Abstract
Cell cycle phase is a critical determinant of the choice between DNA damage repair by non-homologous end joining (NHEJ) or homologous recombination (HR). Here we report that DSBs induce ATM-dependent MOF (a histone H4 acetyl-transferase) phosphorylation (p-T392-MOF) and that phosphorylated MOF co-localizes with γ-H2AX, ATM, and 53BP1 foci. Mutation of the phosphorylation site (MOF-T392A) impedes DNA repair in S- and G2-phase but not G1-phase cells. Expression of MOF-T392A also reverses the reduction in DSB associated 53BP1 seen in wild type S/G2-phase cells, resulting in enhanced 53BP1 and reduced BRCA1 association. Decreased BRCA1 levels at DSB sites correlates with defective repairosome formation, reduced HR repair and decreased cell survival following irradiation. These data support a model whereby ATM mediated MOF-T392 phosphorylation modulates 53BP1 function to facilitate the subsequent recruitment of HR repair proteins, uncovering a regulatory role for MOF in DSB repair pathway choice during S/G2-phase.
Keywords: MOF phosphorylation, 53BP1, DSB repair, NHEJ, HR
Introduction
The MOF (males absent on the first) MYST family member affects ATM function in response to DNA damage and is essential for cell viability, proliferation, autophagy and oncogenesis (Bhadra et al., 2012; Fullgrabe et al., 2013; Gupta et al., 2005; Li et al., 2010; Sharma et al., 2010; Taipale et al., 2005). MOF acetylates the Lysine-16 residue of histone H4 (H4K16), and this modification is critical for the DNA damage response (DDR), as depletion of MOF abrogates repair of double-strand breaks (DSBs) by both the NHEJ and HR pathways (Sharma et al., 2010). Highly up-regulated in the S and G2-phase, DNA DSB repair begins with the nucleolytic resection of the 5’ strands of the DNA break ends to generate 3’ single-stranded DNA. Although NHEJ is also functional in cells at G2 phase (Kakarougkas et al., 2013), the proximity of the appropriate sister chromatid or homologous chromosomes is likely a key regulatory step for preferential initiation of HR. The potential role of chromatin modifying factors in facilitating the HR pathway remains unknown. It is known however that the end resection of DNA DSB to permit repair by HR is activated by cyclin-dependent kinase (CDK) phosphorylation of CtIP (Sartori et al., 2007), the 5’ DNA resection nuclease in S/G2. In G1 this process is blocked by 53BP1 (also called TP53BP1) (Bothmer et al., 2010), a well-known DDR factor that is recruited to nuclear structures at DNA damage sites forming readily visualized IR-induced foci. Depletion of 53BP1 results in genomic instability in human as well as mouse cells and its absence also results in cell-cycle arrest in G2/M phase. The 53BP1 protein is classified as an adaptor/mediator, as it is required for processing of the DDR signal and as a platform for recruiting additional repair factors involved in both DSB repair and cell-cycle control. It has been proposed that the key determinant in choosing the preferred DSB repair pathway lies in the initial processing of the DSB ends, which is regulated during the cell cycle (Symington and Gautier, 2011). For instance, in cells at G1 phase, 53BP1 inhibits DNA end-resection of DSBs, thus favoring repair by the NHEJ pathway. However, in S/G2 phase, end-resection of DNA DSB is initiated at the 5’ DSB to produce a 3’ single-stranded overhang that facilitates HR-mediated repair (Bothmer et al., 2011; Bothmer et al., 2010; Symington and Gautier, 2011; West, 2003). Thus, 53BP1 is a critical determinant in choosing the appropriate DSB repair pathway in G1- and S/G2-phase cells (Chapman et al., 2012).
Inhibition of DNA end-resection by 53BP1 during the G1-phase is required (Anbalagan et al., 2011; Bonetti et al., 2010) in order to shift the repair pathway to the NHEJ pathway (Morales et al., 2003; Nakamura et al., 2006; Oksenych et al., 2012; Ward et al., 2003; Zimmermann et al., 2013), since a homologous DNA template is not available to carry out HR-dependent repair. Conversely, inhibition of 53BP1 activity by BRCA1 in S/G2 phase cells appears to be critical since there is increased genomic instability in BRCA1-deficient cells due to 53BP1-dependent NHEJ repair during S/G2, as a result of resection suppression (Bunting et al., 2010). Callen and colleagues have reported that PTIP (also known as PAXIP) contributes to DNA DSB repair pathway preference, as cells deficient for PTIP are tolerant of DNA damaging agents that are toxic to cells deficient in DSB repair via the HR pathway, arguing that PTIP like 53BP1 play a critical role in NHEJ (Callen et al., 2013; Escribano-Diaz and Durocher, 2013).
The cellular response to IR and DNA DSB repair depends on the protein kinase ATM (Matsuoka et al., 2007; Pandita, 2003; Pandita et al., 2000; Shiloh and Ziv, 2013) which is influenced by MOF. In addition to binding chromatin (Sharma et al., 2010), MOF also associates with ATM (Gupta et al., 2005; Sharma et al., 2010). In this study we present evidence that MOF is phosphorylated at the threonine-392 residue (pT392-MOF) by ATM subsequent to IR-induced DNA damage. Phosphorylation of MOF by ATM is functionally significant as it was required to facilitate recruitment of HR-related repair proteins to DNA damage sites during S/G2 phase. Our results provide new mechanistic insights into the potential role of p-T392-MOF in regulating key proteins involved in choosing the appropriate DNA DSB repair pathway.
Results
ATM dependent phosphorylation of MOF
Given the importance of MOF in DNA repair (Gupta et al., 2014; Peterson and Almouzni, 2013; Sharma et al., 2010), we first asked whether IR exposure induces phosphorylation of MOF. Analysis of MOF immunoprecipitated from human cell extracts with a pan-phospho-serine antibody detected basal level of MOF serine phosphorylation that was unchanged after irradiation and unaffected by cellular ATM status (Figure 1A). In contrast, IR-induced MOF threonine phosphorylation in an ATM dependent manner (Figure 1A). Antibody for ATM specific SQ/TQ phosphorylation sites also detected a post-irradiation increase in MOF phosphorylation in ATM expressing cells. KU55933 blocked the increase in MOF SQ/TQ phosphorylation, confirming the ATM-dependence of MOF phosphorylation after IR exposure (Figure 1B). The human and mouse MOF sequences contain only one SQ and one TQ site (Figure S1A). Substitution of MOF threonine 392 with alanine (T392A) resulted in loss of ATM dependent IR-induced phosphorylation. Western blot analysis of immunoprecipitated MOF from control and irradiated cells with a polyclonal rabbit phosphospecific T392-MOF antibody revealed an increase in endogenous pT392-MOF post-irradiation (Figure 1C). Similarly, Flag-tagged MOF expressed in 293 cells also displayed increased Flag-MOF-T392 phosphorylation post irradiation (Figure 1C), while phosphorylation of Flag-tagged MOF-T392A was not detected (Figure 1C). This result confirms that the single MOF TQ site at threonine 392 is the only IR and ATM dependent MOF phosphorylation site (Figure 1A). In an in vitro kinase assay (Gupta et al., 2005; Lee and Paull, 2004) ATM directly phosphorylates MOF (Figure 1D) but not the MOF-T392A mutant (Figure 1D). Post-irradiation phosphorylation of MOF-T392 is also independent of ATR (Figure 1E) further confirming ATM as the responsible kinase (Figure 1E). Cellular levels of MOF protein in G1 and S/G2-phase cells were similar and unaffected by irradiation but the IR-induced increase in pT392-MOF levels was highest in S/G2 phase cells (Figure 1F). Mutation of the ATM phosphorylation site in MOF-T392A had no effect on MOF acetylation activity, whether measured by in vitro histone H4K16 acetylation (Figure 1G and S1B) or MOF autoacetylation in vivo (Figure 1H). This result is consistent with PONDER analysis of the wild type MOF and MOF-T392A protein structures (Obradovic et al., 2005) which indicated no major structural differences introduced the threonine/alanine substitution induced no major structural changes, whereas the phospho-mimic MOF-T392E mutation induced localized structural changes (Figure S1C). Together these results suggest that ATM-dependent phosphorylation may regulate MOF functions.
