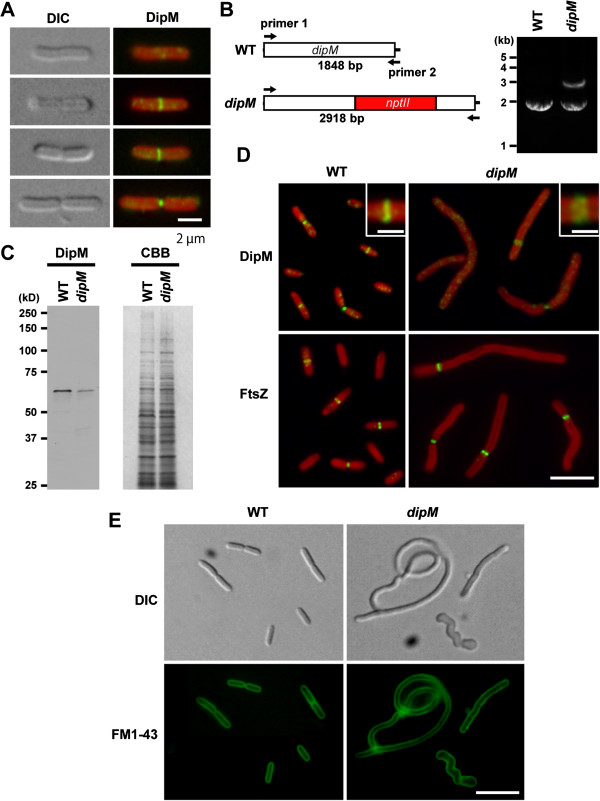
Figure 3.
DipM localizes at the mid cell position and is involved in cell division in the cyanobacterium S. elongatus. (A) Immunofluorescent images showing DipM localization at the mid cell position (the green fluorescence) in S. elongatus. The red color is the autofluorescence of chlorophyll. Differential interference contrast (DIC) images of the same cell are also shown. (B) The mutated S. elongatus dipM locus. nptII gene was inserted into the dipM locus and the insertion was detected with PCR using primer 1 and primer 2. The PCR produced 2918-bp or 1848-bp products from the inserted and intact chromosomes, respectively. S. elongatus has multiple genomes and the amplification of the two bands in the mutant indicates that the mutant cell possesses both mutated and intact dipM loci. (C) Immunoblot analysis showing reduction of the DipM protein level in the dipM mutant. Equal amounts of total protein were loaded in each lane and the equality of loading was confirmed by Coomassie Brilliant Blue (CBB) staining after SDS-PAGE. (D) Analyses of FtsZ ring frequency and DipM localization in wild-type and dipM mutant cells. The dipM mutant cells display an elongated shape because of cell division defect and possesse a single FtsZ ring. Localization of DipM in the dipM mutant is relatively diffusive compared to the wild type. Magnified views of the DipM localization are shown in the insets. (E) FM1-43 staining of the wild type and dipM mutant showing the pattern in the cytoplasmic membrane. The dipM mutant cells have no membrane septa, suggesting that constriction of both the outer and cytoplasmic membranes is impaired. Scale bar = 2 μm (A), 5 μm (D), 1 μm (insets of D), 10 μm (E).