Abstract
Objective:
Women who have low cobalamin (vitamin B12) levels are at increased risk for having children with neural tube defects (NTDs). The transcobalamin II receptor (TCblR) mediates uptake of cobalamin into cells. We evaluated inherited variants in the TCblR gene as NTD risk factors.
Methods:
Case-control and family-based tests of association were used to screen common variation in TCblR as genetic risk factors for NTDs in a large Irish group. A confirmatory group of NTD triads was used to test positive findings.
Results:
We found two tightly linked variants associated with NTDs in a recessive model: TCblR rs2336573 (G220R) (pcorr=0.0080, corrected for multiple hypothesis testing) and TCblR rs9426 (pcorr =0. 0279). These variants were also associated with NTDs in a family-based test prior to multiple test correction (log-linear analysis of a recessive model: rs2336573 (G220R) (RR=6.59, p=0.0037) and rs9426 (RR=6.71, p=0.0035)). We describe a copy number variant (CNV) distal to TCblR and two previously unreported exonic insertion-deletion polymorphisms.
Conclusions:
TCblR rs2336573 (G220R) and TCblR rs9426 represent a significant risk factor in NTD cases in the Irish population. The homozygous risk genotype was not detected in nearly one thousand controls, indicating this NTD risk factor may be of low frequency and high penetrance. Nine other variants are in perfect LD with the associated SNPs. Additional work is required to identify the disease-causing variant. Our data suggest that variation in TCblR plays a role in NTD risk and that these variants may modulate cobalamin metabolism.
Keywords: neural tube defects, spina bifida, transcobalamin II receptor (TCblR), cobalamin, vitamin B12, copy number variant (CNV)
INTRODUCTION
Neural tube defects (NTDs) are common birth defects, affecting ~1 in 1000 pregnancies [1, 2]. The neural tube closes during the fourth week of embryogenesis and gives rise to the spinal cord and brain. Incomplete closure of the neural tube causes a range of birth defects including spina bifida and anencephaly.
NTD etiology is multifactorial and includes environmental and genetic factors [3, 4]. Periconceptional folic acid supplementation in mothers can reduce the risk of an NTD affected pregnancy by up to 70% [5, 6]. Genetic variants relating to folate metabolism have also been implicated in contributing to NTD risk. The most well studied of these is the 677C>T variant in the methylene tetrahydrofolate reductase (MTHFR) gene (reviewed in [7]). In the European population, this variant may account for 26% of NTDs [8].
Cobalamin (vitamin B12) plays an important role in folate metabolism as a cofactor for methionine synthase (MTR), the enzyme that catalyzes the transfer of the methyl group from 5-methyl tetrahydrofolate (5-methyl THF) to homocysteine to produce methionine. Compromised methionine synthase function results in functional intracellular folate deficiency and a deficit of methyl groups for methylation reactions. Low maternal blood [9, 10, 11, 12, 13, 14, 15, 16, 17] or amniotic fluid [18, 19, 20, 21, 22] cobalamin levels have been associated with NTD risk. Levels of the cobalamin-transcobalamin II complex (holoTC) are also reduced in mothers during [23] or after [24] an NTD pregnancy.
Polymorphisms in cobalamin-related genes have been investigated for NTD risk. Several transporter proteins, receptors and converting enzymes are required to ensure that dietary cobalamin is delivered in an active form to methionine synthase and methylmalonyl-CoA mutase (MUT). Transcobalamin II (TCN2) transports cobalamin from the intestinal circulation into the bloodstream and ultimately to target tissues [25]. TCN2 P259R (rs1801198) was reported to be a maternal NTD risk factor [26] although other studies examined multiple TCN2 variants without detecting an effect [24, 27]. Methionine synthase reductase (MTRR) re-reduces oxidized cobalamin, thereby maintaining methionine synthase activity. Although some studies found no association between NTDs and variants in MTR or MTRR [28, 29, 30, 31, 32], others observed independent and/or joint effects [16, 33, 34, 35, 36, 37]. Cubilin (CUBN) is a receptor found on the luminal epithelium of the intestine and kidney that binds the cobalamin-intrinsic factor complex. An intronic single nucleotide polymorphism (SNP, rs1907362) in CUBN has been associated with NTDs [38].
Uptake of the cobalamin-transcobalamin II complex into cells and tissues occurs via binding to the recently identified transcobalamin II receptor (TCblR, also known as CD320 and 8D6) [39]. Because of TCblR’s essential role in cobalamin bioavailability, we evaluated its genetic variation for association with NTDs in a large Irish cohort. We report a significantly associated genetic risk factor for NTDs in TCblR.
MATERIALS AND METHODS
Study Populations
Recruitment of our Irish case and control populations has been previously described [40, 41, 42]. Our complete Irish case population consists of 586 families with an NTD affected child. This includes 442 full family triads (case, mother, father), 57 case-mother pairs, 4 case-father pairs, 44 cases-only and 39 families with parents or single parent only. In total there are 530 spina bifida cases and 21 encephalocele cases. Our control samples [n=999] were randomly selected from a bank of 56,049 blood samples taken from women during their first prenatal visit at the three major maternity hospitals in Dublin between 1986 to 1990, after excluding samples from women whose current or past pregnancies involved an NTD.
An additional 367 case families were recruited in the United Kingdom between 2001 and 2003, with the assistance of the UK Association for Spina Bifida and Hydrocephalus (ASBAH). This cohort includes 258 full family triads, 66 case-mother pairs, 2 case-father pairs, 38 case-only samples and 3 families with parents only. There are 354 spina bifida cases, 8 encephalocele cases and 2 cases with spina bifida and encephalocele. All UK families gave buccal swab samples collected using the protocol of Meulenbelt et al. [43] and completed questionnaires detailing the case’s NTD type, maternal pregnancy history and family history of birth defects.
These cohorts are comparable in regards to folic acid supplementation. Of the Irish NTD mothers, 91 women definitively answered whether they were taking any folic acid or multivitamins in the month before their last monthly period; 23 (25.3%) were on supplements. Of the UK NTD mothers, 73 women definitively answered the same question; 23 (31.5%) were on supplements. Additionally, the distributions of case birth years in the two cohorts were similar and largely predated the recommendation of folate supplementation by the United States Public Health Service in 1992. Case birth years in the Irish NTD cohort ranged from 1938 to 2003 with a median of 1981 and a standard deviation of 11 years. Case birth years in the UK NTD cohort ranged from 1926 to 2001 with a median of 1978 and a standard deviation of 15 years.
The sample collections were approved by the Health Research Board Research Ethics Committee (Dublin, Ireland), the UK Multi-Centre Research Ethics Committee (MREC, University of Newcastle, UK) in collaboration with UK ASBAH, and the Institutional Review Board at the National Human Genome Research Institute (Bethesda, MD, USA). Written consent was obtained from all participants.
Genomic DNA was extracted from all blood samples and buccal swabs using the QIAamp DNA Blood Mini Kit kit (Qiagen).
African American control DNA (HD100AA) and Caucasian control DNA (HD200CAU) samples were purchased from the Coriell Cell Repositories.
SNP Selection and Discovery
To select a set of SNPs to capture common genetic variation in TCblR, we evaluated SNPs genotyped by HapMap [44]. SNPs within and up to 10kb from the gene were considered. A minimum SNP set representing all such HapMap SNPs was manually selected, while allowing exclusion of redundant SNPs (r2>0.8).