Figure 1. ATM dependent MOF phosphorylation.
(A) Cells with (AT221JE-TpEBS7-YZ5) (+ATM) and without (AT221JE-TpEBS7) (−ATM) ATM were irradiated, the MOF protein immunoprecipitated and blotted with anti-phospho-Serine (p-Ser), anti-phospho-Threonine (p-Thr) and anti-phospho-Serine Glutamine/Threonine Glutamine (p-SQ/TQ) antibodies. (B) HEK293 cells with and without KU 55933 treatment were irradiated and MOF protein was immunoprecipitated and blotted with anti-pSQ/TQ antibody. (C) HEK293 cells were irradiated and collected at different time points post irradiation: (i) MOF protein was immunoprecipitated with MOF antibody and blotted with polyclonal anti-pT392-MOF antibody raised in rabbit; (ii) Flag-MOF protein was immunoprecipitated by anti-Flag-antibody and blotted with anti-pT392-MOF antibody and (iii) HEK293 cells expressing Flag-MOF-T392A were irradiated, immunoprecipitated by anti-Flag antibody and blotted with anti-pT392-MOF antibody. (D) The ATM kinase assay was as described (Gupta et al., 2005). The GST-MOF and the recombinant MRN complex was expressed in Sf21 insect cells with a baculovirus system and purified by nickel affinity, ion exchange, and gel filtration chromatography (Gupta et al., 2005): (i) Wild type GST-MOF incubated with different combinations and MOF phosphorylation requires ATM and; (ii) Kinase assay performed with wild type and mutant MOF-T392A. (E) Effect of ATR depletion on IR-induced MOF phosphorylation: (i) Knock down of ATR with specific siRNA; (ii) Western blot showing IR-induced MOF phosphorylation in ATR depleted cells. (F) Detection of phospho-MOF post irradiation in different phases of cell cycle: (i) Western blot showing MOF and phospho-MOF in G1 and S/G2 phase enriched cells; (ii) Quantitation of relative p-T392-MOF levels in G1 and S/G2 phase enriched cells (**P<0.001 as determined by the chi-square test). (G) In vitro acetylation activity assay of mutant and wild type MOF. (H) Cells with and without ATM expressing wild type or mutant MOF were irradiated, MOF immunoprecipitated and analyzed for acetylation status by using pan-acetylation antibody.
Role of MOF phosphorylation during S and G2 phases
Expression of MOF-T392A in cells co-transfected with siRNA specific for the endogenous MOF 3’UTR did not result in any major changes in cell viability 96 hr post-transfection (Figure S2A and S2B and data not shown). Despite a reduction of endogenous MOF to nearly undetectable levels (Figure S2A), H4K16ac levels were largely preserved over this time period (Figure S2C) when MOF-T392A was expressed, which is consistent with the in vitro (Figure 1G and S1C) and in vivo (Figure 1H) acetylation results. However, after exposure to IR, there was significantly reduced survival of MOF-siRNA transfected exponentially growing cells, an outcome partially rescued by MOF-T392A expression (Figure 2A). The partial rescue suggested that the role of MOF phosphorylation may be limited to a specific phase(s) of the cell cycle. This possibility was examined further by analysis of synchronized cell populations. Cells with or without expression of MOF-T392A, with and without depletion of endogenous MOF (Figure S2D and S2E) were enriched in G1-phase by serum depletion, in S-phase by thymidine block and in G2/M phase (Table S1) by releasing from thymidine block for measurement of survival post irradiation. Mutant MOF-T392A expression did not alter the survival of irradiated G1-phase cells (Figure 2B). In contrast, S- as well as G2/M-phase cells expressing mutant MOF-T392A with depleted endogenous MOF (Figure S2D and S2E) had reduced cell survival after IR-exposure as compared to cells expressing wild type MOF (Figure 2C and 2D). The survival differences were not due to dominant negative effects as the endogenous MOF was simultaneously depleted by specific siRNA or cre-mediated or a resistant MOF was expressed (Figure S2C, S2D and S3A–C, data not shown). Furthermore, phospho-mimic MOF-T392E expression in cells depleted of endogenous MOF, did not show increased cell killing post-irradiation, neither in exponential phase cells or G1-, S- or G2-phase enriched cells (Figure 2A–D), confirming the role of phospho-MOF in S- and G2-phase cell survival post irradiation.
Figure 2. Mutant MOF abrogates survival and chromosome repair.
Effect of MOF-T392A on survival, chromosome aberrations post irradiation and DSB repair. (A–D) Clonogenic survival after IR dose of 6 Gy, exponential population (A), G1-phase population (B), S-phase population (C) and G2-phase population (D). (E–G): Metaphase chromosome aberration of G1-type (E), S-type (F) and G2-type (G). (H) Cells treated with caffeine prior exposure to IR. About 50 metaphases were scored for each experiment and the data presented is the mean of three experiments. (I) Proliferation of floxed MOF MEF cells with or without tamoxifen treatment and expression of Mof-T392A. Cell numbers were determined at 96 hr post tamoxifen treatment in cells with and without mMof. (J–L): Metaphase chromosome aberration of G1-type (J), S-type (K) and G2-type (L). (M) I-SceI-induced DSBs by NHEJ. (N) Impairment of I-SceI-induced HR in cells expressing MOF-T392A or cells with MOF depletion by specific siRNA. (O) ChIP analysis in I-SceI-induced cells with and without knockdown of MOF examined for bound MOF and pT392-MOF. (*P<0.05, **P<0.001 ***P<0.0001, as determined by the chi-square test).
We ascertained the effect of MOF-T392A expression on the frequency of IR-induced chromosomal and chromatid-type aberrations observed at metaphase: (a) G1-specific aberrations are mostly of the chromosomal type (dicentric with acentric fragment), with a few involving chromatids in human (Figure S4A); (b) S-type aberrations are both chromosome as well as chromatid type in human (Figure S4B) and; (c) G2-type aberrations are predominantly the chromatid type with the least number of dicentrics in human cells (Figure S4C) (Gupta et al., 2005). Similar aberrations were also examined for G1, S and G2-specific in mouse cells (Figure S4D–H). No differences were found in residual IR-induced G1-type chromosomal aberrations in cells with and without expression of mutant MOF-T392A, whereas cells with MOF depletion alone had an increased number of G1-type chromosome aberrations (Figure 2E). Cells with MOF depletion alone had much higher IR-induced S-phase specific residual chromatid/chromosome aberrations as observed at metaphase (Figure 2F) compared to control siRNA treated cells. Expression of the phosphorylation mutant MOF-T392A in these cells failed to rescue this defect. As in S-phase cells, a higher frequency of chromosomal aberrations was observed in MOF depleted G2-phase cells, which could not be complemented by MOF-T392A mutant expression (Figure 2G). The increase in S- or G2-type chromosome aberrations could possibly be due to cell cycle checkpoint defects that can affect chromosome repair but analysis of irradiated G2 phase cells treated with caffeine indicated the increased frequency of G2-type chromosomal aberrations was independent of mutant MOF-T392A (Figure 2H). This suggests that the higher frequency of G2-specific chromosome aberrations observed in cells with mutant MOF-T392A is not due to a cell cycle checkpoint defect. The ability of the T392A phosphorylation defective MOF mutant to complement wild type MOF depletion in G1-phase but not in S/G2-phase cells suggests an important role for MOF phosphorylation in the HR repair pathway. Further more we also validated the data in mouse cells in which endogenous MOF was subjected to cre-mediated deletion and mutant MOF-T329A or wild type MOF was expressed (Figure S3B and S3C). MOF deletion resulted in loss of cell proliferation (Figure 2I). No defect in G1 chromosome repair post irradiation was observed in mouse cells expressing MOF-T392A (Figure 2E), whereas a defect in repair of S- and G2-type chromosomes was observed (Figure 2J–L), results consistent with those in human cells.
