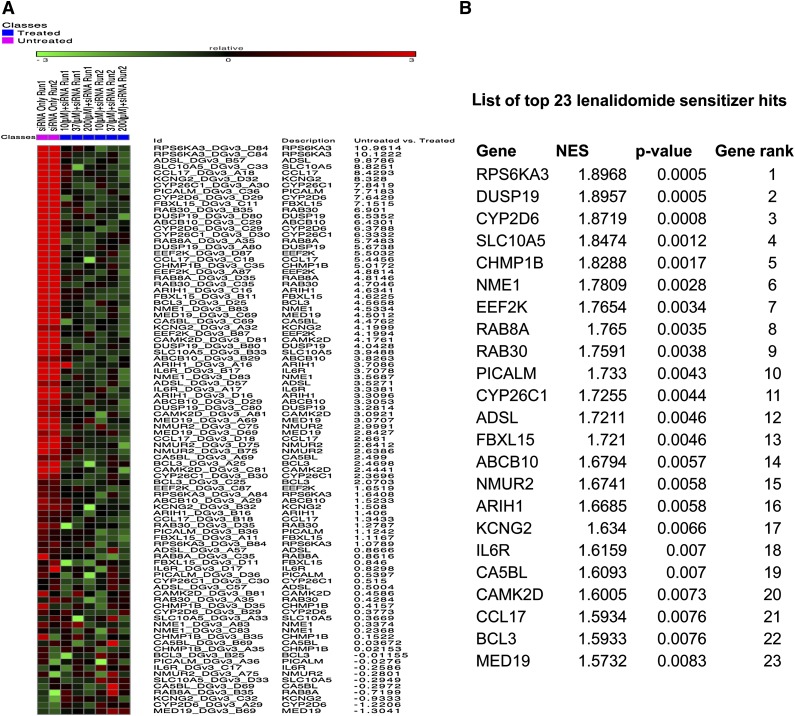
Figure 1.
Top ranking lenalidomide sensitizers identified from druggable genome screens. The data collected from each siRNA were normalized and divided into untreated (siRNA only) and treated (siRNA+drug) classes, followed by analysis using the RIGER algorithm. When a stringent P value of <.01 was applied, 23 genes were selected as top lenalidomide modulators. (A) The 4 siRNAs targeting the top 23 sensitizer hits were ranked using a heat map plot based on their lethal activity changes after lenalidomide treatment. Relative: GENE-E converts values to heat map colors using the mean and maximum values for each row or the standard deviations from the row mean for each row (Source: https://www.broadinstitute.org/cancer/software/GENE-E/doc.html). “Untreated vs treated” are scores generated by the RIGER method for each siRNA. It is a score that is based on siRNA only (untreated) vs siRNA+lenalidomide (treated). (B) A list of top ranked genes with NES and P values are shown.