Figure 1. More DSBs form in zmm mutants.
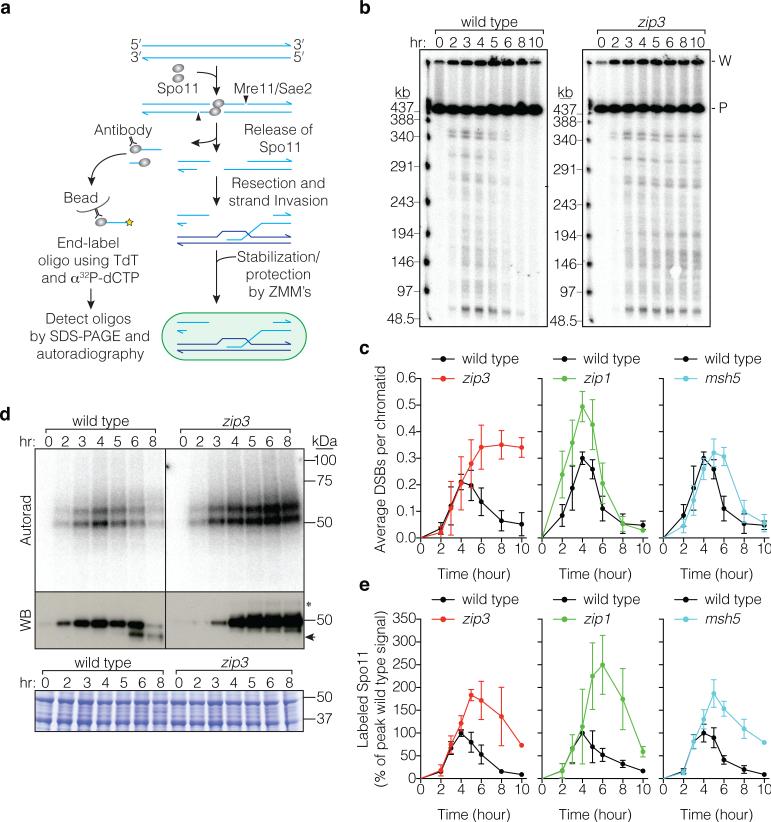
a, Spo11 generates a covalent protein-linked DSB; endonucleolytic cleavage releases Spo11 bound to a short oligo (detection method at left). Resection is followed by strand invasion and ZMM-dependent stabilization of intermediates fated to become crossovers. b, c, Representative pulsed-field gel Southern blots probed for Chr IX are in b and Poisson-corrected DSB quantification in c (mean ± SD, 3 cultures). P, parental; W, wells. d, e, Representative Spo11-oligo complex time courses are in d and quantification in e (mean ± SD for 3 cultures, except at 10 hr for msh5 and zip3 analyses (1 culture)). Radiolabeled Spo11-oligo complexes were detected by autoradiography (top panels) and total Spo11 was detected by anti-flag western blot (WB, middle). The main labeled species differ in oligo size10. Nearly all of the WB signal is Spo11 that has not made a DSB10. Asterisk, species co-migrating with upper Spo11-oligo complexes; arrowhead, proteolytic product. Extract samples run separately and stained with Coomassie control for input to the IPs (bottom). In panels c and e, mutants are plotted with wild-type data collected in parallel.