We directly tested whether mutant MOF-T392A impacts either or both DNA DSB repair pathways by using engineered cell lines that express a fluorescent protein only upon repair of I-SceI-induced DSBs by the appropriate pathway (Sharma et al., 2010). Accurate repair of the NHEJ dependent fluorescent protein gene template was restored by mutant MOF-T392A expression in MOF depleted cells (Figure 2M), further confirming that MOF phosphorylation is not required for DSB repair by the NHEJ pathway. This is also consistent with the normal chromosomal damage repair observed in G1-phase cells complemented with mutant MOF-T392A after endogenous MOF depletion (Figure 2E and 2J).
The reconstitution frequency of a green fluorescent protein (GFP) HR reporter gene within a chromosomally integrated plasmid substrate (Pandita et al., 2006; Pierce et al., 1999) was used to measure the dependence of HR on MOF-T392 phosphorylation. In cells depleted of endogenous MOF, expression of the MOF-T392A failed to rescue the HR repair defect as the number of GFP-positive cells was not significantly increased (Figure 2N), whereas MOF-T392E phospho-mimic expression did rescue the defect. These data suggest that MOF phosphorylation at T392 is required for DNA DSB repair by the HR pathway but not by NHEJ. We also observed that after I-Sce1 induction the total levels of MOF do not change, but the level of phosphorylated MOF significantly increased (Figure 2O).
MOF phosphorylation at DNA DSBs
Endogenous MOF is phosphorylated at T392 post-irradiation (Figure 1C) and a pT392-MOF-specific antibody readily detects IR-induced foci formation (Figure 3A) after immunostaining. Human or mouse cells with MOF depletion did not show any IR-induced pT392-MOF-specific foci (Figure 3B and 3C). Furthermore, in cells expressing either Flag-tagged wild type or Flag-tagged mutant MOF-T392A, the phospho-specific MOF antibody only detected IR-induced pT392-MOF foci in cells expressing wild type MOF, not mutant MOF-T392A (Figure 3D). Ionizing radiation-induced pT392-MOF foci were observed in cells with ATM (Figure S5A) and IR-induced p-T392-MOF foci colocalized with p-S 1981-ATM, γ-H2AX, 53PB1, BRCA1 and RAD51 foci (Figure 3E–G and Figure S5B, C).
Figure 3. IR-induced pT392-MOF foci and alteration of radiation response by MOF-T329A mutant.
(A) 293 cells with and without Flag-MOF-T392A were irradiated with 2 Gy, fixed post irradiation and immunostained for p-T392 MOF. (B) MOF depleted human cells do not show IR-induced p-T329-MOF foci. (C) MEF with cre-mediated MOF deletion do not show IR-induced p-T392-MOF foci. (D) Cells expressing MOF-T392A did not show pT392-MOF foci. MOF-T392A was detected with Flag antibody. (E–G) Colocalization of IR-induced p-T392-MOF foci with p-S1981-ATM foci (E); with γ-H2AX foci (F); and with 53BP1 foci (G) - (i) low magnification and (ii) high magnification.
However the appearance and disappearance kinetics of pT392-MOF foci were slower than those of γ-H2AX foci (Figure 4A) suggesting that phosphorylated MOF foci may have a role in the processing of a specific type of DNA DSB repair. Since MOF is phosphorylated by ATM, we examined whether expression of MOF-T392A influences the appearance and disappearance of p-S 1981-ATM foci and as expected mutant MOF had no influence on ATM foci formation but did delay the disappearance of p-S1918-ATM foci (Figure 4B). Next, we determined whether loss of T392-MOF phosphorylation influenced γ-H2AX foci kinetics (Figure 4C). Cells expressing mutant MOF-T392A did not show any major difference in H4K16ac levels compared to cells expressing wild type MOF (Figure S1B) and the initial appearance of IR-induced γ-H2AX foci was similar at 2 hr post-exposure (Figure 4C). However, by 4 or 6 hr post irradiation there was a higher frequency of residual γ-H2AX foci in cells expressing MOF-T392A as compared to cells expressing wild type MOF (Figure 4C).
Figure 4. Effect of MOF-T392A on repair protein kinetics post-irradiation.
(A) Human cells were irradiated with 2 Gy and immunostained with anti-pT392-MOF and γ-H2AX. (B) Kinetics of appearance and disappearance of p-S 1981-ATM in cells with and without expression of MOF-T392A. (C) Frequency of cells with γ-H2AX foci in cells with and without mutant MOF-T392A expression. (D–E) IR-induced 53BP1 (D), RIF1 (E) in cells with and without expression of MOF-T392A. (F) IR-induced MDC1 foci in MEFs: (i) Comparison of IR-induced MDC1 foci in cells with and without expression of mutant MOF; (ii) Quantification of cells with MDC1 foci. (G) Cells with and without expression MOF-T329A, and MOF depletion by siRNA with expression of YFP 53BP1 were microirradiated. (H) Initial accumulation and relative fluorescence for a 30 min time course of treatment with YFP-53BP1 at laser-generated DSBs. Error bars represent deviation from three different experiments. (*P<0.05, **P<0.001, as determined by the chi-square test).
53BP1 interaction with phosphorylated MOF
It is known that 53BP1 enhances DSB repair by NHEJ and blocks the recruitment of resection proteins associated with DSB repair by HR (Bouwman et al., 2010; Bunting et al., 2010; Tang et al., 2013). Cells expressing mutant MOF-T392A had higher levels of residual 53BP1 foci (Figure 4D) as well as RIF1 foci (Figure 4E) post-irradiation. Since MDC1 depletion results in loss of 53BP1 foci formation post irradiation (Minter-Dykhouse et al., 2008), we examined whether MOF-T392A expression effects MDC1 accumulation, which may impair recruitment of HR related proteins like CtIP and Rad51. Expression of MOF-T392A resulted in a higher frequency of cells with MDC1 foci formation post-irradiation (Figure 4F). We also found that the cells expressing MOF-T392A show delayed kinetics of yellow fluorescent protein (YFP)-53BP1 disappearance at laser track induced DNA damage (Figure 4G and 4H).