Up to eighteen Irish individuals, the majority of which were drawn from NTD families, were screened for unreported polymorphisms. Primers were designed to PCR amplify exons 1 through 5; the resulting amplicons were analyzed via fluorescent automated dideoxy DNA sequencing.
Genotyping
Multiplex Ligation-dependent Probe Amplification (MLPA) was used to estimate the relative copy number for the 14.8kb copy number variant (CNV) immediately downstream from TCblR. Briefly, probe pairs were designed to hybridize to test and control loci (primer sequences available upon request). The multiplex reaction contains probe pairs to detect five test loci within the CNV. Control loci included four single copy regions in TCblR and one single copy probe was placed in TCN2 for relative quantification. A locus in the androgen receptor (AR) was included as an additional control to ensure the method detects copy number differences of the X chromosome.
The MLPA reaction was performed as directed with SALSA kit reagents obtained from MRC-Holland (The Netherlands) using at least 50ng genomic DNA for each sample. After hybridization and ligation, products were PCR amplified and analyzed on an ABI 3100 Genetic Analyzer (Applied Biosystems). Comparisons of amplicon peak areas were used to estimate relative DNA quantity (DQ) scores for each locus. A real time PCR assay using the Applied Biosystems 7900HT Fast Real-Time PCR System (Applied Biosystems) was used to independently estimate copy number in a subset of samples with good correlation.
All other variants were genotyped by detection of allele-specific extension products via Matrix Assisted Laser Desorption/Ionization – Time of Flight (MALDI-TOF) mass spectrometry (Sequenom). Primer sequences and assay conditions are available upon request.
Genotyping quality was assessed in each sample population. More than 10% of Irish samples were repeated with ≥99% concordance for all variants. More than 8% of UK samples were repeated with ≥98% concordance for all variants. More than 7% of Caucasian (Coriell) samples were repeated with 100% concordance for all variants. More than 10% of African American (Coriell) samples were repeated with 100% concordance for all but two variants (E88del and rs9426, one discrepant sample each).
Genotyping call rates (i.e., success rates) averaged 97% for the Irish and UK samples. Call rates were at least 95% for all variants in all groups with the exception of rs4147651 in the Irish NTD cases (94%) and rs2232775 (R8Q) in the Irish controls (93%). Genotyping call rates averaged 98% in the Caucasian Coriell samples and 92% in the African American Coriell samples. Call rates were at least 93% for all variants in both Coriell groups with the exception of rs250503 (87%) in African Americans.
Only the NTD cases from Ireland were observed to deviate from Hardy-Weinberg equilibrium (HWE, p<0.01): rs2232775 (R8Q) NTD cases (χ2=15.4, p=8.9 ×10−05), rs2336573 (G220R) NTD cases (χ2=27.8, p=1.3E−07), rs9426 NTD cases (χ2=21.2, p=7.2E−06), and E88del NTD cases (χ2=51.1, p=8.7E−13). These four SNPs were in HWE in all other genotyped groups. Additionally, the first three SNPs are in strong r2 linkage disequilibrium (r2≥0.94), and similar genotyping results were obtained with these independent genotype assays. The E88del polymorphism has a low minor allele frequency (p~0.02), which may account for its failure to adhere to HWE in the NTD cases.
Discordant genotypes and triads exhibiting non-Mendelian inheritance for any single marker were excluded for that marker. Twenty-four samples with discrepancies for more than one marker were excluded from all analyses.
Haplotype analysis
Linkage disequilibrium (LD) in the region was estimated using Haploview (http://www.broad.mit.edu/mpg/haploview/, [45]. Haplotype blocks were defined based on D’ values using the Solid Spine of LD option in Haploview. Haplotype frequency estimates based on these block definitions were then generated for Irish NTD cases, NTD mothers and controls using PHASE 2.1 [46, 47]. A permutation test within PHASE 2.1 was used to test whether haplotype frequency distributions differed between controls and NTD cases or NTD mothers in the Irish cohort.
Statistical Analyses
A logistic regression model, with the number of risk alleles as the independent variable, was used as a primary test to evaluate each polymorphism for NTD association. In this multiplicative model, the odds ratio of two risk alleles is the square of the odds ratio of one risk allele. Additionally, one degree of freedom (DOF) models of dominant and recessive genetic risk were applied via logistic regression. These tests are not valid when any of the genotypes cells are empty; in these cases, Fisher′s Exact Test was used in place of the logistic regression to test for genotypic differences. These tests were used to perform NTD case-control comparisons as well as NTD mother-control comparisons in the Irish NTD cases, NTD mothers and controls.
The (mother, father, case) triads were analyzed by fitting multiplicative, dominant and recessive log-linear models with one DOF to test for case effects and one DOF to test for direct maternal effects [48]. The case term in the multiplicative log-linear model provides comparable information to the transmission-disequilibrium test (TDT) [49]. These analyses were modified to incorporate data from incomplete triads by using the expectation maximization (EM) algorithm [50]. These family-based tests were applied to Irish and UK NTD triads to test TCblR variants for association with NTDs.
Correction for multiple tests was by permutation (99,999 random permutations). This method accounts for any linkage disequilibrium (non-independence) between evaluated SNPs. Multivariate permuting of triads for log-linear analysis involved treating the test as a one-sample test, and permuting the hypothetical risk allele. Permutations of cases and controls were independent of permutations of triads, and the results were combined by Bonferroni adjustment so that the resulting adjusted p-values accounted for all tests and all SNPs while controlling the chance of any false positive (familywise error) at 5%.
RESULTS
Genomic Analysis
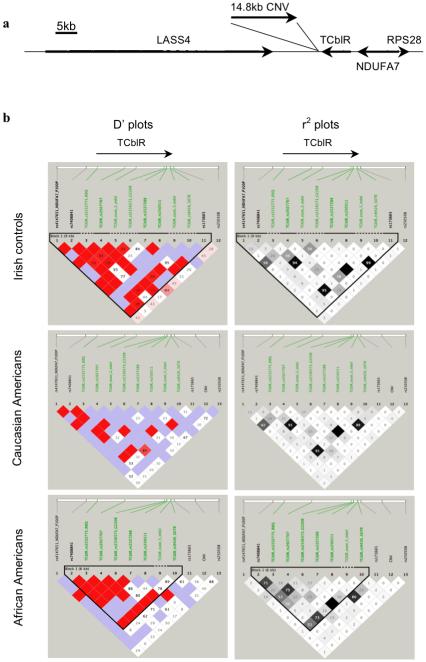
The TCblR gene spans ~6kb on chromosome 19 (Figure 1A). It is flanked by a gene encoding a subunit of the mitochondrial NADH dehydrogenase, NDUFA7, and the LAG1 longevity assurance homolog 4 (LASS4) gene, which encodes a ceramide synthase. TCblR and NDUFA7 are in close proximity (~3 kb) and transcribed from the same strand. In contrast, LASS4 is transcribed from the opposite strand and its 3’ terminus is mapped ~40kb away from the last exon of TCblR. The reference human genomic sequence appears contiguous through this region. Close examination of the genomic sequencing used to “build” this region of the genome revealed that the sequence in this interval had not been completed. The gap in the sequence was annotated in Genbank (contig, NT_077812) as including an undetermined number of a ~15kb repetitive element. During the course of this work, genome annotation was expanded to include areas containing potential segmental duplications. To ensure we were capturing all regional variability related to the TCblR gene, we first asked if this segmental duplication varied in copy number between individuals. Copy number variants (CNVs) have been implicated in disease and have been shown to affect expression for nearby genes. We developed a Multiplex Ligation-dependent Probe Amplification (MLPA) assay for this region and screened Caucasian and African American samples available from Coriell. We found that this region was polymorphic (i.e. this a bona fide CNV) in the African American (minor allele frequency, MAF=0.054) and Caucasian (MAF=0.021) samples tested. MLPA alone does not definitively allow determination of copy number; however, the most parsimonious model consistent with the MLPA derived data is that the common allele has two repeats while the minor allele has three repeats. The CNV was not found to be in strong LD with any other marker (Figure 1B). While it could serve as a candidate variant, we excluded it from further consideration because of its low MAF and the relatively large quantity of DNA consumed by the MLPA assay.
