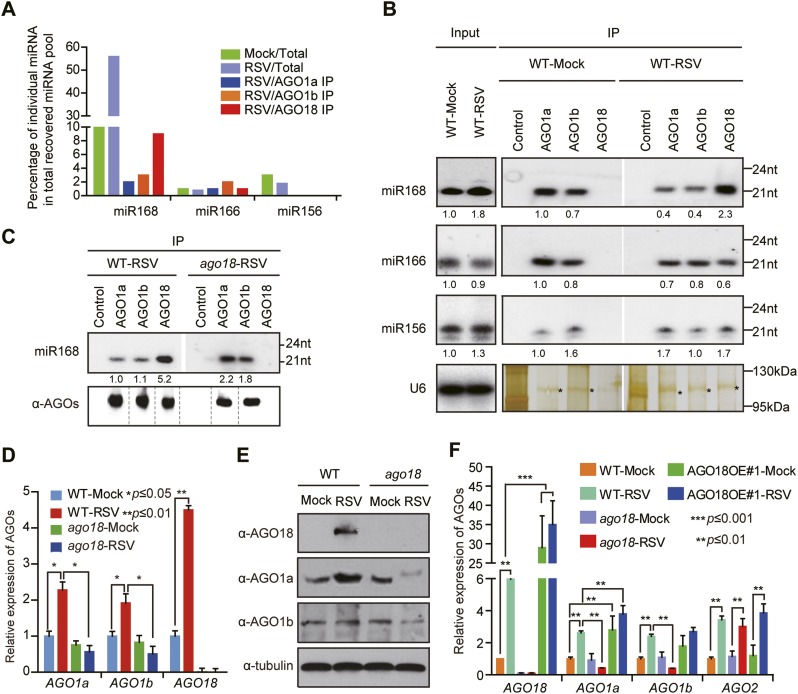
Figure 4. AGO18 competes with AGO1 for miR168 to up-regulate AGO1 upon viral infection.
(A) Percentage of deep sequencing reads matching the indicated miRNAs in total reads obtained from total extracts, AGO1a, AGO1b, and AGO18-associated small RNAs. Samples for deep sequencing were prepared from mock- or RSV-inoculated rice plants. (B) Detection of the indicated miRNAs in total extract (Input), AGO1a, AGO1b, and AGO18 complexes by Northern blot. The blots were stripped and reprobed for multiple times. The silver-stained gel shows that comparable amounts of different AGO complexes were used for RNA preparation. The asterisks indicate the positions of AGO proteins. The positions of RNA size markers are shown on the right of the blots. The RNA signals were quantified, and the relative values were calculated by comparison with those in total extracts or AGO1a complex prepared from mock-inoculated WT (arbitrarily set to 1.0). (C) Northern blot analysis showing miR168 from AGO1a, AGO1b, and AGO18 complexes in RSV-infected WT and ago18 plants (upper panel). Western blot gel shows that comparable amounts of different AGO complexes were used for RNA preparation (lower panel). (D) qRT-PCR analysis of the levels of AGO1a, AGO1b, and AGO18 in WT rice and ago18 mutants with or without RSV infection. The expression levels were normalized using the signal from OsEF-1a. The average (± standard deviation) values from three biological repeats of qRT-PCR are shown. (E) Western blot showing AGO1a, AGO1b, and AGO18 protein levels in WT and ago18 with or without RSV infection. Tubulin was probed and served as a loading control. (F) qRT-PCR analysis of the levels of AGO1a, AGO1b, AGO2, and AGO18 in the indicated plants. The expression levels were normalized using the signal from OsEF-1a. The average (± standard deviation) values from three biological repeats of qRT-PCR are shown.