Figure 2.
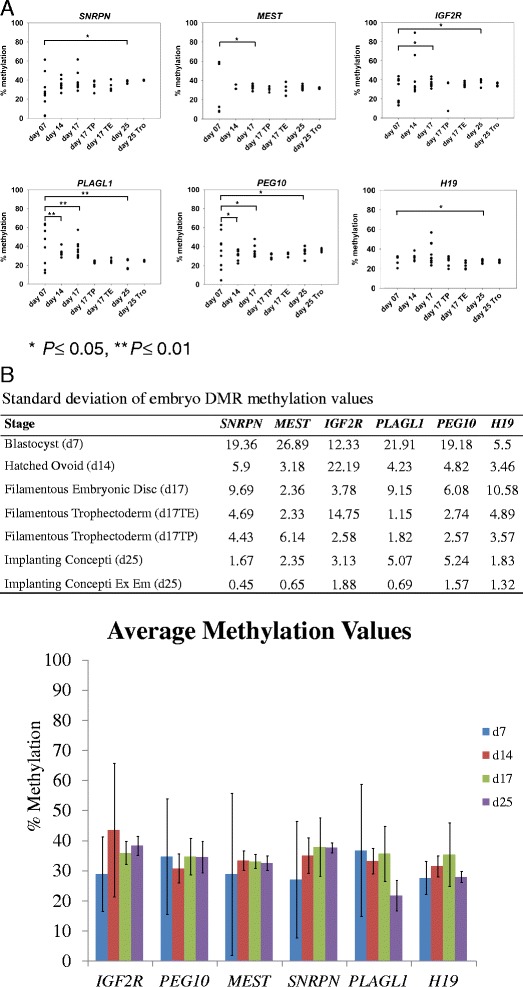
Pyrosequencing analysis of DNA methylation at six bovine imprinted differentially methylated regions during early embryogenesis. Bisulfite PCR and pyrosequencing was performed using genomic DNA isolated from embryos (n = 8 - 10) at four separate time points; blastocyst (Day 7), hatched ovoid embryos (Day 14), filamentous embryos (Day 17) and implanting conceptuses (Day 25). At Day 17, trophectoderm embryonic regions adjacent (TE) and peripheral to the embryonic disc (TP) were analysed. Trophectoderm samples at Day 25 were also included in the analysis. An overall trend of imprint stabilization can be observed for the six genes with increasing Days of embryonic development, post blastocyst. (A) Methylation values for SNRPN, MEST, IGF2R, PLAGL1, PEG10 and H19 at Day 7 had the most significant differences, when compared to Days 14, 17 or 25. (B) Top Panel; Standard deviations were calculated using individual methylation values (n = 4 - 9) for all genes in each group. Overall, standard deviations are greatest in group 1 (Day 7 blastocyst) with the smallest degree of variation occurring in the Day 25 implanting concepti (group 6) and Day 25 implanting concepti extra embryonic region. Bottom panel; Average methylation values, plotted with standard deviation values, for embryos isolated 7, 14, 17 and 25 Days post insemination (groups 1-3 and 6 from top panel).
