Figure 1.
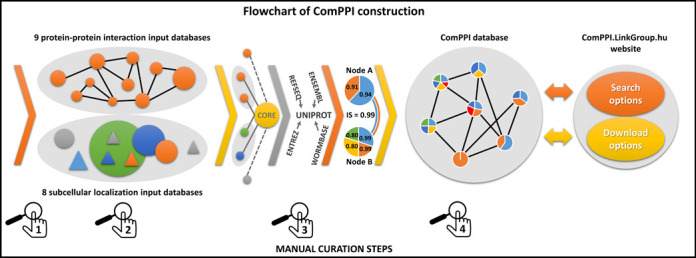
Flowchart of ComPPI construction highlighting the four curation steps. Constructing the ComPPI database we first checked the data content of 24 possible input databases for false entries, data inconsistence and compatible data structure in order to minimize the bias in ComPPI coming from the input sources (1). As a consequence we selected nine protein–protein interaction (BioGRID (29), CCSB (30), DiP (31), DroID (26), HPRD (27), IntAct (32), MatrixDB (18), MINT (33) and MIPS (28)) and eight subcellular localization databases (eSLDB (37), GO (19), Human Proteinpedia (34), LOCATE (38), MatrixDB (18), OrganelleDB (39), PA-GOSUB (36) and The Human Protein Atlas (35)) in order to integrate them into the ComPPI data set. The subcellular localization structure was manually annotated creating a hierarchic, non-redundant subcellular localization tree using >1600 GO cellular component terms (19) for the standardization of the different data resolution and naming conventions (2). All input databases were connected to the ComPPI core database with newly built interfaces in order to improve data consistency, to allow easy extensibility with new databases and to incorporate automatic database updates. As part of the curation steps the filtering efficiency of our newly built interfaces were tested on 200 random proteins for every input databases, and the interfaces were accepted only when all the requested false-entries and data content errors were filtered, in order to establish a more reliable content (Supplementary Table S3). During data integration, different protein naming conventions were mapped to the most reliable protein name. In this process we used publicly available mapping tables (UniProt (24) and HPRD (27)). For 30% of protein names we applied manually built mapping tables with the help of online ID cross-reference services (PICR (25) and Synergizer (http://llama.mshri.on.ca/synergizer/translate/)) (3). After data integration Localization and Interaction Scores were calculated (for detailed description see Figure 2). As an illustration we show the example of Figure 2 with two interacting proteins (nodes A and B corresponding to HSP 90-alpha A2 and Survivin, respectively) with shared cytosolic and nuclear localizations (light blue and orange). Node B has an additional membrane (yellow) subcellular localization and an extracellular localization (green). Numbers in the circles of nodes A and B refer to their Localization Scores. The Interaction Score of nodes A and B is 0.99 (see Figure 2 for details). The integrated ComPPI data set was manually revised by six independent experts (4). During the revision two of the six experts tested our database on 200 random proteins each to ensure high-quality control requirements, and searched for exact matches between the entries in the input sources and the ComPPI data set. All the experts searched for false entries, data inconsistency, protein name mapping errors in the downloadable data and tested the operation of the online services as well. After the revision we updated our source databases, their interfaces, the subcellular localization tree and the algorithm generating the downloadable data, in order to acquire all the changes proposed during the tests. As the final result, the webpage http://ComPPI.LinkGroup.hu is available for search and download options in order to extract the biological information in a user-friendly way.
