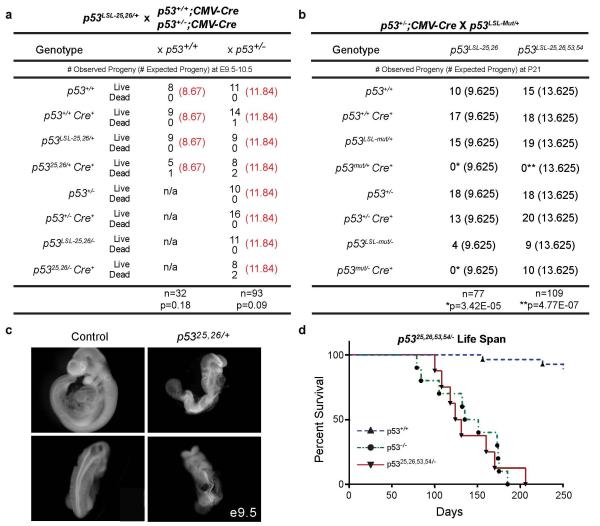
Extended Data Figure 2. p5325,26,53,54/− Mice, but not p5325,26/+ or p5325,26/− Mice, are Viable.
(a) Crosses of p53LSL-25,26/+ with p53+/+;CMV-Cre or p53+/−;CMV-Cre mice reveal a decrease in viable pups expressing p5325,26 at E9.5-E10.5. Observed numbers of live and dead pups compared to the expected numbers of live pups are indicated. [Observed (Expected)] The genotypes of p5325,26/+ and p5325,26/− mice carrying a CMV-Cre transgene lack the LSL designation because the Lox-Stop-Lox element has been deleted from the genome. Significance as assessed by Binomial distribution statistical tests on live pups: p=0.18 and 0.09 (b) Crosses of p53LSL-25,26/+ or p53LSL-25,26,53,54/+ with p53+/−;CMV-Cre mice reveal that p5325,26,53,54/− mice, but not p5325,26/− mice, are viable as assessed at postnatal day 21 (P21). Mut denotes either mutant allele. Observed numbers of pups compared to the expected numbers of pups are indicated. [Observed (Expected)] The genotypes of p53mut/+ and p53mut/− mice carrying a CMV-Cre transgene lack the LSL designation because the Lox-Stop-Lox element has been deleted from the genome. Lack of pups is significant at P21 as assessed by Binomial distribution statistical tests on live pups: p5325,26/+ and p5325,26/−: *p<=3.42E-05, p5325,26,53,54/+: **p=4.77E-07 (c) Whole-mount images of a p5325,26/+ embryo (right) at E9.5 displaying developmental delay (top) and neural tube defects, including exencephaly and kinked spine (bottom), compared to littermate control (left). (d) p5325,26,53,54/− mice display a shortened lifespan (median lifespan 128 days, n=8) compared to wild-type mice (median lifespan 774 days) and a similar lifespan to p53−/− mice (median lifespan 143 days), further indicating that the p5325,26,53,54 allele itself behaves like a p53 null allele. p<0.0001 by Mantel-Cox statistical analysis comparing wild-type and p53 25,26,53,54/− mice