Figure 3.
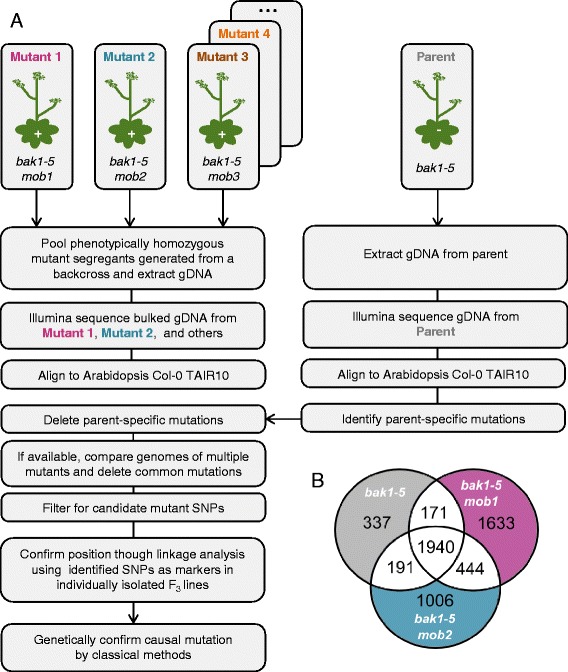
Pipeline for bulking segregants and identification of unique SNPs. (A) The recessive bak1-5 mob1 and bak1-5 mob2 mutants were back-crossed to the parent bak1-5, allowed to self-fertilize in the F1, and phenotypically scored in the F2 for the mob phenotype. Positive segregants were bulk harvested and genomic DNA was prepared and sequenced using the Illumina HiSeq platform. For comparison, the bak1-5 genome was also sequenced. A similar genetics pipeline could be employed for dominant mutants, but material would need to be bulked from segregants that were phenotypically verified as homozygous in the F3 generation. (B) A three-way comparison between the bak1-5, bak1-5 mob1, and bak1-5 mob2 genomes identified the total number of unique SNPs in each genome.
