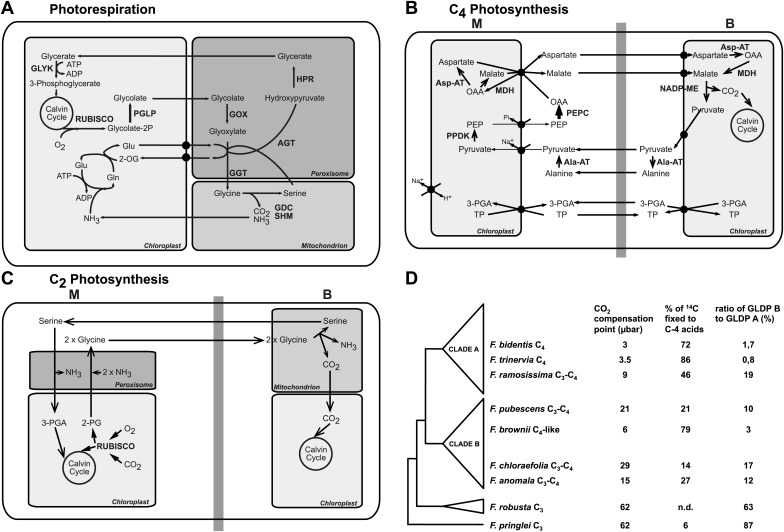
Figure 1. The genus Flaveria as a model organism to study C4 evolution.
Schematic view of the photorespiratory pathway (A), the NADP-ME type C4 pathway as it can be found in C4 Flaveria species (B) and the C2 photosynthesis pathway (C). (D) Phylogeny and physiological properties of selected Flaveria species. The phylogeny was redrawn according to McKown et al. (2005), CO2 compensation points are taken from Ku et al. (1991), incorporation of 14CO2 is from Moore et al. (1987) and the ratios of GLDP B (expressed in all chlorenchyma cells) and GLDP A (expressed in bundle sheath cells only) are from Schulze et al. (2013). (Abbreviations: AGT: serine glyoxylate aminotransferase; AlaAT: alanine aminotransferase; AspAT: aspartate aminotransferase; GDC: glycine decarboxylase complex; GGT: glutamate, glyoxylate-aminotransferase; GLYK: D-glycerate 3-kinase; GOX: glycolate oxidase; HPR: hydroxypyruvate reductase; MDH: malate dehydrogenase; NADP-ME: NADP dependent malic enzyme; PEPC: phosphoenolpyruvate carboxylase; PGLP: 2-phosphoglycerate phosphatase; PPDK pyruvate, phosphate-dikinase; RUBISCO: Ribulose-1,5-bisphosphat-carboxylase/-oxygenase; SHM: serine hydroxymethyltransferase; 2-OG: oxoglutarate; 2-PG 2-phosphoglycolate; 3-PGA: 3-phosphoglycerate; Gln: glutamine; Glu: glutamate; OAA: oxaloacetate; PEP: phosphoenolpyruvate; TP: triosephosphate).