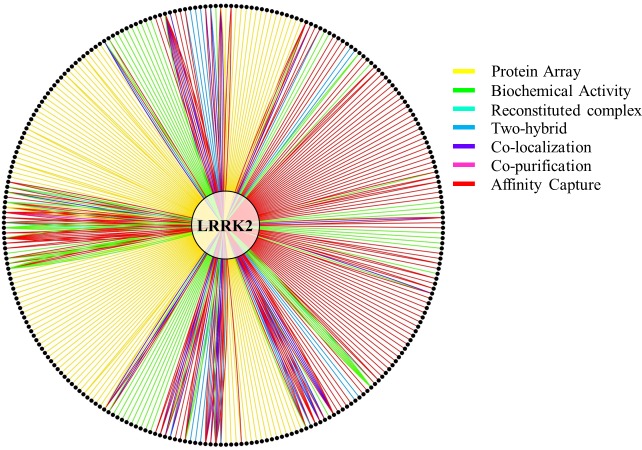
Figure 1. LRRK2 heterologous interactions as reported in the merged dataset.
LRRK2 in the middle of the graph is linked to candidate partners (black dots) through 422 annotations as described in the merged dataset. Some partners have one connection only; others have multiple connections based on the number of annotations. Different colours represent different methods used to infer the interaction; note that since IntAct and BioGRID use different classifications for the interaction detection method, a simplified and harmonized version has been applied to this figure to help the reader. In particular: (i) Affinity Capture stands for—Affinity Capture-Western and Affinity Capture-MS (in BioGRID)—Anti Bait Coimmunoprecipitation, Anti Tag Coimmunoprecipitation, Coimmunoprecipitation, Pull Down and Affinity Chromatography Technology (in IntAct); (ii) Biochemical Activity stands for—Biochemical Activity (in BioGRID)—Protein Kinase Assay (in IntAct); (iii) Co-localization stands for—Co-localization (in BioGRID)—Confocal microscopy and Fluorescence Microscopy (in IntAct); (iv) Reconstituted Complex stands for—Fluorescence Polarization Spectroscopy and Surface Plasmon Resonance (in IntAct).