Figure 1.
Genomic Structure and Linkage Disequilibrium in the TCblR Region. A) Genomic structure of the region including TCblR. Orientation, relative size and distance of LASS4 (53kb), the 14.8kb CNV, TCblR (6kb), NDUFA7 (10kb) and RPS28 (0.9kb) are shown. The 14.8kb CNV is shown in single copy, although two copies are predicted to be present on most chromosomes. B) Linkage disequilibrium plots of polymorphisms within and surrounding TCblR. Genotyping data from Irish controls (n=993), Caucasian Americans (n=79) and African Americans (n=83) were used to construct D’ and r2 LD plots. Labels for variants within TCblR are green. Relative distances between markers are shown with two exceptions: the CNV is approximately 450 bases downstream from TCblR, and rs250508 is approximately 32kb downstream from TCblR.
SNP analysis and Linkage Disequilibrium in the TCblR Region
We selected ten HapMap [44] polymorphisms to cover the region within and flanking TCblR. Additionally, while developing the MLPA assay we discovered a C>GA insertion-deletion (indel) polymorphism in the 3’UTR of exon 5 located 83 bases downstream from the stop codon. Lastly, to search for unreported common variation, limited screening of TCblR coding regions was performed by directly sequencing exons in at least twelve Irish individuals. This revealed a three nucleotide insertion-deletion (indel) polymorphism in exon 2, resulting in the presence or absence of one of the three tandem glutamic acid (E) residues starting at codon 86 (TCblR E88del).
These twelve polymorphisms and the CNV were typed in Caucasians and African Americans to determine the linkage disequilibrium structure in the region (Figure 1B). Blocks defined by measures of D’ revealed that the upstream region and the majority of TCblR are in a single block of LD (Figure 1). In the Irish population, this block encompassed all markers within TCblR. Most of the markers examined are independently informative (r2 < 0.8). The exceptions consist of two sets of markers which are in high r2 LD; 1) rs2232775 (R8Q), rs2336573 (G220R) and rs9426 (r2 ≥ 0.94); and 2) rs250511 and rs173665 (r2 = 0.99). A total of twelve markers were genotyped in the Irish NTD cohort (Table 1). The exon 5 indel had the lowest MAF (0.001) and was excluded from further study.
Table 1.
Genotype Distributions and Allele Frequencies in Irish Controls and NTD Triads*
| Marker | Controls | NTD Children |
NTD Mothers |
NTD Fathers |
|---|---|---|---|---|
| Genotypes | N (freq.) | N (freq.) | N (freq.) | N (freq.) |
| Alleles | Freq. | Freq. | Freq. | Freq. |
| NDVFA7 | ||||
| rs4147651 | ||||
| GG | 636 (0.66) | 314 (0.63) | 310 (0.62) | 289 (0.64) |
| GA | 290 (0.30) | 167 (0.33) | 175 (0.35) | 140 (0.31) |
| AA | 31 (0.03) | 21 (0.04) | 16 (0.03) | 24 (0.05) |
| G | 0.82 | 0.79 | 0.79 | 0.79 |
| A | 0.18 | 0.21 | 0.21 | 0.21 |
| rs7408841 | ||||
| GG | 377 (0.39) | 210 (0.41) | 223 (0.44) | 193 (0.42) |
| GC | 459 (0.48) | 218 (0.43) | 218 (0.43) | 201 (0.44) |
| CC | 129 (0.13) | 81 (0.16) | 63 (0.13) | 61 (0.13) |
| G | 0.63 | 0.63 | 0.66 | 0.65 |
| C | 0.37 | 0.37 | 0.34 | 0.35 |
| TCblR | ||||
| rs2232775 | ||||
| R8Q | ||||
| TT | 846 (0.91) | 479 (0.91) | 479 (0.93) | 423 (0.91) |
| TC | 82 (0.09) | 44 (0.08) | 33 (0.06) | 39 (0.08) |
| CC | 0 (0.00) | 6 (0.01) | 2 (0.00) | 3 (0.01) |
| T | 0.95 | 0.95 | 0.96 | 0.95 |
| C | 0.05 | 0.05 | 0.04 | 0.05 |
| TCblR | ||||
| rs2927707 | ||||
| TT | 481 (0.50) | 259 (0.51) | 243 (0.48) | 227 (0.50) |
| TC | 409 (0.43) | 212 (0.41) | 212 (0.42) | 189 (0.42) |
| CC | 71 (0.07) | 41 (0.08) | 51 (0.10) | 38 (0.08) |
| T | 0.71 | 0.71 | 0.69 | 0.71 |
| C | 0.29 | 0.29 | 0.31 | 0.29 |
| TCblR | ||||
| E88del | ||||
| E/E | 915 (0.96) | 495 (0.95) | 493 (0.97) | 436 (0.96) |
| E/del | 35 (0.04) | 20 (0.04) | 15 (0.03) | 18 (0.04) |
| del/del | 0 (0.00) | 5 (0.01) | 0 (0.00) | 0 (0.00) |
| E | 0.98 | 0.97 | 0.99 | 0.98 |
| del | 0.02 | 0.03 | 0.01 | 0.02 |
| TCblR | ||||
| rs2336573 | ||||
| G220R | ||||
| CC | 889 (0.92) | 464 (0.90) | 468 (0.93) | 417 (0.91) |
| CT | 80 (0.08) | 42 (0.08) | 35 (0.07) | 39 (0.08) |
| TT | 0 (0.00) | 8 (0.02) | 1 (0.00) | 3 (0.01) |
| C | 0.96 | 0.94 | 0.96 | 0.95 |
| T | 0.04 | 0.06 | 0.04 | 0.05 |
| TCblR | ||||
| rs2227288 | ||||
| GG | 780 (0.81) | 417 (0.82) | 404 (0.80) | 364 (0.80) |
| GC | 179 (0.19) | 90 (0.18) | 97 (0.19) | 89 (0.20) |
| CC | 6 (0.01) | 4 (0.01) | 6 (0.01) | 2 (0.00) |
| G | 0.90 | 0.91 | 0.89 | 0.90 |
| C | 0.10 | 0.10 | 0.11 | 0.10 |
| TCblR | ||||
| rs250511 | ||||
| TT | 802 (0.82) | 453 (0.86) | 428 (0.83) | 392 (0.84) |
| TA | 175 (0.18) | 72 (0.14) | 83 (0.16) | 76 (0.16) |
| AA | 7 (0.01) | 4 (0.01) | 6 (0.01) | 0 (0.00) |
| T | 0.90 | 0.92 | 0.91 | 0.84 |
| A | 0.10 | 0.08 | 0.09 | 0.16 |
| TCblR exon 5 | ||||
| indel | ||||
| del/del | 982 (1.00) | 508 (1.00) | 504 (1.00) | 456 (1.00) |
| del/ins | 1 (0.00) | 1 (0.00) | 1 (0.00) | 1 (0.00) |
| ins/ins | 0 (0.00) | 0 (0.00) | 0 (0.00) | 0 (0.00) |
| del | 1.00 | 1.00 | 1.00 | 1.00 |
| ins | 0.00 | 0.00 | 0.00 | 0.00 |
| TCblR rs9426 | ||||
| CC | 882 (0.91) | 470 (0.90) | 473 (0.93) | 425 (0.91) |
| CT | 87 (0.09) | 44 (0.08) | 35 (0.07) | 39 (0.08) |
| TT | 0 (0.00) | 7 (0.01) | 1 (0.00) | 3 (0.01) |
| C | 0.95 | 0.94 | 0.96 | 0.95 |
| T | 0.05 | 0.06 | 0.04 | 0.05 |
| rsl73665 | ||||
| GG | 790 (0.82) | 445 (0.86) | 421 (0.83) | 389 (0.84) |
| GA | 171 (0.18) | 66 (0.13) | 79 (0.16) | 74 (0.16) |
| AA | 8 (0.01) | 4 (0.01) | 6 (0.01) | 1 (0.00) |
| G | 0.90 | 0.93 | 0.91 | 0.92 |
| A | 0.10 | 0.07 | 0.09 | 0.08 |
| rs250508 | ||||
| CC | 304 (0.32) | 176 (0.35) | 166 (0.34) | 140 (0.30) |
| CT | 482 (0.50) | 222 (0.44) | 235 (0.48) | 222 (0.47) |
| TT | 169 (0.18) | 107 (0.21) | 87 (0.18) | 107 (0.23) |
| C | 0.57 | 0.57 | 0.58 | 0.54 |
| T | 0.43 | 0.43 | 0.42 | 0.46 |
Due to rounding, group frequencies may not sum to 1.
