and characterization of a fifteen kilodalton beta-peptide tetramer. J Am Chem Soc. 2008;130:821–823. doi: 10.1021/ja077245x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Cheng RP, Gellman SH, DeGrado WF. beta-Peptides: from structure to function. Chem Rev. 2001;101:3219–3232. doi: 10.1021/cr000045i. [DOI] [PubMed] [Google Scholar]
- 4.Huc I. Aromatic oligoamide foldamers. European Journal of Organic Chemistry. 2004:17–29. [Google Scholar]
- 5.Hecht S, Huc I. Foldamers: Structure, Properties, and Applications. Wiley-VCH; Weinheim, Germany: 2007. [Google Scholar]
- 6.Lee EF, et al. Structural basis of Bcl-xL recognition by a BH3-mimetic alpha/beta-peptide generated by sequence-based design. Chembiochem. 2011;12:2025–2032. doi: 10.1002/cbic.201100314. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Tew GN, Scott RW, Klein ML, DeGrado WF. De novo design of antimicrobial polymers, foldamers, and small molecules: from discovery to practical applications. Acc Chem Res. 2010;43:30–39. doi: 10.1021/ar900036b. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Moore DT, Berger BW, DeGrado WF. Protein-protein interactions in the membrane: sequence, structural, and biological motifs. Structure. 2008;16:991–1001. doi: 10.1016/j.str.2008.05.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Horne WS, Gellman SH. Foldamers with heterogeneous backbones. Accounts of chemical research. 2008;41:1399–1408. doi: 10.1021/ar800009n. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Goodman CM, Choi S, Shandler S, DeGrado WF. Foldamers as versatile frameworks for the design and evolution of function. Nat Chem Biol. 2007;3:252–262. doi: 10.1038/nchembio876. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Scott RW, DeGrado WF, Tew GN. De novo designed synthetic mimics of antimicrobial peptides. Curr Opin Biotechnol. 2008;19:620–627. doi: 10.1016/j.copbio.2008.10.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Choi S, et al. De novo design and in vivo activity of conformationally restrained antimicrobial arylamide foldamers. Proc Natl Acad Sci U S A. 2009;106:6968–6973. doi: 10.1073/pnas.0811818106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Tang H, Doerksen RJ, Jones TV, Klein ML, Tew GN. Biomimetic facially amphiphilic antibacterial oligomers with conformationally stiff backbones. Chem Biol. 2006;13:427–435. doi: 10.1016/j.chembiol.2006.02.007. [DOI] [PubMed] [Google Scholar]
- 14.Farrusseng D. Metal-organic frameworks: applications from catalysis to gas storage. Wiley-VCH; 2011. [Google Scholar]
- 15.Su Y, DeGrado WF, Hong M. Orientation, dynamics, and lipid interaction of an antimicrobial arylamide investigated by 19F and 31P solid-state NMR spectroscopy. J Am Chem Soc. 2010;132:9197–9205. doi: 10.1021/ja103658h. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Betz SF, DeGrado WF. Controlling topology and native-like behavior of de novo-designed peptides: design and characterization of antiparallel four-stranded coiled coils. Biochemistry. 1996;35:6955–6962. doi: 10.1021/bi960095a. [DOI] [PubMed] [Google Scholar]
- 17.Walters RF, DeGrado WF. Helix-packing motifs in membrane proteins. Proc Natl Acad Sci U S A. 2006;103:13658–13663. doi: 10.1073/pnas.0605878103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Meyer EA, Castellano RK, Diederich F. Interactions with aromatic rings in chemical and biological recognition. Angewandte Chemie. 2003;42:1210–1250. doi: 10.1002/anie.200390319. [DOI] [PubMed] [Google Scholar]
- 19.King NP, et al. Computational design of self-assembling protein nanomaterials with atomic level accuracy. Science. 2012;336:1171–1174. doi: 10.1126/science.1219364. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Lanci CJ, et al. Computational design of a protein crystal. Proc Natl Acad Sci U S A. 2012;109:7304–7309. doi: 10.1073/pnas.1112595109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Lai YT, Tsai KL, Sawaya MR, Asturias FJ, Yeates TO. Structure and flexibility of nanoscale protein cages designed by symmetric self-assembly. J Am Chem Soc. 2013;135:7738–7743. doi: 10.1021/ja402277f. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Desiraju GR. Crystal engineering: a holistic view. Angewandte Chemie. 2007;46:8342–8356. doi: 10.1002/anie.200700534. [DOI] [PubMed] [Google Scholar]
- 23.Desiraju GR. Crystal design: structure and function. Wiley; 2003. [Google Scholar]
- 24.Desiraju GR, Vittal JJ, Ramanan A. Crystal engineering: a textbook. World Scientific; IISc Press; 2011. [Google Scholar]
- 25.Chartrand D, Theobald I, Hanan GS. Bis [4′-(3,5-dibromophenyl)-2,2′:6′,2′′-terpyridine]ruthenium(II) bis(hexafluorophosphate) acetonitrile disolvate. Acta Crystallographica Section E Structure Reports Online. 2007;63:m1561–m1561. doi: 10.1107/s1600536807020259. [DOI] [Google Scholar]
- 26.Desiraju GR. Crystal Engineering: From Molecule to Crystal. Journal of the American Chemical Society. 2013;135:9952–9967. doi: 10.1021/ja403264c. [DOI] [PubMed] [Google Scholar]
- 27.Ganguly P, Desiraju GR. Long-range synthon Aufbau modules (LSAM) in crystal structures: systematic changes in C6H6-nFn (0 ≤n≤ 6) fluorobenzenes. CrystEngComm. 2010;12:817. doi: 10.1039/b910915c. [DOI] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