MOF phosphorylation affects ATM dynamics and DNA retention
Expression of mutant MOF-T392A delayed the disappearance of pS1981-ATM foci post irradiation (Figure 4B) while not affecting H4K16 acetylation (Figure S1B and Figure S1B 1H). Live cell imaging of microirradiated cells expressing ATM tagged with YFP (Figure 5A and 5B) did not detect any major effect of mutant MOF-T392A expression on ATM accumulation but the kinetics of YFP-tagged ATM accumulation at the damage sites was modestly reduced. To determine whether the altered ATM kinetics was due to altered retention of mutant MOF post-irradiation, we expressed Flag-tagged wild type and mutant MOF-T392A in HEK293 cells (Figure S2D), exposed the cells to 5 Gy and after increasing intervals isolated and cross-linked the DNA for Flag antibody immunoprecipitation. The association of wild type MOF with DNA increased post irradiation whereas MOF-T392A association with DNA did not significantly increase (Figure 5C), suggesting that phosphorylation enhances MOF chromatin binding. We examined MOF retention in G1 and S/G2-phase enriched cells (Figure 5D and 5E) and found a significant increase in the retention of wild type MOF in S/G2 phase cells, where as no significant increase in mutant MOF retention was observed. To avoid any competitive interference with MOF-T392A binding to chromatin, endogenous MOF was depleted using UTR-specific siRNA. Conversely, we determined whether expression of mutant MOF-T329A effects the association of wild type MOF with chromatin by quantitating chromatin bound MOF and MOF-T392A in cells expressing Flag-tagged MOF-T392A and HA-tagged wild type MOF after endogenous MOF depletion cells by UTR-specific siRNA (Figure S2D) and quantitated MOF and MOF-T392A bound to chromatin (Figure 5B). Prior to irradiation MOF-T392A expression does decrease wild type MOF association with chromatin but upon irradiation wild type MOF and not mutant MOF-T392A showed enhanced association with chromatin (Figure S6). We further examined the effect of the MOF-T392A mutation on ATM chromatin association in irradiated cells by isolating soluble and chromatin bound fractions (Figure 5F and 5G). In Flag-tagged MOF-T392A expressing cells, as compared to wild type transfected cells, western blot analysis of the chromatin bound proteins indicated there was a significant decrease in chromatin associated ATM. In addition we found that MRE11 binding to chromatin is also reduced post-irradiation in cells expressing MOF-T392A cells (Figure 5H and 5I). Phosphorylation of MOF T392 post irradiation, therefore, enhances chromatin retention of MOF and a subset of other critical proteins required for DNA damage repair.
Figure 5. Effect of MOF-T392A on ATM dynamics and association with chromatin post-irradiation.
(A) U2OS cells expressing YFP-ATM were transfected with MOF-T392A, microirradiated and accumulation of YFP-ATM. (B) Accumulation kinetics and relative fluorescence for a 60 min course of treatment with YFP-ATM at laser-generated DSBs. Error bars represent the standard deviations from three independent different experiments. (C–E) Comparison of MOF and MOF-T392A association with chromatin after cells were exposed to 5 Gy. ChIP was performed with an MOF or Flag-MOF-T392A or Rb antibody, and DNA was quantified by absorbance at 260 nm by NanoDrop 2000 in: (C) Cells in exponential phase; (D) G1-phase and; (E) S/G2-phase. (F, G) Cells with and without expression of MOF-T392 were analyzed for chromatin fraction vs soluble fraction of ATM by Western blotting: (F) ATM was detected with a specific ATM antibody and; (G) Relative levels of ATM. (H, I) Cells with and without expression of MOF-T392 were analyzed for the retention of MRE11 by Western blotting post irradiation (H) and histogram showing relative accumulation MRE11 (I). The mean are from three different experiments. (*P<0.05, **P<0.001 ***P<0.0001, as determined by the chi-square test).
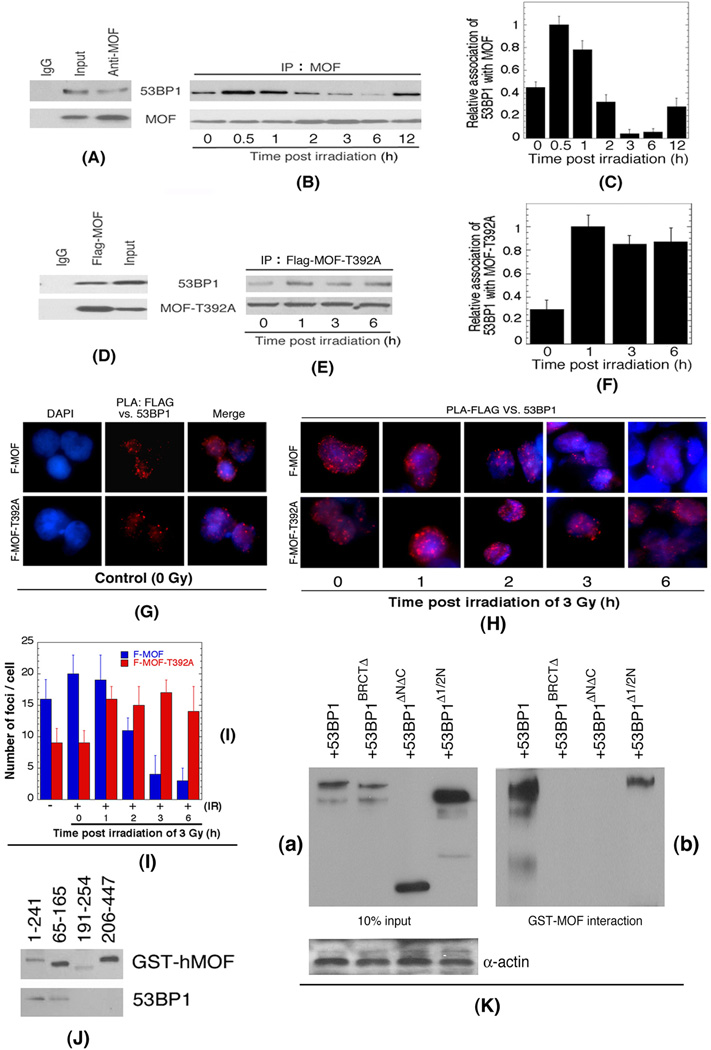
Expression of mutant MOF-T392A affected DNA DSB repair in S and G2-phase cells, where homologous recombination is upregulated. Since 53BP1 is known to suppress DSB repair by HR, we examined the cellular interaction of wild type and mutant MOF with 53BP1 before and after irradiation. Immunoprecipitation studies indicated a fraction of both wild type (Figure 6A–C) as well as mutant MOF-T392A (Figure 6D–F) associated with 53BP1 under non-stress conditions and the interaction was not disrupted by ethidium bromide (Figure S7A). The interaction of wild type MOF with 53BP1 increases immediately post irradiation and then decreases until, by 3 hr post-irradiation, the complex largely disappeared. Only later, at 12 hr post-irradiation, did the MOF/53BP1 complex reform to control levels (Figure 6B and 6C). In contrast, the association of mutant MOF-T392A with 53BP1, while also increasing following irradiation does not decline, being stable over the next 6 hr period (Figure 6E and 6F). To further confirm the in-cell association of wild type MOF and mutant MOF-T329A, we have used an in situ proximity ligation assay (PLA), in which the close physical association of two proteins is visualized by a fluorescent signal (Fredriksson et al., 2002; Soderberg and Lang, 2006). Both wild type and mutant MOF-T392A associated with 53BP1, but while the 53BP1/MOF interaction declined by 2 hr post-irradiation, the mutant MOF-T392A did not dissociate from 53BP1 over the course of 6 hr post-irradiation recovery, a result similar to the immunoprecipitation studies (Figure 6G–I). Together, these results suggest a mechanism whereby phosphorylation of MOF at T392 may be required for 53BP1 release from DSB sites, a step required prior to recruitment of proteins involved in DSB repair by HR.
Figure 6. MOF phosphorylation decreases interaction with 53BP1 post-irradiation.
