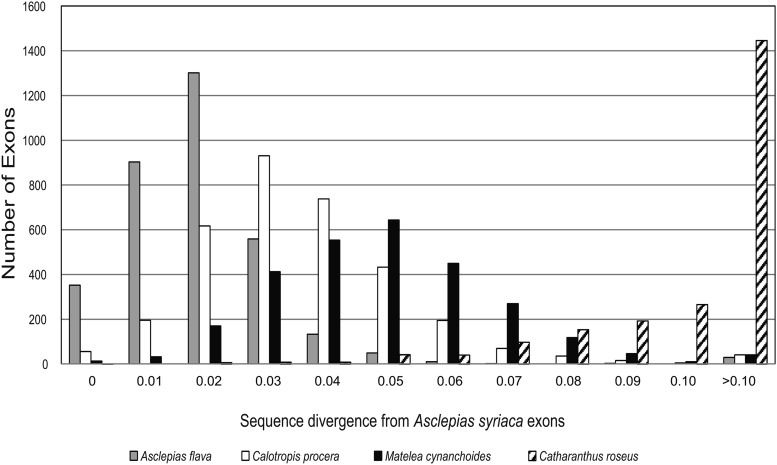
Fig. 1.
Histogram of exon sequence divergence between the species used for probe design, Asclepias syriaca, and four other species: the most divergent species of Asclepias, A. flava; another member of Asclepiadinae (Asclepiadeae: Asclepiadoideae), Calotropis procera; a member of Gonolobinae (Asclepiadeae: Asclepiadoideae), Matelea cynanchoides; and a member of a different subfamily, Catharanthus roseus (Rauvolfioideae). Note that a maximum sequence divergence of 75% was allowed for BLAT and that exons with >10% divergence were less likely to be observed in Calotropis and Matelea because they were less likely to be enriched by the probes, while the Catharanthus data were from transcriptome sequences of multiple tissues and not subject to target enrichment bias.