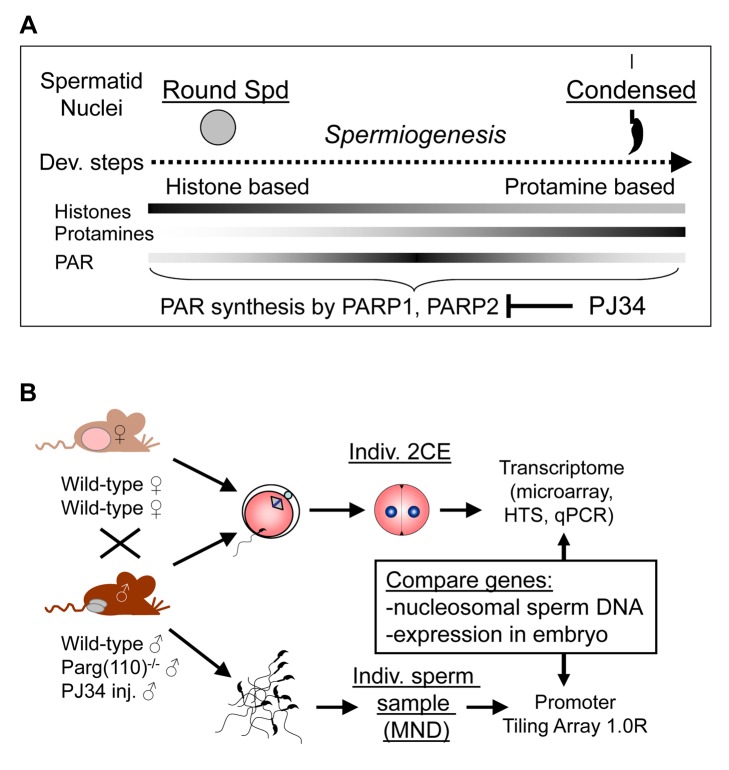
Figure 1. Experimental design to ascertain the impact of sperm chromatin structure on early embryonic gene expression.
(A) Efficiency of histone-to-protamine exchange in spermiogenesis depends in part on levels of poly(ADP-ribose) (PAR) formed transiently by the interplay of PAR polymerases (PARP1, PARP2) and PAR glycohydrolase (PARG). Inhibition of PAR synthesis by PJ34 or disruption of normal PARG activity in the Parg(110)−/− mouse leads to abnormal chromatin remodeling with retention of histones in sperm [29]. (B) Natural mating of Parg(110)−/− males or males treated with PJ34 with wild-type control females was used to obtain 2-cell stage embryos (2CE) for genome-wide transcriptional profiling at the individual embryo level using microarrays and high throughput sequencing (HTS). Cauda epididymal sperm from the fathers were used to identify genes associated with nucleosomes rather than protamines using micrococcal nuclease digestion (MND). Aberrant histone association of gene loci with differential expression of genes in two-cell embryos was assessed and compared to embryo expression data.