Figure 2.
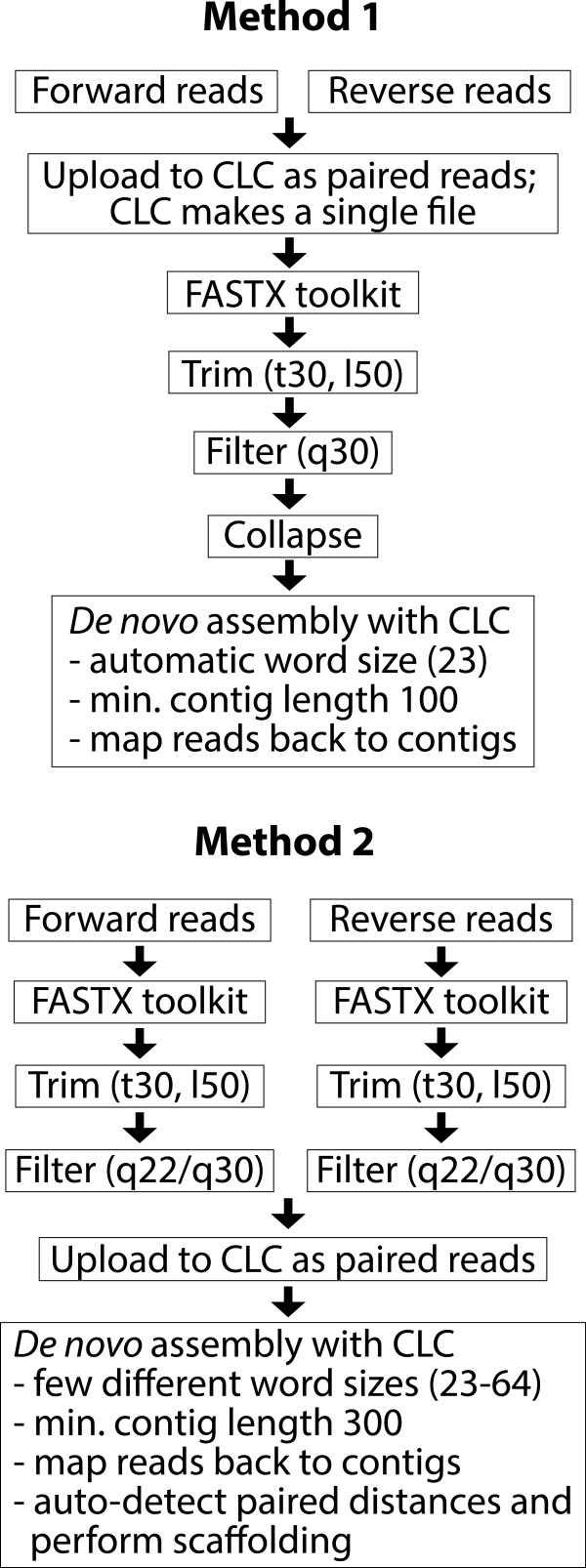
Flow diagram showing the two different assembly methods used. In the first method raw reads were combined before trimming, filtering, and collapsing. Then they were assembled using CLC Genomics using the automatic word size (23) set by the program indicating the minimum contig length as 100. In the second method forward and reverse reads were trimmed and filtered individually. High or low quality filtering parameters were tried. Trimmed and filtered reads were assembled using CLC Genomics. Several different word sizes were tried and the minimum contig length was set to 300. Paired distances were automatically detected by the program and scaffolding was performed.
