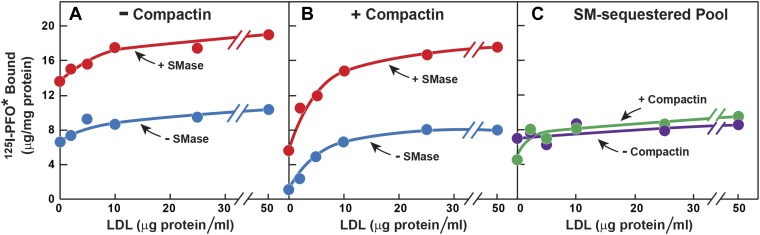
Figure 4. Effect of LDL on SM-sequestered pool of PM cholesterol.
(A and B) On day 0, SV-589 cells were setup in medium A at 1 × 105 cells per 60-mm dish. On day 2, cells were switched to lipoprotein-deficient medium C. On day 3, cells were treated with fresh medium C containing 50 μM mevalonate in the absence (A) or presence (B) of 50 μM compactin. On day 4, each monolayer received fresh medium D containing 50 μM mevalonate in the absence (A) or presence (B) of 50 μM compactin and the indicated concentration of LDL. After incubation for 5 hr at 37°C, the cells were washed twice with PBS at room temperature and treated with fresh medium D containing 50 μM mevalonate in the absence (A) or presence (B) 50 μM compactin and with or without 180 milliunits/ml SMase as indicated. After incubation for 1 hr at 37°C, the cells were washed five times as described in ‘Materials and methods’ and then incubated with 2 ml of ice-cold buffer A containing 25 μg/ml 125I-PFO* (8 × 103 cpm/μg). After 2 hr at 4°C, the total amount of cell surface binding of 125I-PFO* was determined. Each value represents the mean of duplicate incubations. (C) Graph showing the SM-sequestered pool of cholesterol in the absence and presence of compactin plotted as a function of the concentration of LDL. The SM-sequestered pool was calculated by subtracting the amount of 125I-PFO* bound in the absence of SMase from that bound after SMase treatment.