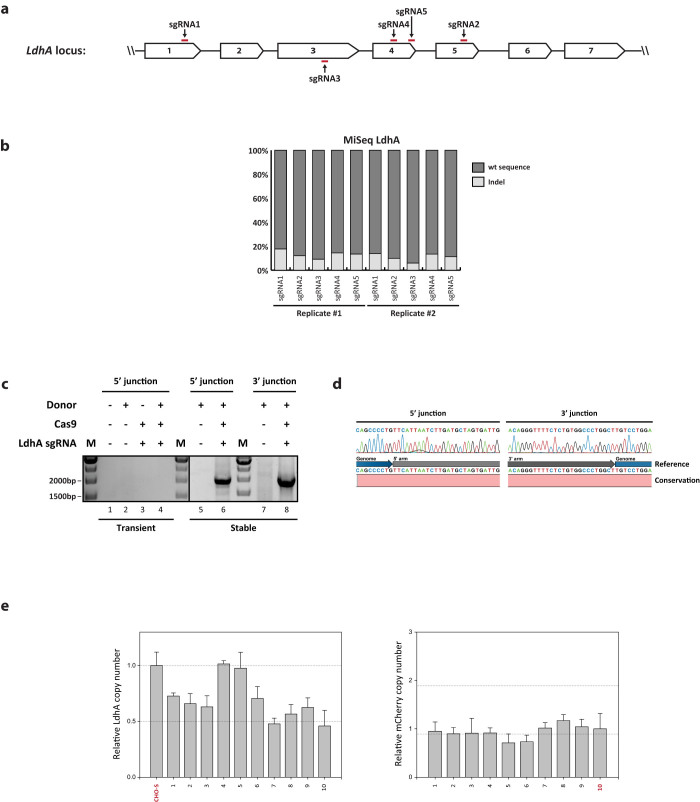
Figure 3. Targeted integration into LdhA locus using CRISPR/Cas9.
(a) Illustration of the five sgRNA target genomic sites in LdhA locus. (b) Indel frequency in LdhA locus analyzed by deep sequencing. Genomic DNA was extracted 3 days after transfection with plasmids expressing Cas9 gene and sgRNA. The genomic regions covering sgRNA target sites were amplified, then subjected to deep sequencing analysis using Miseq. The percentage of wt and indel sequences are described in the bar plot. The values from control samples transfected only with plasmid expressing Cas9 were subtracted from test samples. Investigation of target specific knock-in in transiently transfected and stable cell pools analyzed by (c) Agarose gel of 5′/3′ junction PCR (d) Sanger sequencing of the 5′/3′ junction PCR amplicons. Amplicons from the stable cell pool were purified and directly sequenced after PCR amplification. M, 1 kb DNA ladder. (e) Relative copy number of LdhA and mCherry regions in clonal cells, as described in Fig. 1(e).