(A–F) The interactions between 53BP1 with wild type MOF (A) and MOF-T392A (D). (B) Cells irradiated with 10 Gy and fixed at different times for immunoprecipitation. (C) Relative association of 53BP1 with MOF. (E) Interaction between 53BP1 and MOF-T392A as determined by immunoprecipitation and western blot. Cells irradiated with 10 Gy and fixed at different times for immunoprecipitation. (F) Relative association of 53BP1 with MOF-T392A. (G) Control (unirradiated) cells showing PLA foci of FLAG (for MOF) vs. 53BP1 detected by the described procedure (Fredriksson et al., 2002; Hegde et al., 2012). The nuclei are counterstained with DAPI, and the PLA signals are visualized in a fluorescence microscope. (H) Cells irradiated with 3 Gy and examined for PLA foci at different times post irradiation. (I) Histogram showing number of PLA-FLAG (MOF) vs. 53BP1 foci per cell in cells expressing wild type and mutant MOF were determined by co-immunoprecipitation as well as by proximity ligation assay (PLA). (J) Mapping of 53BP1 binding site in hMOF. The domains of hMOF and interaction with 53BP1 were conducted as described previously (Gupta et al., 2005). (K) Mapping of hMOF binding site in 53BP1:
(a) 293T cells were transected with either 53BP1 WT-FLAG or truncated 53BP1-FLAG. (b) hMOF-GST fusion protein was expressed in bacteria and immobilized hMOF was incubated with the whole cell lysates prepared from :53BP1 WT-FLAG (1–1973 AA), 53BP1 ΔBRCT- FLAG (1–1711 AA), 53BP1 ΔN/ΔC FLAG - (1220 – 1711 AA) , 53BP1Δ1/2N FLAG -(618–1711 AA). After extensive washing, bound fractions were resolved in SDS-PAGE and Western blot with FLAG M5 (Sigma) antibody.
The protein domains involved in the MOF and 53BP1 interaction were mapped by deletion analysis. Binding of purified 53BP1 to GST tagged MOF deletion mutants identified the MOF N-terminal region between amino acids 65–165 as mediating 53BP1 binding (Figure 6J). Similar analysis using whole cell extracts expressing mutants of 53BP1 identified the 53BP1 N-terminal domain as required for interaction with added, purified GST-MOF (Figure 6K).
pT392-MOF is critical for resection during HR repair
Since depletion of MOF reduces the formation of IR-induced RAD51 foci, a hallmark of HR (Sharma et al., 2010), we examined further the effect of pT392-MOF mutation on recruitment of HR components to DSBs. The frequency of RIF1 co-localization with 53BP1 is significantly higher in post-irradiated cells expressing MOF-T392A than in control cells (Figure 7A and S7B). Consistent with results that p-T392-MOF has a role in HR, a reduced frequency of cells with IR-induced BRCA1, CtIP, RAD51, RPA2, MRE11, RAP80 and FANCD2 foci was found in mutant MOF-T392A expressing cells (Figure 7B–H), supporting a direct link between DSB repair by HR and phosphorylation of MOF-T392.
Figure 7. Phosphorylated MOF-T392 is required for HR and HR related protein recruitment.
(A) Co-localization of 53BP1 and RIF1 in cells with and without MOF-T392A. (B, C) Cells with and without expression of mutant MOF were irradiated with 10 Gy and examined for BRCA1 foci (B) and CtIP foci (C) at the indicated times. (D) RAD51 foci induced after different doses of ionizing radiation exposure in cells with and without expression of mutant MOF-T392A. (E–H) Cells with and without expression of mutant MOF-T392A were irradiated with 10 Gy and quantified for foci at different time points post irradiation. (E) RPA2, (F) MRE11, (G) RAP80 and (H) FANCD2. (I–L) Detection of repair proteins at I-SceI induced DSB sites by ChIP. Cell synchronization, cell cycle analysis, ISec1 induced DSB and ChIP analysis was done according to the described procedure (Rodrigue et al., 2006). The closest PCR product to the DSB site is 94–378. (I) G1 phase cells expressing wild type MOF. (J) G1/S phase cells expressing wild type MOF. (K) G1/S phase cells expressing mutant MOF. (L) S/G2 phase cells expressing wild type MOF. (M) S/G2 phase cells expressing mutant MOF-T392A. Error bars represent the standard deviations from three different experiments. (*P<0.05, **P<0.001, as determined by the chi-square test).
To further determine the role of MOF phosphorylation in DNA repair protein recruitment, we compared the levels of BRCA1, RAD51, MRE11 and KU80 at different distances from an I-SceI induced DSB site using ChIP analysis with specific primers (Rodrigue et al., 2006). Cells enriched in G1 or S/G2 phase (Supplementary Table 1), with and without expression of mutant MOF-T392A, were induced for I-SceI site cleavage and analyzed by ChIP for RAD51, MRE11, KU80, MOF and Flag-tagged MOF-T392A. In G1-phase cells, the localization and increase in KU80 at close proximity to the DSB break was unaltered by MOF-T392A expression whereas levels of RAD51 at the break site were significantly reduced in G1-phase cells expressing wild type or mutant MOF (Figure 7I). A modest impact of MOF-T392A on the recruitment of RAD51 was observed in G1/S cells, where the levels were relatively low as compared to cells expressing wild type MOF (Figure 7J and 7K). However, in S/G2 phase cells the high levels of RAD51 or MRE11 that localized near the break in cells expressing wild type MOF were significantly reduced in cells expressing mutant MOF-T392A (Figure 7L and 7M).
The reduced level of RAD51 at the DSB site in MOF-T392A expressing cells could be due to altered recruitment of BRAC1, therefore, we examined the localization of 53BP1 and BRCA1 in G1 and S/G2 phase cells expressing wild or mutant MOF-T392A. Although BRCA1 co-localizes with p-T329-MOF foci post-irradiation, no direct interaction was observed between BRCA1 and MOF by co-immunoprecipitation (Figure S7C). The levels of RAD51 was similar in cells expressing wild type or mutant MOF-T392A (Figure S7D), maximum 53BP1 association at the DSB site was observed in G1-phase cells expressing either wild type or mutant MOF-T392A whereas minimum levels of BRCA1 were observed (Figure S7E). In contrast, maximum levels of BRCA1 and minimum levels of the 53BP1 at the DSB site were observed in S/G2 phase cells expressing wild type MOF, but MOF-T392A expression reduced BRCA1 levels and increased 53BP1 levels at the DSB site (Figure S7F).
Discussion
We have identified a mechanism whereby ATM-dependent phosphorylation of MOF threonine-392 regulates 53BP1-dependent recruitment of proteins involved in the cell cycle specific selection between NHEJ and HR DNA DSB repair. Cells depleted for MOF fail to recruit MDC1 and its downstream effectors 53BP1 and BRCA1 to DNA damage foci (Li et al., 2010). Similar to ATM deficiency, endogenous MOF depletion also results in an overall defect in DNA DSB repair throughout all phases of the cell cycle (Gupta et al., 2005; Pandita and Hittelman, 1992a, b; Pandita et al., 1999; Sharma et al., 2010). The loss of ATM-dependent MOF phosphorylation has no effect on IR-induced G1-phase cell killing, chromosome damage repair, or MDC1 foci formation. In contrast, MOF-T392 phosphorylation is critical for cell survival and chromosome damage repair in S- and G2-phase. Interestingly, ATM dependent-MOF phosphorylation increases MOF retention on DNA post-irradiation in S/G2-phase cells. Furthermore, the fact that there is a reduced frequency of cells with HR-related repairsome foci after MOF-T392 expression, strongly supports the argument that ATM-dependent phosphorylation of MOF regulates recruitment of HR-related proteins during the S/G2-phases of the cell cycle.
The 53BP1 protein has been implicated in the regulation of DNA DSB pathway choice (Morales et al., 2003; Ward et al., 2003; Zimmermann et al., 2013) and the first effector of 53BP1 was identified to be RIF1 (Chapman et al., 2013; Di Virgilio et al., 2013; Escribano-Diaz and Durocher, 2013; Feng et al., 2013; Zimmermann et al., 2013). In BRCA1-deficient cells, suppression of DNA end resection (Bunting et al., 2010) is critical, and 53BP1 as well as the 53BP1-interacting protein RIF1 have been shown to be required for inhibition of resection. Accumulation of RIF1 at DSB sites results from its binding to phosphorylated 53BP1 (Anbalagan et al., 2011; Bonetti et al., 2010), and the frequency of RIF1/53BP1 foci co-localization increases in cells expressing the MOF-T392A mutant. These results support the notion that phosphorylated-MOF plays an essential role in regulating recruitment of HR-related proteins to the damaged sites. Furthermore, we have shown that expression of mutant MOF-T392A results in failure of 53BP1 displacement and reduced BRCA1 accumulation at DSB site during the S/G2-phase of the cell cycle. Cumulatively, we have shown that in response to DNA damage, ATM phosphorylates MOF, which promotes dissociation of a complex containing 53BP1 and MOF from the site of damage in S/G2 phase cells in order to allow for subsequent recruitment of the resection machinery facilitating HR-dependent repair. As such, this ATM dependent MOF post-translational modification mediates DNA DSB repair pathway choice according to the cell cycle position.