Evaluation of Individual Markers for Association with NTDs
Each marker (n=11) was tested for case-control associations with NTDs in the Irish population using three models of logistic regression: a multiplicative model, a dominant model and a recessive model. Significant associations are shown in Table 2. Essentially, three positive associations were found. First, the three highly linked SNPs were found to be significantly associated in a recessive model of case effect (rs2232775 (R8Q), OR=10.6 [95% CI 1.28-88.56], p=0.029; rs2336573 (G220R), Fisher’s Exact Test, p=0.0002; rs9426, Fisher’s Exact Test, p=0.0006). These SNPs share very high r2 values, and as predicted, yield similar association results. Second, TCblR E88del is also significantly associated with NTDs in a recessive model of case effect (OR=9.2 [95% CI 1.3-88.9], p=0.04). Third, the highly linked SNP pair (rs250511, OR=0.7 [95% CI 0.56-0.99], p=0.04; rs173665, OR=0.7 [95% CI 0.51-0.94], p=0.017) is significantly associated with NTDs in a dominant model of case effect. Additionally, rs173665 was associated when a multiplicative model of case effect was applied (OR=0.7 [95% CI 0.55-0.96], p=0.02). Upon correcting for multiple tests, two markers from the highly linked trio remained significant: rs2336573 (G220R) (p=0.008) and rs9426 (p=0.03), by Fisher’s Exact Test of a recessive model in cases vs. controls. No other comparisons were significant after adjusting for multiple tests.
Table 2.
Case-Control Logistic Regression of Significantly Associated TCblR Variants in the Irish Population.
| Logistic regression | Fisher’s Exact Test | |||||
|---|---|---|---|---|---|---|
| Model | OR* (95% CI) | p-value | corrected p-value |
p-value | corrected p-value |
|
| rs2232775 (R8Q) | Recessive | 10.6 (1.28-88.56) | 0.0288 | 0.3747 | ||
| E88del | Recessive | 9.2 (1.08-79.16) | 0.0428 | 0.6887 | ||
| rs2336573 (G220R) | Recessive | FTC** | 0.0002 | 0.0080 | ||
| rs250511 | Dominant | 0.7 (0.55-0.99) | 0.0422 | 1.0000 | ||
| rs9426 | Recessive | FTC | 0.0006 | 0.0279 | ||
| rsl73665 | Dominant | 0.7 (0.51-0.94) | 0.0170 | 0.7217 | ||
| rsl73665 | Multiphcative | 0.7 (0.55-0.96) | 0.0249 | 1.0000 | ||
OR for the Recessive Model is the odds of disease with 2 copies divided by odds of disease with 0 or 1 copy; OR for the Dominant Model is the odds of disease with 1 or 2 copies divided by odds of disease with 0 copies; OR for the Multiplicative Model is the odds ratio for disease for each copy of the allele.
FTC = failed to converge
Examining the same markers in NTD mothers and controls revealed no significant associations (data not shown).
A family-based test of association was also used to evaluate these eleven markers for case and maternal NTD risk (Table 3). Multiplicative (data not shown), dominant and recessive models of log-linear analysis were applied. Depending on the applied model and corresponding genotype frequencies, the test failed to converge (no result obtained) for some SNPs. One positive result was observed among the successfully tested SNPs. All three highly linked SNPs were significantly associated with a strong relative risk in log-linear analysis of the recessive model in cases (rs2232775 (R8Q), RR=6.0, p=0.0086; rs2336573 (G220R), RR=6.6, p=0.0037; and rs9426, RR=6.7, p=0.0035). However, these results did not withstand correction for multiple tests (data not shown).
Table 3.
