Figure 1.
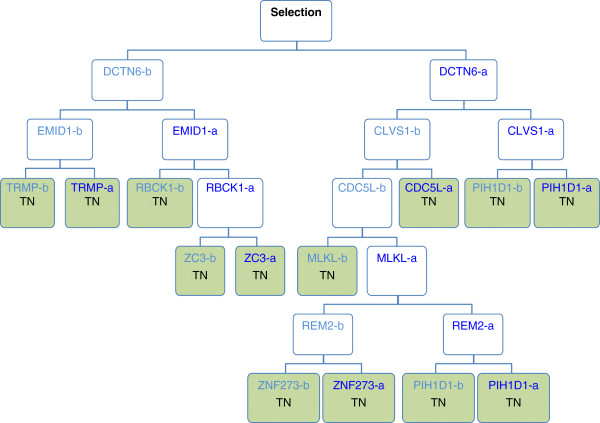
Depiction of a classification tree. For a tree within the random forest a bootstrap or subset of cell lines (samples) and genes (variables) is randomly selected. The gene from this random subset whose expression best divides the cell lines as cleanly as possible into sensitive and resistant is then selected. In the example above this gene was DCTN6. The two groupings or daughter nodes are not pure, they have some resistant and some sensitive cell lines, so a new subset of genes is selected and the gene which best divides the cell lines is selected; in the example above these genes were EMID1 and CLVS1. This recursive partitioning continues until terminal nodes (TN; green colored nodes) are reached, these being groupings in which only sensitive or resistant cell lines are found. In this manner classification trees consider the expression of one gene in relation to another (“co-acting genes”) in determining the biological response (i.e. TRAIL-sensitivity).
