Figure 2.
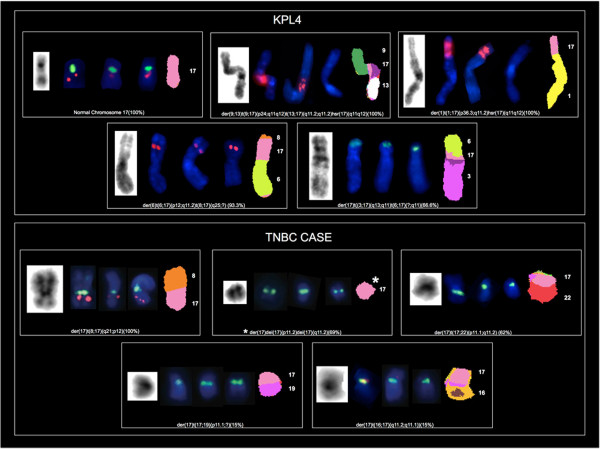
Analysis of Chr17 using G-Banding, dual-color FISH ( HER2 /CEP17 , STARD3 /CEP17 and TOP2A /CEP17) and M-FISH in KPL4 HER2 amplified breast cancer cell line showing four translocated Chr17 in addition to the normal-appearing copies of Chr17 and in one triple negative breast cancer case (TNBC) showing five rearranged copies of Chr17. Rearranged chromosomes containing a portion of Chr17 are visualized by G-Banding technique on the left and by M-FISH on the right. For M-FISH the classified color of Chr17 is shown in pink, the translocation partners are numbered on the right hand side of the chromosomes and the frequency at which each abnormality was observed is indicated in brackets at the end of each abnormality. CEP17, HER2, STARD3 and TOP2A are shown in the middle by dual-color FISH (HER2/CEP17, STARD3/CEP17, TOP2A/CEP17, respectively) whenever mapped to the corresponding derivatives (CEP17 is green-labeled; HER2, STARD3 and TOP2A genes are red-labeled). In the TNBC cells the chromosome in which we identified Chr17 material only is a der(17)del(17)(p11.2)del(17)(q11.2) with a deletion on both short and long arm involving 17q12-q21.
