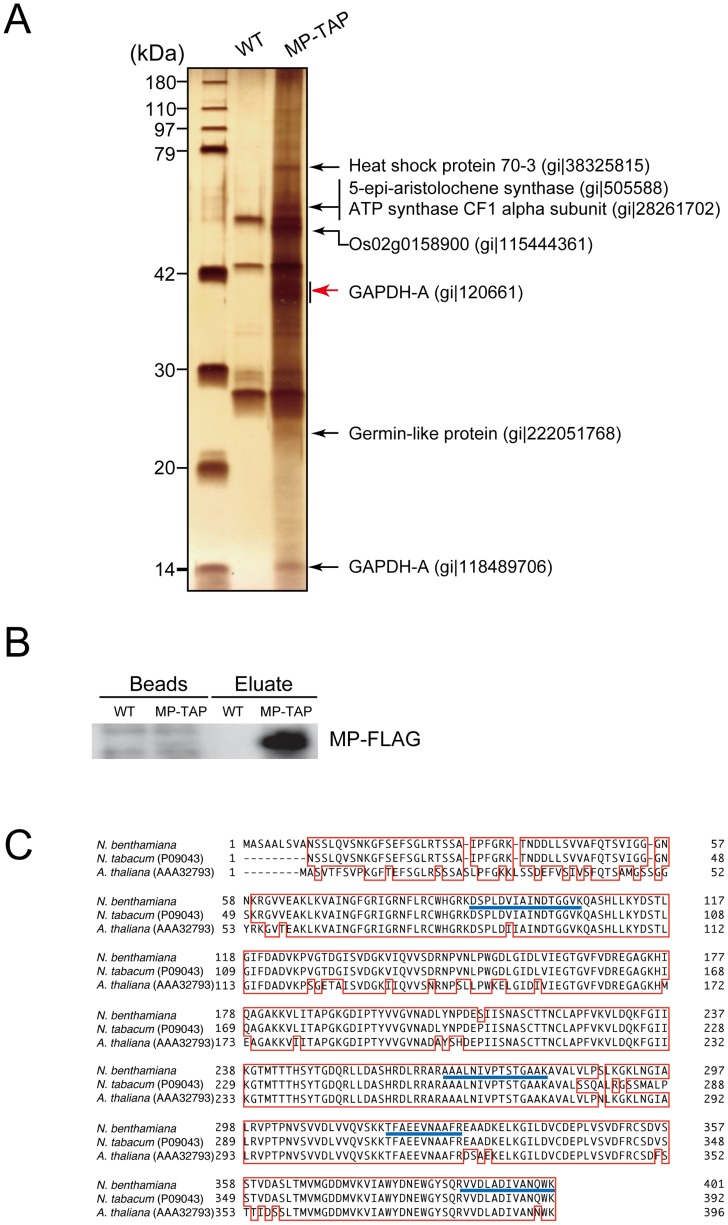
Figure 2. Identification of proteins that were copurified with RCNMV MP.
Protein extracts prepared from Agrobacterium-infiltrated leaves that expressed RCNMV RNA1 plus RNA2 (WT, pBICR12, Figure S1C), or RNA1 plus recombinant RNA2 encoding MP tagged with TAP tag sequences (MP-TAP, pBICR12/MP-TAP, Figure S1D), were subjected to two-step affinity purification using an anti-HA antibody followed by an anti-FLAG antibody. (A) One fifth of the affinity-purified fractions eluted from the anti-FLAG beads (Eluate) and the proteins contained with the beads after elution (Beads) were subjected to Western blotting using anti-FLAG antibody. (B) Four fifths of the affinity-purified fractions were subjected to SDS-PAGE and stained using MS-compatible silver staining. The protein bands of interests were excised, subjected to in-gel digestion, and analyzed by tandem mass spectrometry. Proteins with Mascot search scores >50, which were absent from the control protein bands, and proteins with significantly higher scores than the control proteins are indicated on the right-hand sides of the panels. The NCBI accession numbers of the identified proteins are also indicated. (C) The GAPDH-A gene of N. benthamiana was cloned and the deduced amino acid sequence was aligned using GENETYX-Mac ver. 14.0.1 with its closest homolog from N. tabacum (accession number P09043) and that from Arabidopsis thaliana (accession number AAA32793). The blue lines show the peptide sequences detected.