MATERIALS AND METHODS
Cell survival and chromosomal aberration analysis
Cell survival and chromosomal aberrations analysis were carried out as described previously (Pandita et al., 2006). Cell enrichment in different phases was achieved by serum deprivation as well as thymidine block (Supplementary Table 1). For serum deprivation, cells were incubated for till confluence, washed and incubated for 48 h in serum free medium. Cell cycle analysis by flow revealed that more than 95% cells were in G1-phase of the cell cycle. For S- and G2-phase cells were enriched by thymidine block and then released. Cell cycle analysis was performed by flow cytometric analysis (Pandita and Hittelman, 1992a, b). MOF mutant (MOF-T329A and MOF-T392E) were generated by site-directed mutagenesis, then cloned into pcDNA3.1 vector and transfected in 293, RKO and U2OS cells. siRNA treatment of cell lines was performed as described (Gupta et al., 2008; Gupta et al., 2005; Pandita, 2006). Mutant generation, siRNA transfection, immunofluorescence, and protein retention assay were carried out by described procedures (Agarwal et al., 2008; Gupta et al., 2008; Gupta et al., 2005; Gupta et al., 2009; Hunt et al., 2007; Kumar et al., 2011; Pandita et al., 2006; Pandita, 2006; Pandita et al., 1999; Sharma et al., 2003a; Sharma et al., 2003b; Sharma et al., 2010; Singh et al., 2013). Site directed mutagenesis of MOF-T392 as well as in vitro kinase assays were performed to confirm that MOF-T392 phosphorylation is ATM dependent. A rabbit pT392-MOF polyclonal antibody was generated. Phosphorylated MOF-T392 foci were determined by immunostaining. The impact of MOF phosphorylation on IR response was determined by clonogenic assay and by measuring chromosome aberrations at metaphase in cells expressing mutant or wild type MOF. Cells in different phases were enriched by serum starvation and thymidine block. Endogenous MOF was depleted by UTR-specific MOF-siRNA in cells expressing mutant MOF.
Immunofluorescence, microirradiation and protein retention assay
The procedure for immunofluorescence analysis was the same as previously described (Agarwal et al., 2008; Gupta et al., 2009; Hunt et al., 2007; Pandita et al., 2006; Pandita et al., 1999). For microirradiation, cells transfected with YFP labeled ATM or 53BP1 were grown on glass bottom culture dishes then microirradiated and the signal was quantified as described (Sharma et al., 2010). The frequency of co-localization between two proteins was determined by the procedure described previously (Asaithamby et al., 2011). The procedure for protein retention assay is same as described (Sharma et al., 2003a; Sharma et al., 2003b).
Western blot analysis, IP, ChIP and GST Pull down Assay
Western blot analysis, immunoprecipitation (IP) chromatin immunoprecipitation (ChIP) and GST pull down assay were performed as described in the experimental procedures of supplemental information.
Histone acetyl-transferase (HAT) assay
Plasmid pGEX-4T1 carrying the open reading frame for GST-MOF was mutagenized to generate T392A and T392E versions. The wild type and mutant GST-MOF proteins were expressed in BL21(DE3) cells and were affinity purified on glutathione Sepharose beads. Equal amounts of the proteins were incubated with Xenopus core histones, 50mM Tris pH 7.5, 100mM NaCl, and acetylCoA. Acetylation efficiency was determined by western blotting with antisera specific for H4K16ac.
In situ proximity ligation assay (PLA)
293 cells transiently expressing FLAG-tagged WT or MOF-T392A grown overnight in 16-well chamber slides were fixed with 4% paraformaldehyde, and permeabilized with 0.2% Tween-20, followed by incubation with a primary antibody for FLAG for MOF [mouse; SIGMA] vs. γH2AX (rabbit; GENTEX) or FLAG vs. 53BP1 (rabbit) and the PLA assay performed using the Duolink PLA kit from OLink Bioscience (Uppsala, Sweden).
Supplementary Material
Highlights.
DNA DSBs induce ATM-dependent MOF phosphorylation at Threonine 392.
Mutant MOF-T392A expression abrogates DSB repair only in S/G2-phase cells.
MOF-T392A mutant delayed dissociation with 53BP1, and less association with DNA.
MOF phosphorylation regulates recruitment of proteins (BRCA1, RAD51) required for HR.
Acknowledgements
We thank Maria Jasin, Simmon Boulton, Daniel Durocher, Matthew Porteus, Tanya Paull, Yali Duo and Simon Powell for providing reagents. Sincere thanks are due to Daniel Durocher, Andre Nussenzweig and to members of Pandita laboratory for critical reading and suggestions for the manuscript. This work has been supported by NIH grants CA129537 and CA154320.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
Author contributions: A.G., C.R.H., M.L.H., and T.K.P. designed research; A.G., C.R.H., M.L.H., S.C., D.U., N.H., M.S., D.R., W.N.H., A.A., S.N., T.K.H., T.L., R.K.P., performed experiments; and A.G., N.H., C.R.H., J.K.T. and T.K.P. wrote paper.
