Figure 2.
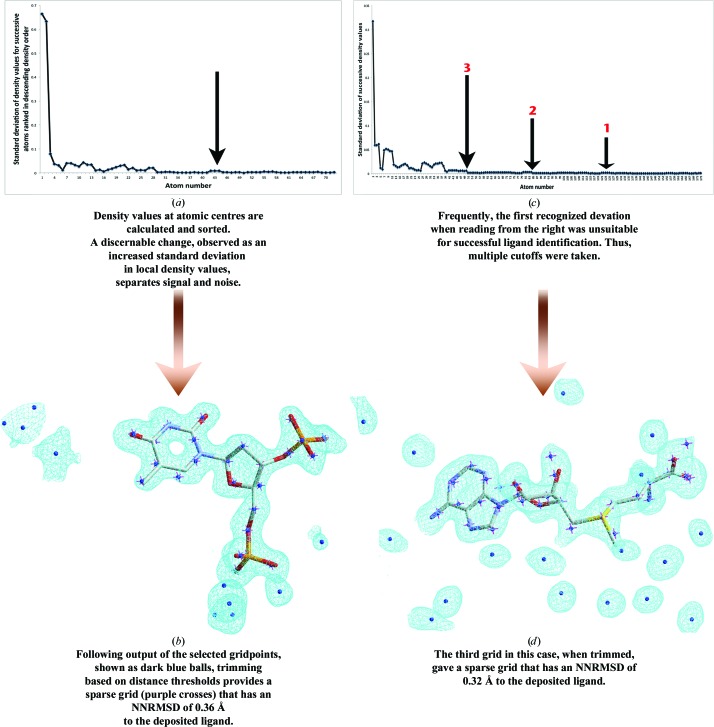
Trimming of pseudo-atomic grid clusters for feature comparison with ligand features. Difference (F o − F c, αc) maps are shown contoured at 2.5σ above the mean. (a) Density values for placed free atoms are sorted in descending order and the differences in adjacent values are calculated. The standard deviations of density differences are plotted, and only those atoms with density higher than the marked point are output. The data are shown for PDB entry 4iun (Li et al., 2010 ▶). (b) The output atoms, shown as balls, are trimmed further based on distance cutoffs to produce the final shape for screening, shown as crosses. It is an excellent match to the deposited ligand, THP. (c) As in (a) but with three clusters identified for the data in PDB deposition 3mb5 (Guelorget et al., 2010 ▶) are marked with arrows. (d) The third cluster, marked by arrow 3, is a good match to the final ligand, SAM.