Figure 7. The human SAS-1 homolog C2CD3 is impaired by mutations important for SAS-1 function.
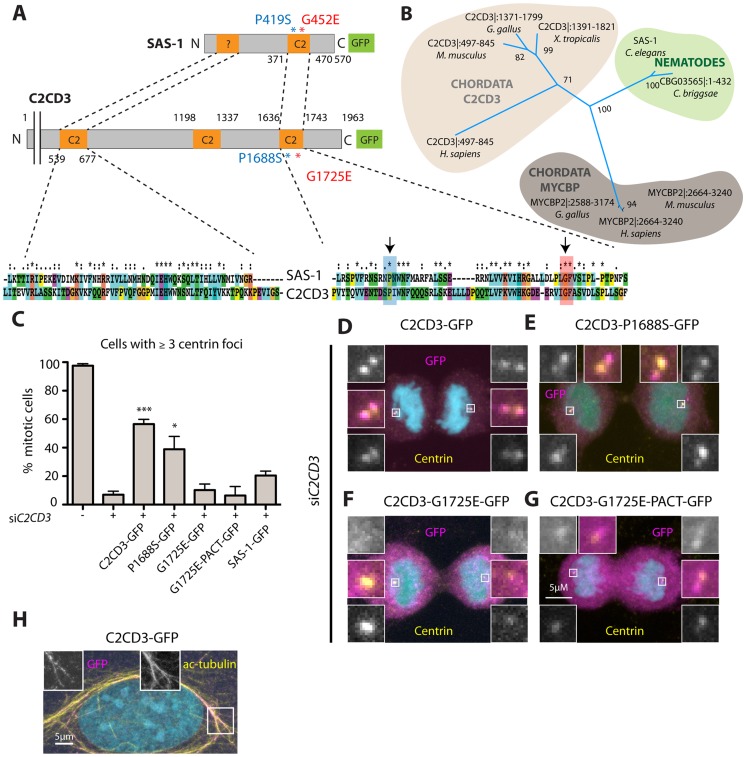
(A) Schematics and alignment of the most homologous regions of C. elegans SAS-1 and human C2CD3. The arrows indicate the two sas-1 mutations. Asterisks in the alignment indicate identity, colons strongly similar properties, dots weakly similar properties. Among the two proteins, the N-terminal C2 domain (C2CD3) and related region (SAS-1) share ∼30% identity, the C-terminal C2 domains ∼33% identity. (B) Phylogenetic tree based on PSI-BLAST analysis performed with full length SAS-1. The numbers indicate support values for the respective tree branches. Only selected species are shown; note that although widely present across metazoan evolution, a homolog was not identified in Drosophila. Note that we also identified a homology with MYCBP. However, when performing PSI-BLAST analysis with the N-terminus only, MYCBP did not come up, in contrast to C2CD3. (C) U2OS cells expressing the indicated fusion proteins were subjected to siC2CD3, stained for Centrin and GFP; the number of Centrin foci in the resulting mitotic cells expressing GFP were scored. Data from ≥3 experiments;>25 cells were counted in each condition, except for two experiments with G1725E PACT, where only very few cells (7 and 4, respectively) were positive for GFP at centrioles. Error bars indicate SEM, *** p<0.001, * p<0.05. (D–G) Mitotic cells depleted of C2CD3 and expressing C2CD3-GFP (D) or the indicated proteins harboring single amino acid substitutions corresponding to the sas-1 mutant alleles (E–G) stained for GFP (magenta), centrin (yellow). DNA is in cyan. (H) An interphase cell expressing C2CD3-GFP showing microtubule co-localization of GFP (magenta) and acetylated tubulin (yellow).