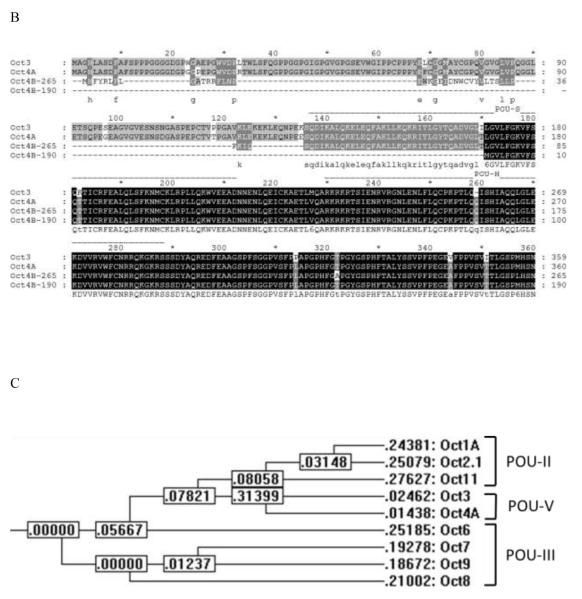
Figure 2.
A, B. Alignments of all human Octamer-binding factors (Octs) (A) and Oct3 and Oct4 isoforms (B). The GenBank protein identification numbers of these sequences are NP_002688 (Oct1A), NP_002689.1 (Oct2.1), NP_001153014.1 (Oct3), NP_002692.2 (Oct4A), NP_002690.3 (Oct6), NP_005595 (Oct7), NP_006227.1 (Oct8), NP_000298.2 (Oct9), NP_055167.2 (Oct11), CAA77952 (Oct4B-265) and NP_001167002.1 (Oct4B-190). The alignment was performed with the CLUSTAL-W program with an open gap cost = 10 and a gap extension cost = 0.2. Residues that are highlighted with black shading represent conserved amino acids, and the gray shading indicates 5 or more (A) or 2 or more (B) conserved residues at that position. Positions of the POU-specific domain (POU-S) and the POU homeodomain (POU-H) are given by dashed lines at the top of the sequence alignment. In addition, the conserved amino acids are shown on the bottom of the sequence alignment. C. Phylogenetic tree of Oct factors drawn from the multiple sequence alignment using CLUSTAL-W. The numbers represent tree weights.