Dominant and Recessive Models of Log-Linear Analysis of TCblR Variants
| Republic of Ireland | United Kingdom | |||||||
|---|---|---|---|---|---|---|---|---|
| Variant | Dom RR* |
Dom p-value |
Rec RR |
Rec p-value |
Dom RR |
Dom p-value |
Rec RR |
Rec p-value |
| rs4147651 | ||||||||
| Case | 1.1048 | 0.4329 | 0.7598 | 0.3496 | FTC** | FTC | 0.6830 | 0.3386 |
| Mother | 1.0342 | 0.5795 | 0.5868 | 0.1055 | 0.9765 | 0.5303 | 1.0637 | 0.8976 |
| rs7408841 | ||||||||
| Case | FTC | FTC | 1.3588 | 0.0824 | 1.2350 | 0.2127 | 1.0259 | 0.9091 |
| Mother | FTC | FTC | 0.8482 | 0.4135 | 1.1464 | 0.3378 | 0.8033 | 0.3594 |
| rs2232775 | ||||||||
| Case | FTC | FTC | 6.000 | 0.0086 | 0.7965 | 0.4081 | FTC | FTC |
| Mother | FTC | FTC | 0.4567 | 0.3357 | 0.8394 | 0.1738 | FTC | FTC |
| rs2927707 | ||||||||
| Case | 1.0561 | 0.6148 | 0.7133 | 0.1091 | 0.9483 | 0.7146 | 0.9659 | 0.9014 |
| Mother | FTC | FTC | 1.2461 | 0.3263 | 0.9582 | 0.6013 | 0.8331 | 0.4988 |
| E88del | ||||||||
| Case | 1.3405 | 0.7462 | FTC | FTC | ND*** | ND | ND | ND |
| Mother | FTC | FTC | FTC | FTC | ND | ND | ND | ND |
| rs2336573 | ||||||||
| Case | FTC | FTC | 6.5871 | 0.0037 | 0.7064 | 0.2550 | FTC | FTC |
| Mother | FTC | FTC | 0.2469 | 0.1858 | 0.7518 | 0.1386 | FTC | FTC |
| rs2227288 | ||||||||
| Case | 1.0210 | 0.8971 | 0.6856 | 0.4932 | 0.8320 | 0.3962 | 0.4258 | 0.2315 |
| Mother | FTC | FTC | 2.6990 | 0.2006 | 0.8055 | 0.5434 | 0.4428 | 0.3369 |
| rs250511 | ||||||||
| Case | 0.7980 | 0.2625 | FTC | FTC | ND | ND | ND | ND |
| Mother | FTC | FTC | FTC | FTC | ND | ND | ND | ND |
| rs9426 | ||||||||
| Case | FTC | FTC | 6.7127 | 0.0035 | ND | ND | ND | ND |
| Mother | FTC | FTC | 0.2471 | 0.1656 | ND | ND | ND | ND |
| rsl73665 | ||||||||
| Case | 0.7253 | 0.1313 | FTC | FTC | 0.9480 | 0.8021 | 0.7332 | 0.7084 |
| Mother | FTC | FTC | FTC | FTC | FTC | FTC | 2.4150 | 0.4509 |
| rs250508 | ||||||||
| Case | 0.8428 | 0.1861 | 1.2416 | 0.1621 | ND | ND | ND | ND |
| Mother | 0.9832 | 0.6401 | 0.7748 | 0.1239 | ND | ND | ND | ND |
RR for the Recessive Model is the risk of disease with 2 copies divided by risk of disease with 0 or 1 copy; RR for the Dominant Model is the risk of disease with 1 or 2 copies divided by risk of disease with 0 copies.
FTC = Failure to converge
ND = not done
Note: none of the reported p-values are <0.05 upon correction for multiple tests (permutation)
Further evidence was sought by evaluating TCblR variation for NTD risk in an independent population. Seven SNPs were tested in a United Kingdom cohort of NTD triads (Table 3). Again, some tests failed to converge, including log-linear analysis of the recessive model as applied to two of the associated SNPs (rs2232775 (R8Q) and rs2336573 (G220R)). The remaining models of SNPs that were successfully tested were not found to be associated with case or maternal NTD risk in the UK population.
Haplotype Analysis of TCblR Variants for NTD Risk
To test for TCblR haplotype association, a haplotype was constructed based on the single block of LD in the Irish population (Figure 1B). rs2336573 (G220R) was retained to tag rs2232775 (R8Q) and rs9426, which were not included in these analyses. Similarly, rs173665 was retained as a tag for rs250511. The resulting haplotype block is defined by the remaining seven markers: rs4147651, rs7408841, rs2927707, E88del, rs2336573 (G220R), rs2227288, rs173665. Haplotype frequencies were estimated in Irish controls, NTD cases and mothers (Table 4). Five haplotypes account for >90% of the variation present in this haplotype block in the Irish population. A permutation test did not detect differences in haplotype frequencies between NTD cases and controls (p=0.3) or NTD mothers and controls (p=1.0).
Table 4.
Haplotype Frequency Estimates of the TCblR D’ Block*
| Haplotype | Controls | NTD cases | NTD mothers |
|---|---|---|---|
| GCTdelCGG | 0.33 | 0.32 | 0.30 |
| GGTdelCGG | 0.24 | 0.24 | 0.24 |
| AGCdelCGG | 0.19 | 0.21 | 0.21 |
| GGTdelCCG | 0.10 | 0.10 | 0.11 |
| GGCdelCGA | 0.10 | 0.07 | 0.09 |
| GCTdelTGG | 0.02 | 0.03 | 0.02 |
| GCTinsTGG | 0.02 | 0.03 | 0.01 |
| GGCdelCGG | 0.01 | 0.01 | 0.01 |
| p-value | 0.3** | 1.0** |
The haplotype block consists of the following markers: rs4147651, rs7408841, rs2927707, E88del, rs2336573 (G220R), rs2227288, rsl73665.
Result of permutation test for significant differences in haplotype frequencies in NTD cases vs. controls or NTD mothers vs. controls.
DISCUSSION
We evaluated the association between genetic variation in the transcobalamin II receptor gene, TCblR, and NTD risk. When considered singly, several independent (r2<0.8) TCblR polymorphisms were implicated as NTD risk factors in the Irish population. One pair of redundant SNPs (rs2336573 (G220R) and rs9426, r2~1) remained significantly associated with NTDs even after rigorous correction for multiple tests. One possible concern about the validity of this result is that the NTD cases are not in HWE for these SNPs (see Methods), which can result from genotyping error. We consider this unlikely because these two highly linked SNPs were genotyped with two independent assays, producing concordant results. Moreover, these variants were also significantly associated in an unadjusted recessive model of log-linear analysis that does not use control data, eliminating the possible problem of population stratification. Furthermore, this finding may not have withstood correction because power is reduced when examining a variant with a low MAF. Thus, we conclude that the signal seen with the rs2336573 (G220R) and rs9426 variants represents a true genetic risk factor in TCblR in the Irish population.
We sought to replicate this association in a sample of triads from the UK. As this sample did not include population controls, we were restricted to family-based tests of association. The lack of individuals homozygous for the minor allele of rs2336573 (G220R) precluded the use of the log-linear test for a recessive case effect. Because testing of the Irish cohort revealed a case effect, we also performed the TDT [49] but did not detect a case effect for either rs2336573 G220R or rs2232775 R8Q in the UK cohort (data not shown). The rs9426 SNP was not directly tested, but due to its high LD with the other two markers we would predict the same result in the UK NTD cohort. This lack of replication may be due to decreased power to detect an effect (~525 Irish NTD triads vs. ~345 UK NTD triads). Alternatively, because of differences in dietary factors, prenatal screening and/or greater ethnic heterogeneity, these variants may not contribute to NTDs in the UK population.
Haplotype analysis of all TCblR variants did not yield further evidence of association; therefore the identified SNP pair exhibits the strongest signal for NTD risk. It is possible that these variants do not alter gene function and a yet unidentified “causal” risk SNP may be linked to these two and reside on a haplotype we have yet to test. In addition, these SNPs (rs2336573 G220R and rs9426) have been typed in the HapMap CEU (Caucasian) population, and are in very high LD (r2~1) with five additional TCblR SNPs: rs17160390 (intron 1), rs2232783 (T149T), rs2232784 (S161S), rs2232785 (intron 3) and rs2227289 (T279T). Additionally, there are four such SNPs in introns of upstream genes: rs7249111, rs7250792, rs2288414 in NDUFA7 and rs4147645 in ribosomal protein S28 (RPS28).
