Figure 9.
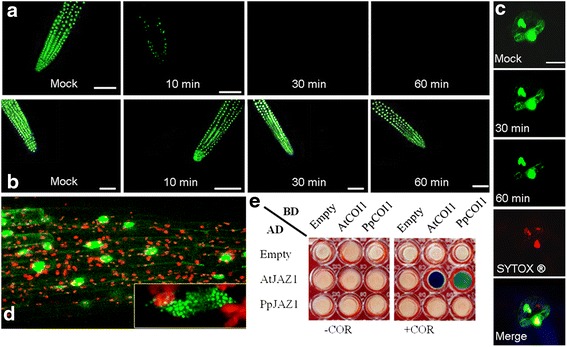
PpJAZ1 is not subject to JA-induced degradation, and does not interact with PpCOI1. (a-b) Arabidopsis roots expressing AtJAZ1-GFP (a) or PpJAZ1-GFP (b) were monitored with confocal laser scanning microscopy after treatment with water (mock) or MeJA (50 μM) for 10, 30 and 60 minutes. (c) The stability of PpJAZ1 was also examined in peach leaves transiently expressing PpJAZ1-GFP chimeric protein. Two guard cells showing GFP fluorescence were monitored after 10, 30 and 60 minutes of MeJA (50 μM) treatment (upper three panels). The location of the nucleus is indicated by SYTOX® red staining (fourth panel). The localization of PpJAZ1 in the nucleus was verified with the merged view (bottom panel), scale bars = 10 μm. (d) In Arabidopsis roots, PpJAZ1-GFP was present in the nucleus. At the bottom right is a magnified view of the nucleus showing nuclear protein bodies (NPBs) of different sizes. Scale bar = 50 μm. (e) Yeast two-hybrid assays revealed no interaction of PpJAZ1 with either PpCOI1 or AtCOI1. No blue colonies were observed in COR-free medium for all assays. In media supplemented with COR (200 μM)), however, only positive controls with AtJAZ1-AtCOI1 proteins showed dark blue colonies. PpCOI1 with AtJAZ1 gave rise to light blue colonies, indicative of lower interaction affinity between the corresponding proteins. JA, jasmonic acid; MeJA methyl jasmonate.
