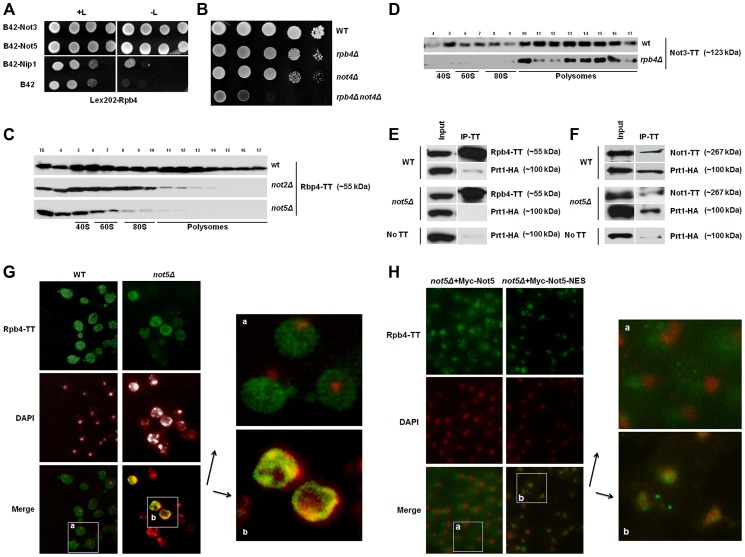
Figure 1. Rpb4 interacts with Not5 and the cytoplasmic functions of Rpb4 require Not5.
A. Serial dilutions of exponentially growing reporter cells expressing LexA-Rpb4 as a bait, and the indicated proteins fused to B42 as preys, were spotted either on medium selective for the plasmids (left panel +L) or selective for the plasmids and indicative of an interaction between bait and prey (right panel -L). B. Serial dilutions of exponentially growing cells from the indicated strains were spotted on plates and left to grow for several days at 30°C. C and D. Fractions from 7–47% sucrose gradients of extracts from wild-type or mutant strains expressing the indicated Tap-tagged (TT) proteins were precipitated with TCA and analyzed by western blotting with PAP antibodies. The positions of 40S, 60S, 80S and polysomes are indicated under the blots. The numbers of the gradient fractions tested or the total extract (TE) are indicated at the top. The polysome profiles for these experiments are available in Fig. S15 along with a typical distribution of a ribosomal protein (Rps3) in the wt and not5Δ gradients. Rpb4-TT (E) or Not1-TT (F) were immunoprecipitated from extracts of wild-type or mutant cells expressing HA-tagged Prt1. Wild-type cells expressing untagged Rpb4 or Not1 were used as a control. Similar negative controls were obtained with not5Δ cells not expressing any Tap-tagged protein (Fig. S16). The immunoblots were developed using anti-CBP or HA antibodies. G and H. Wild-type and not5Δ cells expressing Rpb4-TT (G) or the indicated (H) Not5 derivatives, were grown exponentially and stained with anti-CBP antibodies (upper panels) or DAPI (middle panels). The pictures were merged (lower panels) and the indicated section from wild-type (a) or not5Δ (b) was enlarged for better visualization. The localization of the Not5 derivatives is presented in Fig. S2.