We aligned the TCblR protein sequences from multiple species to determine whether any of these variants are in highly conserved regions. The “Conservation” track of the University of California - Santa Cruz Genome Browser (http://genome.ucsc.edu/; [ 51] displays syntenic regions of up to 17 species, and full alignments for TCblR were obtained only for chimp, rhesus, dog, cow, mouse, rat, tenrec and elephant. The only intronic SNP with a non-zero phastCons conservation score is rs17160390 (TCblR intron 1); its signal is relatively low. In contrast, rs2232783 (T149T), rs2232784 (S161S) are the only other SNPs in this set with phastCons conservation scores (0.742 and 0.677, respectively). These SNPs are part of the coding region for the second Low Density Lipoprotein Receptor Class A (LDLRa) domain in TCblR. As the name implies, these domains are shared by proteins in the LDL receptor superfamily, which includes the LDL receptor (LDLR), the LDL receptor related protein (LRP1), megalin/LRP2, the apolipoprotein 2 receptor (LRP8), and the VLDL receptor (reviewed in [52]). The LDLRa domains shared by these proteins contain cysteine-rich regions of approximately 40 amino acids and are involved in ligand binding. Whilst rs2232783 (T149T) and rs2232784 (S161S) fall into this well-conserved region, they are synonymous coding SNPs and appear to be weak candidates for affecting functionality.
Perhaps the most likely causal candidate SNP is rs2336573 (G220R), the nonsynonymous coding SNP directly tested in this study. This amino acid substitution replaces a glycine with arginine and is predicted to reside ten residues from the transmembrane and cytoplasmic domains found at the carboxyl terminus of the protein. We aligned and examined the TCblR orthologs from 23 mammalian species. This position contains glycine in 14 species. The other species contain glutamic acid, arginine, alanine or tryptophan at this residue. The moderate conservation of this amino acid implies functional tolerance of the rs2336573 (G220R ) variant in TCblR While such speculation is intriguing, functional analyses of these individual variants will be required to determine which of these NTD-associated SNPs may be the direct contributor to NTD risk.
Strengths of the current study include a large, homogenous Irish population, as well as a large UK population for replication. One limitation was the lack of UK population-based controls, which prevented replication of the same case-control analyses that identified rs2336573 (G220R) and rs9426 as NTD risk SNPs in the Irish group. Thus replication studies in independent cohorts are required to determine whether these TCblR polymorphisms contribute to NTDs in other populations. Additionally, the lack of metabolite measurements in either population prevented testing whether the risk genotype influences circulating cobalamin levels.
In summary, we identified a highly linked SNP pair (rs2336573 (G220R) and rs9426) in TCblR that is significantly associated in a recessive model of NTD case risk. The risk genotype (minor allele homozygote) was undetected in the large control group (n=979), contributing to the large relative risk (RR~6) observed in NTD cases. As a complex disease, multiple genetic risk factors are expected to contribute to the development of NTDs. Because of the relative rarity of the TCblR 2336573 (G220R) and TCblR rs9426 risk genotypes (~1% in NTD cases), we lack the power to perform meaningful interaction analyses of these genotypes with other established ntd risk factors such as MTHFR 677C>T. Its low frequency also means the risk genotype does not contribute greatly to the total genetic risk for NTDs in the general population. However, for those few individuals carrying the risk genotype, risk is increased ~6-fold.
Finding a genetic risk factor in the receptor required for cobalamin bioavailability further establishes the role of cobalamin in the development of NTDs. This raises the possibility that women with the risk variant may need additional dietary cobalamin to provide sufficient intracellular cobalamin for the developing embryo at the time of neural tube closure. There may also be individuals with these polymorphisms in the general population who need dietary supplements to maintain normal cobalamin homeostasis. These identified NTD risk SNPs are prime candidates to investigate in any disease state influenced by cobalamin metabolism.
Acknowledgements
These studies would not be possible without the participation of the affected families, and their recruitment by the Irish Association of Spina Bifida and Hydrocephalus and the Irish Public Health Nurses in Ireland, and by the UK Association for Spina Bifida and Hydrocephalus. The authors would like to thank Amanda Samuels and David Bernard for technical assistance in screening TCblR coding regions for polymorphisms.
Funding: The authors acknowledge research support from the intramural research programs of the National Human Genome Research Institute, the Eunice Kennedy Shriver National Institute of Child Health and Human Development, NIH, DHHS and the Health Research Board, Ireland. EVQ and JMS are supported by NIH grant DK064732.
Footnotes
Competing interests: None.
REFERENCES
- 1.Congenital malformations worldwide: A report from the International Clearinghouse for Birth Defects Monitoring Systems. Elsevier Science Publishers; Amsterdam: 1991. [Google Scholar]
- 2.Busby A, Abramsky L, Dolk H, Armstrong B, Addor MC, Anneren G, Armstrong N, Baguette A, Barisic I, Berghold A, Bianca S, Braz P, Calzolari E, Christiansen M, Cocchi G, Daltveit AK, De Walle H, Edwards G, Gatt M, Gener B, Gillerot Y, Gjergja R, Goujard J, Haeusler M, Latos-Bielenska A, McDonnell R, Neville A, Olars B, Portillo I, Ritvanen A, Robert-Gnansia E, Rosch C, Scarano G, Steinbicker V. Preventing neural tube defects in Europe: a missed opportunity. Reprod Toxicol. 2005;20(3):393–402. doi: 10.1016/j.reprotox.2005.03.009. [DOI] [PubMed] [Google Scholar]
- 3.Cowchock S, Ainbender E, Prescott G, Crandall B, Lau L, Heller R, Muir WA, Kloza E, Feigelson M, Mennuti M, Cederquist L. The recurrence risk for neural tube defects in the United States: a collaborative study. Am J Med Genet. 1980;5(3):309–14. doi: 10.1002/ajmg.1320050314. [DOI] [PubMed] [Google Scholar]
- 4.Seller MJ. Recurrence risks for neural tube defects in a genetic counseling clinic population. J Med Genet. 1981;18(4):245–8. doi: 10.1136/jmg.18.4.245. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Prevention of neural tube defects: results of the Medical Research Council Vitamin Study. MRC Vitamin Study Research Group. Lancet. 1991;338(8760):131–7. [PubMed] [Google Scholar]
