Figure 1.
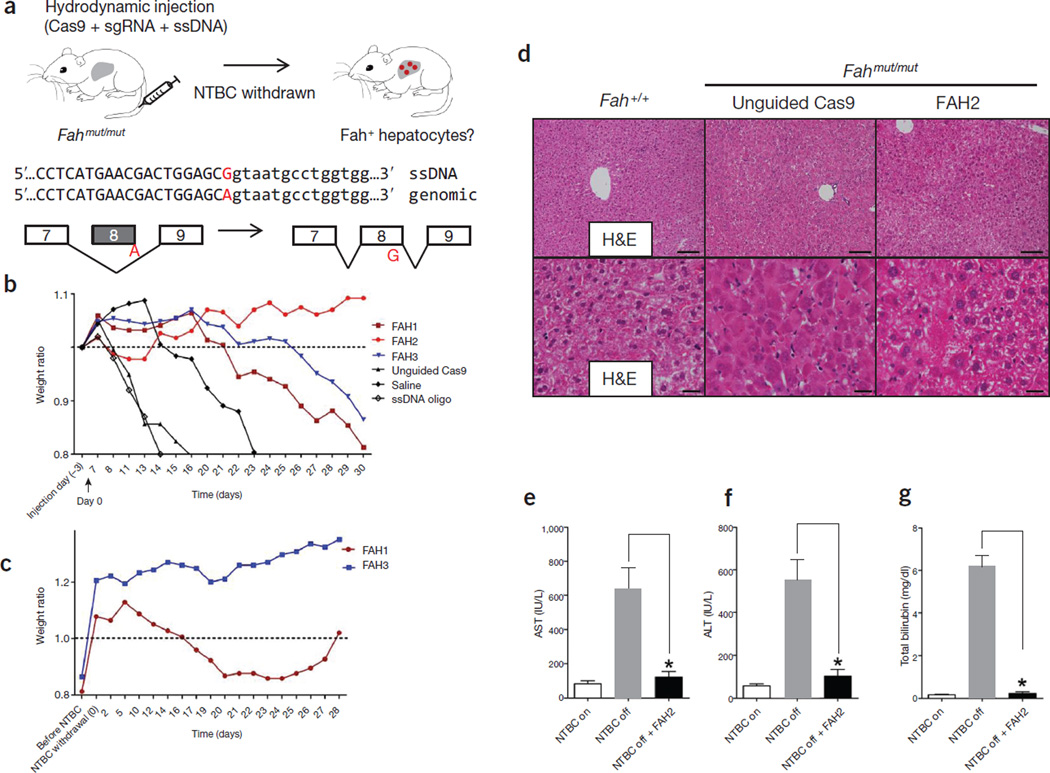
Hydrodynamic injection of CRISPR components rescues lethal phenotype of Fah-deficient mice. (a) Experimental design. Fahmut/mut mice harbor a homozygous G→A point mutation at the last nucleotide of exon 8 (red), causing skipping of exon 8 during splicing. pX330 plasmids expressing Cas9 and a sgRNA targeting the Fah locus are delivered to the liver by hydrodynamic tail vein injection. A ssDNA oligo with the correct fragment of Fah sequence (i.e., the G allele) is co-injected to serve as a donor template to repair the ‘A’ mutation. Exon and intron sequences are in upper and lower cases, respectively. (b) Fahmut/mut mice were injected with saline only, ssDNA oligo only, ssDNA oligo plus pX330 (unguided Cas9), or ssDNA oligo plus pX330 expressing Cas9 and one of the three Fah sgRNAs (FAH1, FAH2 and FAH3). Body weight was monitored over time and normalized to pre-injection weight. Arrow indicates withdrawal of NTBC water (defined as day 0, which is 3 d after injection). (c) Mice injected with FAH1 or FAH3 in b were put back on NTBC water for 7 d and then again withdrawn from NTBC for 28 d. (d) H&E staining of liver sections from wild-type (Fah+/+) or Fahmut/mut mice injected with unguided Cas9 or Cas9 with the FAH2 sgRNA and kept off NTBC water. The FAH2 sample is from a mouse 30 d after NTBC withdrawal. Scale bars, 100 µm for upper panels, 20 µm for lower panels. (e–g) Liver damage markers (aspartate aminotransferase (AST), alanine aminotransferase (ALT) and bilirubin) were measured in peripheral blood from Fahmut/mut mice injected with saline or ssDNA oligo only or unguided Cas9 (NTBC off) or FAH2 (NTBC off + FAH2, day 30). Fahmut/mut mice on NTBC water (NTBC on) served as a control. * P < 0.01 (n = 3 mice) using one-way ANOVA. Error bars, mean ± s.e.m.