Table 3.
Summary of Mendelian inheritance error rates on SNPs derived from the new and old versions of GATK and different mappers
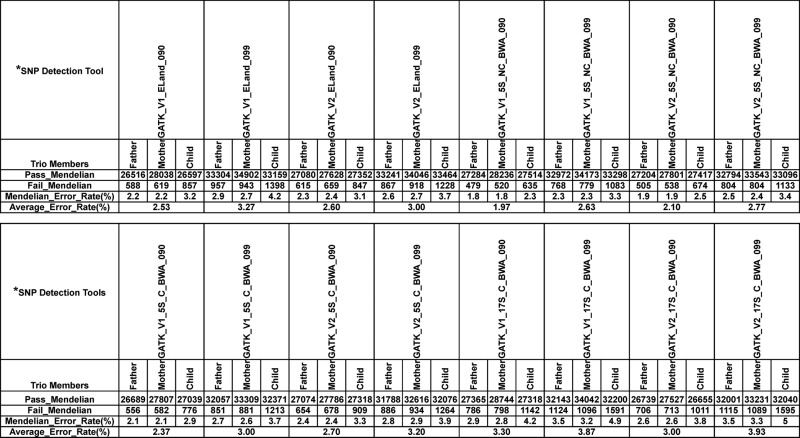 |
Summary of Mendelian inheritance error rates amongst the chosen family trio members (samples #9, #10 and #2; Supplementary Figure S1) for the SNPs detected with the new (V2.0 up to V2.2.4) or old (up to V1.6.7) versions of GATK and different mappers including Eland and BWA.
*Name designation for variations of GATK versions and mappers in the following format:
GATK_version_NumSample(option)_MapperContigOptionsForBWA(Option)_Mapper_VQSRThreshold. Version: V1 or V2 of GATK; NumSample: optional, if 5S, 5 samples with trio relations; if 17S, 17 samples in larger family; MapperContigOptionsForBWA: optional, if NC; no contigs in genome reference; C: with contigs in genome reference;
Mapper: BWA or Eland methods; VQSRThreshold: VQSR thresholds as 099 for 0.99 or 090 for 0.90. Similar results were obtained for the other two trio sets (with sample #3 or #4 as child) available in the family (data not shown).