- 6.Czeizel AE, Dudas I. Prevention of the first occurrence of neural-tube defects by periconceptional vitamin supplementation. N Engl J Med. 1992;327(26):1832–5. doi: 10.1056/NEJM199212243272602. [DOI] [PubMed] [Google Scholar]
- 7.Botto LD, Yang Q. 5,10-Methylenetetrahydrofolate reductase gene variants and congenital anomalies: a HuGE review. Am J Epidemiol. 2000;151(9):862–77. doi: 10.1093/oxfordjournals.aje.a010290. [DOI] [PubMed] [Google Scholar]
- 8.Kirke PN, Mills JL, Molloy AM, Brody LC, O'Leary VB, Daly L, Murray S, Conley M, Mayne PD, Smith O, Scott JM. Impact of the MTHFR C677T polymorphism on risk of neural tube defects: case-control study. BMJ. 2004;328(7455):1535–6. doi: 10.1136/bmj.38036.646030.EE. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Gaber KR, Farag MK, Soliman SE, El-Bassyouni HT, El-Kamah G. Maternal vitamin B12 and the risk of fetal neural tube defects in Egyptian patients. Clin Lab. 2007;53(1-2):69–75. [PubMed] [Google Scholar]
- 10.Hao L, Liu M, Liu X, Chen X, Tang Y, Li Z. [Relationship between folate, vitamin B12, total plasma homocysteine and mutation of reductase] Zhonghua Yu Fang Yi Xue Za Zhi. 2000;34(1):22–4. [PubMed] [Google Scholar]
- 11.Kirke PN, Molloy AM, Daly LE, Burke H, Weir DG, Scott JM. Maternal plasma folate and vitamin B12 are independent risk factors for neural tube defects. Q J Med. 1993;86;(11):703–8. [PubMed] [Google Scholar]
- 12.Wright ME. A case-control study of maternal nutrition and neural tube defects in Northern Ireland. Midwifery. 1995;11(3):146–52. doi: 10.1016/0266-6138(95)90029-2. [DOI] [PubMed] [Google Scholar]
- 13.Zhang T, Xin R, Gu X, Wang F, Pei L, Lin L, Chen G, Wu J, Zheng X. Maternal serum vitamin B12, folate and homocysteine and the risk of neural tube defects in the offspring in a high-risk area of China. Public Health Nutr. 2009;12(5):680–6. doi: 10.1017/S1368980008002735. [DOI] [PubMed] [Google Scholar]
- 14.Suarez L, Hendricks K, Felkner M, Gunter E. Maternal serum B12 levels and risk for neural tube defects in a Texas-Mexico border population. Ann Epidemiol. 2003;13(2):81–8. doi: 10.1016/s1047-2797(02)00267-3. [DOI] [PubMed] [Google Scholar]
- 15.Groenen PM, van Rooij IA, Peer PG, Gooskens RH, Zielhuis GA, Steegers-Theunissen RP. Marginal maternal vitamin B12 status increases the risk of offspring with spina bifida. Am J Obstet Gynecol. 2004;191(1):11–7. doi: 10.1016/j.ajog.2003.12.032. [DOI] [PubMed] [Google Scholar]
- 16.Candito M, Rivet R, Herbeth B, Boisson C, Rudigoz RC, Luton D, Journel H, Oury JF, Roux F, Saura R, Vernhet I, Gaucherand P, Muller F, Guidicelli B, Heckenroth H, Poulain P, Blayau M, Francannet C, Roszyk L, Brustie C, Staccini P, Gerard P, Fillion-Emery N, Gueant-Rodriguez RM, Van Obberghen E, Gueant JL. Nutritional and genetic determinants of vitamin B and homocysteine metabolisms in neural tube defects: a multicenter case-control study. Am J Med Genet A. 2008;146A(9):1128–33. doi: 10.1002/ajmg.a.32199. [DOI] [PubMed] [Google Scholar]
- 17.Molloy AM, Kirke PN, Troendle JF, Burke H, Sutton M, Brody LC, Scott JM, Mills JL. Maternal vitamin B12 status and risk of neural tube defects in a population with high neural tube defect prevalence and no folic Acid fortification. Pediatrics. 2009;123(3):917–23. doi: 10.1542/peds.2008-1173. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Gardiki-Kouidou P, Seller MJ. Amniotic fluid folate, vitamin B12 and transcobalamins in neural tube defects. Clin Genet. 1988;33(6):441–8. doi: 10.1111/j.1399-0004.1988.tb03478.x. [DOI] [PubMed] [Google Scholar]
- 19.Dawson EB, Evans DR, Van Hook JW. Amniotic fluid B12 and folate levels associated with neural tube defects. Am J Perinatol. 1998;15(9):511–4. doi: 10.1055/s-2007-993975. [DOI] [PubMed] [Google Scholar]
- 20.Economides DL, Ferguson J, Mackenzie IZ, Darley J, Ware II, Holmes-Siedle M. Folate and vitamin B12 concentrations in maternal and fetal blood, and amniotic fluid in second trimester pregnancies complicated by neural tube defects. Br J Obstet Gynaecol. 1992;99(1):23–5. doi: 10.1111/j.1471-0528.1992.tb14386.x. [DOI] [PubMed] [Google Scholar]
- 21.Steen MT, Boddie AM, Fisher AJ, Macmahon W, Saxe D, Sullivan KM, Dembure PP, Elsas LJ. Neural-tube defects are associated with low concentrations of cobalamin (vitamin B12) in amniotic fluid. Prenat Diagn. 1998;18(6):545–55. [PubMed] [Google Scholar]
- 22.Weekes EW, Tamura T, Davis RO, Birch R, Vaughn WH, Franklin JC, Barganier C, Cosper P, Finley SC, Finley WH. Nutrient levels in amniotic fluid from women with normal and neural tube defect pregnancies. Biol Neonate. 1992;61(4):226–31. doi: 10.1159/000243747. [DOI] [PubMed] [Google Scholar]
- 23.Ray JG, Wyatt PR, Thompson MD, Vermeulen MJ, Meier C, Wong PY, Farrell SA, Cole DE. Vitamin B12 and the risk of neural tube defects in a folic-acid-fortified population. Epidemiology. 2007;18(3):362–6. doi: 10.1097/01.ede.0000257063.77411.e9. [DOI] [PubMed] [Google Scholar]
- 24.Afman LA, Lievers KJ, van der Put NM, Trijbels FJ, Blom HJ. Single nucleotide polymorphisms in the transcobalamin gene: relationship with transcobalamin concentrations and risk for neural tube defects. Eur J Hum Genet. 2002;10(7):433–8. doi: 10.1038/sj.ejhg.5200830. [DOI] [PubMed] [Google Scholar]
- 25.Quadros EV, Regec AL, Khan KM, Quadros E, Rothenberg SP. Transcobalamin II synthesized in the intestinal villi facilitates transfer of cobalamin to the portal blood. Am J Physiol. 1999;277(1):G161–6. doi: 10.1152/ajpgi.1999.277.1.G161. Pt 1. [DOI] [PubMed] [Google Scholar]
- 26.Pietrzyk JJ, Bik-Multanowski M. 776C>G polymorphism of the transcobalamin II gene as a risk factor for spina bifida. Mol Genet Metab. 2003;80(3):364. doi: 10.1016/S1096-7192(03)00131-8. [DOI] [PubMed] [Google Scholar]
- 27.Swanson DA, Pangilinan F, Mills JL, Kirke PN, Conley M, Weiler A, Frey T, Parle-McDermott A, O'Leary VB, Seltzer RR, Moynihan KA, Molloy AM, Burke H, Scott JM, Brody LC. Evaluation of transcobalamin II polymorphisms as neural tube defect risk factors in an Irish population. Birth Defects Res A Clin Mol Teratol. 2005;73(4):239–44. doi: 10.1002/bdra.20122. [DOI] [PubMed] [Google Scholar]
- 28.De Marco P, Merello E, Calevo MG, Mascelli S, Raso A, Cama A, Capra V. Evaluation of a methylenetetrahydrofolate-dehydrogenase 1958G>A polymorphism for neural tube defect risk. J Hum Genet. 2006;51(2):98–103. doi: 10.1007/s10038-005-0329-6. [DOI] [PubMed] [Google Scholar]
- 29.O'Leary VB, Mills JL, Pangilinan F, Kirke PN, Cox C, Conley M, Weiler A, Peng K, Shane B, Scott JM, Parle-McDermott A, Molloy AM, Brody LC. Analysis of methionine synthase reductase polymorphisms for neural tube defects risk association. Mol Genet Metab. 2005;85(3):220–7. doi: 10.1016/j.ymgme.2005.02.003. [DOI] [PubMed] [Google Scholar]