Competing financial interests:
The authors declare no competing financial interests and have no conflict of interest
References
- Agarwal M, Pandita S, Hunt CR, Gupta A, Yue X, Khan S, Pandita RK, Pratt D, Shay JW, Taylor JS, et al. Inhibition of telomerase activity enhances hyperthermia-mediated radiosensitization. Cancer Res. 2008;68:3370–3378. doi: 10.1158/0008-5472.CAN-07-5831. [DOI] [PubMed] [Google Scholar]
- Anbalagan S, Bonetti D, Lucchini G, Longhese MP. Rif1 supports the function of the CST complex in yeast telomere capping. PLoS genetics. 2011;7:e1002024. doi: 10.1371/journal.pgen.1002024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Asaithamby A, Hu B, Chen DJ. Unrepaired clustered DNA lesions induce chromosome breakage in human cells. Proceedings of the National Academy of Sciences of the United States of America. 2011;108:8293–8298. doi: 10.1073/pnas.1016045108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bhadra MP, Horikoshi N, Pushpavallipvalli SN, Sarkar A, Bag I, Krishnan A, Lucchesi JC, Kumar R, Yang Q, Pandita RK, et al. The role of MOF in the ionizing radiation response is conserved in Drosophila melanogaster. Chromosoma. 2012;121:79–90. doi: 10.1007/s00412-011-0344-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bonetti D, Clerici M, Anbalagan S, Martina M, Lucchini G, Longhese MP. Shelterin-like proteins and Yku inhibit nucleolytic processing of Saccharomyces cerevisiae telomeres. PLoS genetics. 2010;6:e1000966. doi: 10.1371/journal.pgen.1000966. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bothmer A, Robbiani DF, Di Virgilio M, Bunting SF, Klein IA, Feldhahn N, Barlow J, Chen HT, Bosque D, Callen E, et al. Regulation of DNA end joining, resection, and immunoglobulin class switch recombination by 53BP1. Molecular cell. 2011;42:319–329. doi: 10.1016/j.molcel.2011.03.019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bothmer A, Robbiani DF, Feldhahn N, Gazumyan A, Nussenzweig A, Nussenzweig MC. 53BP1 regulates DNA resection and the choice between classical and alternative end joining during class switch recombination. The Journal of experimental medicine. 2010;207:855–865. doi: 10.1084/jem.20100244. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bouwman P, Aly A, Escandell JM, Pieterse M, Bartkova J, van der Gulden H, Hiddingh S, Thanasoula M, Kulkarni A, Yang Q, et al. 53BP1 loss rescues BRCA1 deficiency and is associated with triple-negative and BRCA-mutated breast cancers. Nat Struct Mol Biol. 2010;17:688–695. doi: 10.1038/nsmb.1831. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bunting SF, Callen E, Wong N, Chen HT, Polato F, Gunn A, Bothmer A, Feldhahn N, Fernandez-Capetillo O, Cao L, et al. 53BP1 inhibits homologous recombination in Brca1-deficient cells by blocking resection of DNA breaks. Cell. 2010;141:243–254. doi: 10.1016/j.cell.2010.03.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Callen E, Di Virgilio M, Kruhlak MJ, Nieto-Soler M, Wong N, Chen HT, Faryabi RB, Polato F, Santos M, Starnes LM, et al. 53BP1 mediates productive and mutagenic DNA repair through distinct phosphoprotein interactions. Cell. 2013;153:1266–1280. doi: 10.1016/j.cell.2013.05.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chapman JR, Barral P, Vannier JB, Borel V, Steger M, Tomas-Loba A, Sartori AA, Adams IR, Batista FD, Boulton SJ. RIF1 is essential for 53BP1-dependent nonhomologous end joining and suppression of DNA double-strand break resection. Molecular cell. 2013;49:858–871. doi: 10.1016/j.molcel.2013.01.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chapman JR, Taylor MR, Boulton SJ. Playing the end game: DNA double-strand break repair pathway choice. Molecular cell. 2012;47:497–510. doi: 10.1016/j.molcel.2012.07.029. [DOI] [PubMed] [Google Scholar]
- Di Virgilio M, Callen E, Yamane A, Zhang W, Jankovic M, Gitlin AD, Feldhahn N, Resch W, Oliveira TY, Chait BT, et al. Rif1 prevents resection of DNA breaks and promotes immunoglobulin class switching. Science. 2013;339:711–715. doi: 10.1126/science.1230624. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Escribano-Diaz C, Durocher D. DNA repair pathway choice--a PTIP of the hat to 53BP1. EMBO reports. 2013;14:665–666. doi: 10.1038/embor.2013.99. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feng L, Fong KW, Wang J, Wang W, Chen J. RIF1 counteracts BRCA1-mediated end resection during DNA repair. J Biol Chem. 2013;288:11135–11143. doi: 10.1074/jbc.M113.457440. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fredriksson S, Gullberg M, Jarvius J, Olsson C, Pietras K, Gustafsdottir SM, Ostman A, Landegren U. Protein detection using proximity-dependent DNA ligation assays. Nat Biotechnol. 2002;20:473–477. doi: 10.1038/nbt0502-473. [DOI] [PubMed] [Google Scholar]
- Fullgrabe J, Lynch-Day MA, Heldring N, Li W, Struijk RB, Ma Q, Hermanson O, Rosenfeld MG, Klionsky DJ, Joseph B. The histone H4 lysine 16 acetyltransferase hMOF regulates the outcome of autophagy. Nature. 2013;500:468–471. doi: 10.1038/nature12313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gupta A, Guerin-Peyrou TG, Sharma GG, Park C, Agarwal M, Ganju RK, Pandita S, Choi K, Sukumar S, Pandita RK, et al. The mammalian ortholog of Drosophila MOF that acetylates histone H4 lysine 16 is essential for embryogenesis and oncogenesis. Mol Cell Biol. 2008;28:397–409. doi: 10.1128/MCB.01045-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gupta A, Hunt CR, Chakraborty S, Pandita RK, Yordy J, Ramnarain DB, Horikoshi N, Pandita TK. Role of 53BP1 in the Regulation of DNA Double-Strand Break Repair Pathway Choice. Radiation research. 2014;181:1–8. doi: 10.1667/RR13572.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gupta A, Sharma GG, Young CS, Agarwal M, Smith ER, Paull TT, Lucchesi JC, Khanna KK, Ludwig T, Pandita TK. Involvement of human MOF in ATM function. Mol Cell Biol. 2005;25:5292–5305. doi: 10.1128/MCB.25.12.5292-5305.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gupta A, Yang Q, Pandita RK, Hunt CR, Xiang T, Misri S, Zeng S, Pagan J, Jeffery J, Puc J, et al. Cell cycle checkpoint defects contribute to genomic instability in PTEN deficient cells independent of DNA DSB repair. Cell Cycle. 2009;8:2198–2210. doi: 10.4161/cc.8.14.8947. [DOI] [PubMed] [Google Scholar]
- Hegde ML, Banerjee S, Hegde PM, Bellot LJ, Hazra TK, Boldogh I, Mitra S. Enhancement of NEIL1 protein-initiated oxidized DNA base excision repair by heterogeneous nuclear ribonucleoprotein U (hnRNP-U) via direct interaction. J Biol Chem. 2012;287:34202–34211. doi: 10.1074/jbc.M112.384032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hunt CR, Pandita RK, Laszlo A, Higashikubo R, Agarwal M, Kitamura T, Gupta A, Rief N, Horikoshi N, Baskaran R, et al. Hyperthermia activates a subset of ataxia-telangiectasia mutated effectors independent of DNA strand breaks and heat shock protein 70 status. Cancer Res. 2007;67:3010–3017. doi: 10.1158/0008-5472.CAN-06-4328. [DOI] [PubMed] [Google Scholar]
- Kakarougkas A, Ismail A, Klement K, Goodarzi AA, Conrad S, Freire R, Shibata A, Lobrich M, Jeggo PA. Opposing roles for 53BP1 during homologous recombination. Nucleic acids research. 2013 doi: 10.1093/nar/gkt729. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kumar R, Hunt CR, Gupta A, Nannepaga S, Pandita RK, Shay JW, Bachoo R, Ludwig T, Burns DK, Pandita TK. Purkinje cell-specific males absent on the first (mMof) gene deletion results in an ataxia-telangiectasia-like neurological phenotype and backward walking in mice. Proc Natl Acad Sci U S A. 2011;108:3636–3641. doi: 10.1073/pnas.1016524108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee JH, Paull TT. Direct activation of the ATM protein kinase by the Mre11/Rad50/Nbs1 complex. Science. 2004;304:93–96. doi: 10.1126/science.1091496. [DOI] [PubMed] [Google Scholar]
