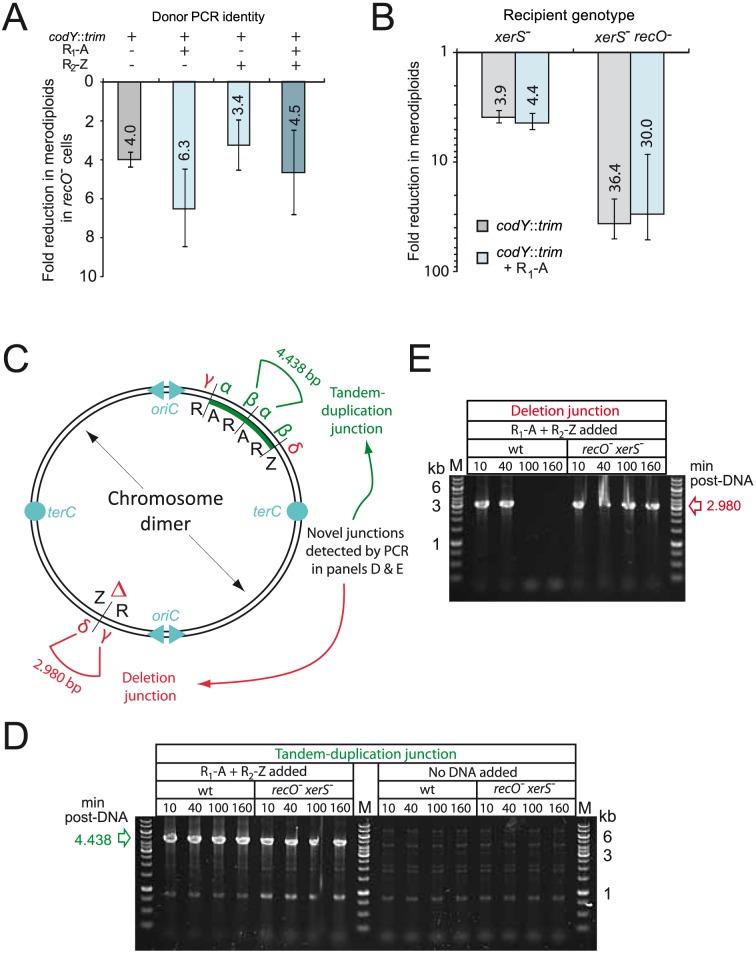
Figure 3. RecO and XerS are involved in pneumococcal chromosome dimer resolution.
(A) Fold reduction in merodiploids formed in recO mutant compared to wildtype cells. Merodiploid formation was triggered using the R1-A PCR fragment as donor and the codY::trim donor cassette was used to score merodiploid transformants. Recipient strains used: wildtype, R246; recO-, R3170. (B) Fold reduction in merodiploids formed in xerS and recO xerS mutant compared to wildtype cells. Same experimental set up as in panel A. Recipient strains used: xerS-, R3214; xerS- recO-; R3873. Since merodiploid frequency was low in the double mutant, representing few clones, we confirmed on a few randomly chosen clones that they were indeed merodiploid by detection of the tandem-duplication junction by PCR (panel D), as well as confirming that they had retained the antibiotic sensitivities associated with the parent. (C) Schematic of chromosome dimer and novel tandem duplication and deletion junctions detected by PCR. α and β represent primers used to detect the novel junction created by tandem duplication. δ and γ represent primers used to detect the deletion junction created as a consequence of tandem duplication. Both junctions are present on the chromosome dimer, while the merodiploid chromosome possesses only the tandem-duplication junction, and the abortive chromosome possesses only the deletion junction. (D) Detection of tandem-duplication junction on cultures of wildtype or recO- xerS- cells transformed with either R1-A and R2-Z PCR fragments or no DNA. PCRs carried out at different time-points after DNA addition with primers CJ242 and merod-b to amplify a 4,438 bp DNA fragment diagnostic of the tandem duplication. (E) Detection of deletion junction on same cells as in panel A (DNA added only). PCRs carried out with primers CJ244 and CJ245 to amplify a 2,980 bp DNA fragment diagnositic of the deletion junction. Strains used in both panels: wildtype; R246, recO- xerS-; R3873.