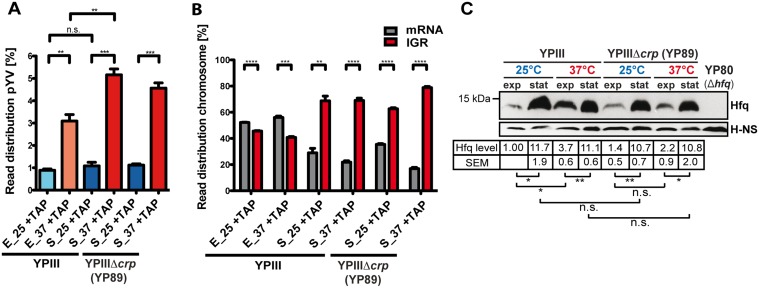
Fig 1. Global cDNA read count distribution and Hfq levels in Y. pseudotuberculosis during growth under environmental and infection-relevant conditions.
(A) Percentage of uniquely mapped reads of the virulence plasmid pYV obtained by RNA-seq of Y. pseudotuberculosis YPIII and YPIIIΔcrp cDNA libraries. Libraries were generated from TAP-treated (+TAP) rRNA depleted RNA obtained from cultures grown to exponential (E) or stationary (S) growth phase at 25°C (25) and 37°C (37) (for detailed statistics see S1 Dataset). The data represent the mean ± SEM from three independent biological replicates and were analyzed with Student’s t-test. **: P<0,01; ***: P<0,001; n.s.: not significant. (B) Percentage of uniquely mapped reads to annotated ORFs (mRNA) or intergenic regions (IGR) of the YPIII genome. IGR reads do not include 5’-UTRs of mRNAs, rRNAs and tRNAs, and mostly represent trans-encoded sRNAs. The data represent the mean ± SEM from three independent biological replicates and were analyzed with Student’s t-test. **: P<0,01; ***: P<0,001; ****: P<0.0001. (C) Growth phase- and temperature-responsive expression of Hfq in Y. pseudotuberculosis YPIII and the isogenic crp deletion strain YP89. Equal amounts of whole cell extracts of YPIII or YP89 grown to exponential (exp) or stationary phase (stat) at 25°C or 37°C were separated by SDS-PAGE prior to Western blotting using polyclonal Hfq or H-NS antibodies (loading control). As a negative control YP80 (YPIII Δhfq) was included. Relative protein amounts were quantified using ImageJ. The data represent the mean ± SEM from three independent experiments and were analyzed with Student’s t-test. *: P<0,05; **: P<0,01; n.s.: not significant.