- 30.Boyles AL, Billups AV, Deak KL, Siegel DG, Mehltretter L, Slifer SH, Bassuk AG, Kessler JA, Reed MC, Nijhout HF, George TM, Enterline DS, Gilbert JR, Speer MC. Neural tube defects and folate pathway genes : family-based association tests of gene-gene and gene-environment interactions. Environ Health Perspect. 2006;114(10):1547–52. doi: 10.1289/ehp.9166. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Dunlevy LP, Chitty LS, Burren KA, Doudney K, Stojilkovic-Mikic T, Stanier P, Scott R, Copp AJ, Greene ND. Abnormal folate metabolism in foetuses affected by neural tube defects. Brain. 2007;130:1043–9. doi: 10.1093/brain/awm028. Pt 4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Brouns R, Ursem N, Lindemans J, Hop W, Pluijm S, Steegers E, Steegers-Theunissen R. Polymorphisms in genes related to folate and cobalamin metabolism and the associations with complex birth defects. Prenat Diagn. 2008;28(6):485–93. doi: 10.1002/pd.2006. [DOI] [PubMed] [Google Scholar]
- 33.Pietrzyk JJ, Bik-Multanowski M, Sanak M, Twardowska M. Polymorphisms of the 5,10-methylenetetrahydrofolate and the methionine synthase reductase genes as independent risk factors for spina bifida. J Appl Genet. 2003;44(1):111–3. [PubMed] [Google Scholar]
- 34.van der Linden IJ, den Heijer M, Afman LA, Gellekink H, Vermeulen SH, Kluijtmans LA, Blom HJ. The methionine synthase reductase 66A>G polymorphism is a maternal risk factor for spina bifida. J Mol Med. 2006;84(12):1047–54. doi: 10.1007/s00109-006-0093-x. [DOI] [PubMed] [Google Scholar]
- 35.Doolin MT, Barbaux S, McDonnell M, Hoess K, Whitehead AS, Mitchell LE. Maternal genetic effects, exerted by genes involved in homocysteine remethylation, influence the risk of spina bifida. Am J Hum Genet. 2002;71(5):1222–6. doi: 10.1086/344209. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Zhu H, Wicker NJ, Shaw GM, Lammer EJ, Hendricks K, Suarez L, Canfield M, Finnell RH. Homocysteine remethylation enzyme polymorphisms and increased risks for neural tube defects. Mol Genet Metab. 2003;78(3):216–21. doi: 10.1016/s1096-7192(03)00008-8. [DOI] [PubMed] [Google Scholar]
- 37.Gueant-Rodriguez RM, Rendeli C, Namour B, Venuti L, Romano A, Anello G, Bosco P, Debard R, Gerard P, Viola M, Salvaggio E, Gueant JL. Transcobalamin and methionine synthase reductase mutated polymorphisms aggravate the risk of neural tube defects in humans. Neurosci Lett. 2003;344(3):189–92. doi: 10.1016/s0304-3940(03)00468-3. [DOI] [PubMed] [Google Scholar]
- 38.Franke B, Vermeulen SH, Steegers-Theunissen RP, Coenen MJ, Schijvenaars MM, Scheffer H, den Heijer M, Blom HJ. An association study of 45 folate-related genes in spina bifida: Involvement of cubilin (CUBN) and tRNA aspartic acid methyltransferase 1 (TRDMT1) Birth Defects Res A Clin Mol Teratol. 2009;85(3):216–26. doi: 10.1002/bdra.20556. [DOI] [PubMed] [Google Scholar]
- 39.Quadros EV, Nakayama Y, Sequeira JM. The protein and the gene encoding the receptor for the cellular uptake of transcobalamin-bound cobalamin. Blood. 2009;113(1):186–92. doi: 10.1182/blood-2008-05-158949. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Shields DC, Kirke PN, Mills JL, Ramsbottom D, Molloy AM, Burke H, Weir DG, Scott JM, Whitehead AS. The "thermolabile" variant of methylenetetrahydrofolate reductase and neural tube defects: An evaluation of genetic risk and the relative importance of the genotypes of the embryo and the mother. Am J Hum Genet. 1999;64(4):1045–55. doi: 10.1086/302310. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Brody LC, Conley M, Cox C, Kirke PN, McKeever MP, Mills JL, Molloy AM, O'Leary VB, Parle-McDermott A, Scott JM, Swanson DA. A polymorphism, R653Q, in the trifunctional enzyme methylenetetrahydrofolate dehydrogenase/methenyltetrahydrofolate cyclohydrolase/formyltetrahydrofolate synthetase is a maternal genetic risk factor for neural tube defects: report of the Birth Defects Research Group. Am J Hum Genet. 2002;71(5):1207–15. doi: 10.1086/344213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Pangilinan F, Geiler K, Dolle J, Troendle J, Swanson DA, Molloy AM, Sutton M, Conley M, Kirke PN, Scott JM, Mills JL, Brody LC. Construction of a high resolution linkage disequilibrium map to evaluate common genetic variation in TP53 and neural tube defect risk in an Irish population. Am J Med Genet A. 2008;146A(20):2617–25. doi: 10.1002/ajmg.a.32504. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Meulenbelt I, Droog S, Trommelen GJ, Boomsma DI, Slagboom PE. High-yield noninvasive human genomic DNA isolation method for genetic studies in geographically dispersed families and populations. Am J Hum Genet. 1995;57(5):1252–4. [PMC free article] [PubMed] [Google Scholar]
- 44.The International HapMap Project. Nature. 2003;426(6968):789–96. doi: 10.1038/nature02168. [DOI] [PubMed] [Google Scholar]
- 45.Barrett JC, Fry B, Maller J, Daly MJ. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics. 2005;21(2):263–5. doi: 10.1093/bioinformatics/bth457. [DOI] [PubMed] [Google Scholar]
- 46.Stephens M, Smith NJ, Donnelly P. A new statistical method for haplotype reconstruction from population data. Am J Hum Genet. 2001;68(4):978–89. doi: 10.1086/319501. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Stephens M, Donnelly P. A comparison of bayesian methods for haplotype reconstruction from population genotype data. Am J Hum Genet. 2003;73(5):1162–9. doi: 10.1086/379378. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Weinberg CR, Wilcox AJ, Lie RT. A log-linear approach to case-parent-triad data: assessing effects of disease genes that act either directly or through maternal effects and that may be subject to parental imprinting. Am J Hum Genet. 1998;62(4):969–78. doi: 10.1086/301802. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Spielman RS, McGinnis RE, Ewens WJ. Transmission test for linkage disequilibrium: the insulin gene region and insulin-dependent diabetes mellitus (IDDM) Am J Hum Genet. 1993;52(3):506–16. [PMC free article] [PubMed] [Google Scholar]
- 50.Dempster A, Laird NM, Rubin DB. Maximum likelihood from incomplete data via EM algorithm. Journal of the Royal Statistical Society, Series B. 1977;39(1):1–38. [Google Scholar]
- 51.Kent WJ, Sugnet CW, Furey TS, Roskin KM, Pringle TH, Zahler Am, Haussler D. The human genome browser at UCSC. Genome Res. 2002;12(6):996–1006. doi: 10.1101/gr.229102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.May P, Woldt E, Matz RL, Boucher P. The LDL receptor-related protein (LRP) family: an old family of proteins with new physiological functions. Ann Med. 2007;39(3):219–28. doi: 10.1080/07853890701214881. [DOI] [PubMed] [Google Scholar]