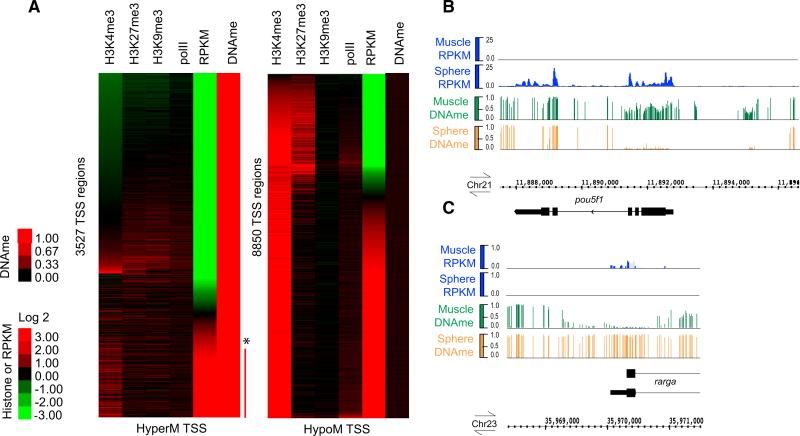
Figure 4. Relationship of DNAme and Histone Modifications at the TSS to Gene Expression during Sphere/PostMBT.
(A) TSS regions (±250bp) at sphere stage were separated into two groups based on their methylation status: either HyperM ≥ 0.8 or HypoM ≤ 0.2 (mean fraction CG methylation, scale 0 to 1) (note: TSSs with partial methylation are extremely rare at sphere). They were then subjected to separate k-means clustering with data sets for histone modifications and gene expression levels. Promoter histone modification status (mean log 2 ratio, array data; [Lindeman et al., 2011]) only available just after sphere/MBT (50% epiboly, 5.3 hpf). Gene expression RPKM levels (first exon, log 2 converted) from our total RNA-seq at sphere stage. Red bar and asterisk indicate loci with high DNAme and high RPKM, which are “false positives”as they mostly represent alternative or incorrectly annotated TSS with high DNAme (see Resultsand Supplemental Information).
(B and C) Example of developmentally regulated DNA demethylation or remethylation at the TSS upon differentiation into muscle with correlated gene expression. Browser snapshots of mean base fraction CG methylation tracks (scale 0 to 1) and relative RPKM values obtained from RNA-seq on total RNA from sphere stage (4 hpf) and adult muscle visualized on Integrated Genome Browser (IGB) for pou5f1 (RPKM scale 0–25) and rarga (RPKM scale 0 to 1).
See also Table S7 for (A) and Figure S5 and Tables S5 and S7 for (B) and (C).