- Li X, Corsa CA, Pan PW, Wu L, Ferguson D, Yu X, Min J, Dou Y. MOF and H4 K16 acetylation play important roles in DNA damage repair by modulating recruitment of DNA damage repair protein Mdc1. Mol Cell Biol. 2010;30:5335–5347. doi: 10.1128/MCB.00350-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matsuoka S, Ballif BA, Smogorzewska A, McDonald ER, 3rd, Hurov KE, Luo J, Bakalarski CE, Zhao Z, Solimini N, Lerenthal Y, et al. ATM and ATR substrate analysis reveals extensive protein networks responsive to DNA damage. Science. 2007;316:1160–1166. doi: 10.1126/science.1140321. [DOI] [PubMed] [Google Scholar]
- Minter-Dykhouse K, Ward I, Huen MS, Chen J, Lou Z. Distinct versus overlapping functions of MDC1 and 53BP1 in DNA damage response and tumorigenesis. J Cell Biol. 2008;181:727–735. doi: 10.1083/jcb.200801083. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morales JC, Xia Z, Lu T, Aldrich MB, Wang B, Rosales C, Kellems RE, Hittelman WN, Elledge SJ, Carpenter PB. Role for the BRCA1 C-terminal repeats (BRCT) protein 53BP1 in maintaining genomic stability. J Biol Chem. 2003;278:14971–14977. doi: 10.1074/jbc.M212484200. [DOI] [PubMed] [Google Scholar]
- Nakamura K, Sakai W, Kawamoto T, Bree RT, Lowndes NF, Takeda S, Taniguchi Y. Genetic dissection of vertebrate 53BP1: a major role in non-homologous end joining of DNA double strand breaks. DNA Repair (Amst) 2006;5:741–749. doi: 10.1016/j.dnarep.2006.03.008. [DOI] [PubMed] [Google Scholar]
- Obradovic Z, Peng K, Vucetic S, Radivojac P, Dunker AK. Exploiting heterogeneous sequence properties improves prediction of protein disorder. Proteins. 2005;61(Suppl 7):176–182. doi: 10.1002/prot.20735. [DOI] [PubMed] [Google Scholar]
- Oksenych V, Alt FW, Kumar V, Schwer B, Wesemann DR, Hansen E, Patel H, Su A, Guo C. Functional redundancy between repair factor XLF and damage response mediator 53BP1 in V(D)J recombination and DNA repair. Proceedings of the National Academy of Sciences of the United States of America. 2012;109:2455–2460. doi: 10.1073/pnas.1121458109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pandita RK, Sharma GG, Laszlo A, Hopkins KM, Davey S, Chakhparonian M, Gupta A, Wellinger RJ, Zhang J, Powell SN, et al. Mammalian Rad9 plays a role in telomere stability, S- and G2-phase-specific cell survival, and homologous recombinational repair. Mol Cell Biol. 2006;26:1850–1864. doi: 10.1128/MCB.26.5.1850-1864.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pandita TK. A multifaceted role for ATM in genome maintenance. Expert Rev Mol Med. 2003;5:1–21. doi: 10.1017/S1462399403006318. [DOI] [PubMed] [Google Scholar]
- Pandita TK. Role of mammalian Rad9 in genomic stability and ionizing radiation response. Cell Cycle. 2006;5:1289–1291. doi: 10.4161/cc.5.12.2862. [DOI] [PubMed] [Google Scholar]
- Pandita TK, Hittelman WN. The contribution of DNA and chromosome repair deficiencies to the radiosensitivity of ataxia-telangiectasia. Radiat Res. 1992a;131:214–223. [PubMed] [Google Scholar]
- Pandita TK, Hittelman WN. Initial chromosome damage but not DNA damage is greater in ataxia telangiectasia cells. Radiat Res. 1992b;130:94–103. [PubMed] [Google Scholar]
- Pandita TK, Lieberman HB, Lim DS, Dhar S, Zheng W, Taya Y, Kastan MB. Ionizing radiation activates the ATM kinase throughout the cell cycle. Oncogene. 2000;19:1386–1391. doi: 10.1038/sj.onc.1203444. [DOI] [PubMed] [Google Scholar]
- Pandita TK, Westphal CH, Anger M, Sawant SG, Geard CR, Pandita RK, Scherthan H. Atm inactivation results in aberrant telomere clustering during meiotic prophase. Mol Cell Biol. 1999;19:5096–5105. doi: 10.1128/mcb.19.7.5096. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peterson CL, Almouzni G. Nucleosome dynamics as modular systems that integrate DNA damage and repair. Cold Spring Harb Perspect Biol. 2013;5 doi: 10.1101/cshperspect.a012658. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pierce AJ, Johnson RD, Thompson LH, Jasin M. XRCC3 promotes homology-directed repair of DNA damage in mammalian cells. Genes Dev. 1999;13:2633–2638. doi: 10.1101/gad.13.20.2633. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rodrigue A, Lafrance M, Gauthier MC, McDonald D, Hendzel M, West SC, Jasin M, Masson JY. Interplay between human DNA repair proteins at a unique double-strand break in vivo. The EMBO journal. 2006;25:222–231. doi: 10.1038/sj.emboj.7600914. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sartori AA, Lukas C, Coates J, Mistrik M, Fu S, Bartek J, Baer R, Lukas J, Jackson SP. Human CtIP promotes DNA end resection. Nature. 2007;450:509–514. doi: 10.1038/nature06337. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sharma GG, Gupta A, Wang H, Scherthan H, Dhar S, Gandhi V, Iliakis G, Shay JW, Young CS, Pandita TK. hTERT associates with human telomeres and enhances genomic stability and DNA repair. Oncogene. 2003a;22:131–146. doi: 10.1038/sj.onc.1206063. [DOI] [PubMed] [Google Scholar]
- Sharma GG, Hwang KK, Pandita RK, Gupta A, Dhar S, Parenteau J, Agarwal M, Worman HJ, Wellinger RJ, Pandita TK. Human heterochromatin protein 1 isoforms HP1(Hsalpha) and HP1(Hsbeta) interfere with hTERT-telomere interactions and correlate with changes in cell growth and response to ionizing radiation. Mol Cell Biol. 2003b;23:8363–8376. doi: 10.1128/MCB.23.22.8363-8376.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sharma GG, So S, Gupta A, Kumar R, Cayrou C, Avvakumov N, Bhadra U, Pandita RK, Porteus MH, Chen DJ, et al. MOF and histone H4 acetylation at lysine 16 are critical for DNA damage response and double-strand break repair. Mol Cell Biol. 2010;30:3582–3595. doi: 10.1128/MCB.01476-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shiloh Y, Ziv Y. The ATM protein kinase: regulating the cellular response to genotoxic stress, and more. Nature reviews Molecular cell biology. 2013;14:197–210. [PubMed] [Google Scholar]
- Singh M, Hunt CR, Pandita RK, Kumar R, Yang CR, Horikoshi N, Bachoo R, Serag S, Story MD, Shay JW, et al. Lamin a/c depletion enhances DNA damage-induced stalled replication fork arrest. Molecular and cellular biology. 2013;33:1210–1222. doi: 10.1128/MCB.01676-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Soderberg M, Lang MA. Megaprimer-based methodology for deletion of a large fragment within a repetitive polypyrimidine-rich DNA. Mol Biotechnol. 2006;32:65–71. doi: 10.1385/MB:32:1:065. [DOI] [PubMed] [Google Scholar]
- Symington LS, Gautier J. Double-strand break end resection and repair pathway choice. Annu Rev Genet. 2011;45:247–271. doi: 10.1146/annurev-genet-110410-132435. [DOI] [PubMed] [Google Scholar]
- Taipale M, Rea S, Richter K, Vilar A, Lichter P, Imhof A, Akhtar A. hMOF histone acetyltransferase is required for histone H4 lysine 16 acetylation in mammalian cells. Mol Cell Biol. 2005;25:6798–6810. doi: 10.1128/MCB.25.15.6798-6810.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tang J, Cho NW, Cui G, Manion EM, Shanbhag NM, Botuyan MV, Mer G, Greenberg RA. Acetylation limits 53BP1 association with damaged chromatin to promote homologous recombination. Nat Struct Mol Biol. 2013;20:317–325. doi: 10.1038/nsmb.2499. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ward IM, Minn K, van Deursen J, Chen J. p53 Binding protein 53BP1 is required for DNA damage responses and tumor suppression in mice. Molecular and cellular biology. 2003;23:2556–2563. doi: 10.1128/MCB.23.7.2556-2563.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- West SC. Molecular views of recombination proteins and their control. Nature reviews Molecular cell biology. 2003;4:435–445. doi: 10.1038/nrm1127. [DOI] [PubMed] [Google Scholar]
- Zimmermann M, Lottersberger F, Buonomo SB, Sfeir A, de Lange T. 53BP1 regulates DSB repair using Rif1 to control 5’ end resection. Science. 2013;339:700–704. doi: 10.1126/science.1231573. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.