Figure 1.
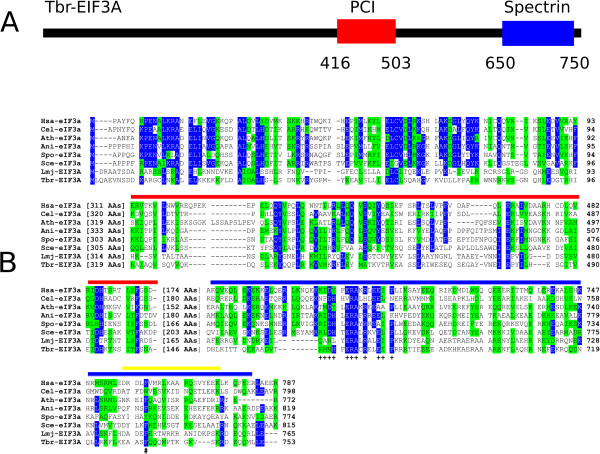
Conserved and diverged features between the trypanosomatid EIF3As and various eukaryotic orthologues. A Schematic representation of T. brucei EIF3A highlighting the PCI and Spectrin domains (red and blue boxes, respectively). B Amino acid sequence alignment comparing the N-terminus and the PCI and Spectrin domains from the various eIF3a homologues selected for this study. Amino acids identical in more than 60% of the sequences are shown with a blue background, while amino acids defined as similar, based on the BLOSUM 62 Matrix, in more than 60% of the sequences are highlighted with a green background. The red and blue lines define the PCI and Spectrin domains, respectively, whilst the yellow line indicates the segment implicated in the binding of human eIF3a to eIF3b and eIF3i [36]. The conserved residues within the Spectrin domain mentioned in the text are marked with “+”, whilst the “#” symbol marks the position of the conserved phenylalanine residue required for the interaction with human eIF3b.
